diff --git a/.gitignore b/.gitignore
index 96760692..ce31fcc6 100644
--- a/.gitignore
+++ b/.gitignore
@@ -1,5 +1,7 @@
output/
dataset/
+test_report.md
+test_results/
root.log
report.log
*.sqlite*
diff --git a/README.rst b/README.rst
index 4b1b4ef3..56417b0c 100644
--- a/README.rst
+++ b/README.rst
@@ -85,3 +85,15 @@ joining the **PyAutoFit** `Slack channel `_, where
Slack is invitation-only, so if you'd like to join send an `email `_ requesting an
invite.
+
+Build Configuration
+-------------------
+
+The ``config/`` directory contains two files used by the automated build and test system
+(CI, smoke tests, and pre-release checks). These are not relevant to normal workspace usage.
+
+- ``config/build/no_run.yaml`` — scripts to skip during automated runs. Each entry is a filename stem
+ or path pattern with an inline comment explaining why it is skipped.
+- ``config/build/env_vars.yaml`` — environment variables applied to each script during automated runs.
+ Defines default values (e.g. test mode, small datasets) and per-script overrides for scripts
+ that need different settings.
diff --git a/config/build/env_vars.yaml b/config/build/env_vars.yaml
new file mode 100644
index 00000000..d956d4e5
--- /dev/null
+++ b/config/build/env_vars.yaml
@@ -0,0 +1,21 @@
+# Per-script environment variable configuration for automated runs
+# (smoke tests, pre-release checks, CI).
+#
+# "defaults" are applied to every script on top of the inherited environment.
+# "overrides" selectively unset or replace vars for matching path patterns.
+#
+# Pattern convention (same as no_run.yaml):
+# - Patterns containing '/' do a substring match against the file path
+# - Patterns without '/' match the file stem exactly
+
+defaults:
+ PYAUTOFIT_TEST_MODE: "2" # 0=normal, 1=reduced iterations, 2=skip sampler (fastest)
+ PYAUTO_WORKSPACE_SMALL_DATASETS: "1" # Cap grids/masks to 15x15, reduce MGE gaussians
+ PYAUTO_DISABLE_CRITICAL_CAUSTICS: "1" # Skip critical curve/caustic overlays in plots
+ PYAUTO_DISABLE_JAX: "1" # Force use_jax=False, avoid JIT compilation overhead
+ PYAUTO_FAST_PLOTS: "1" # Skip tight_layout() in subplots
+ JAX_ENABLE_X64: "True" # Enable 64-bit precision in JAX
+ NUMBA_CACHE_DIR: "/tmp/numba_cache" # Writable cache dir for numba
+ MPLCONFIGDIR: "/tmp/matplotlib" # Writable config dir for matplotlib
+
+overrides: []
diff --git a/config/build/no_run.yaml b/config/build/no_run.yaml
new file mode 100644
index 00000000..81649f7c
--- /dev/null
+++ b/config/build/no_run.yaml
@@ -0,0 +1,17 @@
+# Scripts to skip during automated runs (smoke tests, pre-release checks, CI).
+# Each entry is matched against script paths:
+# - Entries with '/' do a substring match against the file path
+# - Entries without '/' match the file stem exactly
+# Add an inline # comment to document the reason for skipping.
+
+- GetDist # Cant get it to install, even in optional requirements.
+- Zeus # Test Model Iniitalization no good.
+- ZeusPlotter # Test Model Iniitalization no good.
+- UltraNestPlotter # Test Model Iniitalization no good.
+- DynestyPlotter # Test Model Iniitalization no good.
+- start_point # bug https://github.com/rhayes777/PyAutoFit/issues/1017
+- tutorial_8_astronomy_example # Requires dataset/howtofit/chapter_1/astro/simple/data.npy (not auto-generated)
+- searches/mle/PySwarmsGlobal # PySwarms does not support JAX.
+- searches/mle/PySwarmsLocal # PySwarms does not support JAX.
+- searches/nest/UltraNest # UltraNest does not support JAX.
+- plot/PySwarmsPlotter # PySwarms does not support JAX.
diff --git a/projects/cosmology/example_1_intro.py b/projects/cosmology/example_1_intro.py
index f72f1def..8f89a264 100644
--- a/projects/cosmology/example_1_intro.py
+++ b/projects/cosmology/example_1_intro.py
@@ -1,372 +1,368 @@
-"""
-Project: Cosmology
-==================
-
-This project uses the astrophysical phenomena of Strong Gravitational Lensing to illustrate multi-level model
-composition and fitting with **PyAutoFit**.
-
-A strong gravitational lens is a system where two (or more) galaxies align perfectly down our line of sight from Earth
-such that the foreground galaxy's mass deflects the light of a background source galaxy(s).
-
-When the alignment is just right and the lens is massive enough, the background source galaxy appears multiple
-times. The schematic below shows such a system, where light-rays from the source are deflected around the lens galaxy
-to the observer following multiple distinct paths.
-
-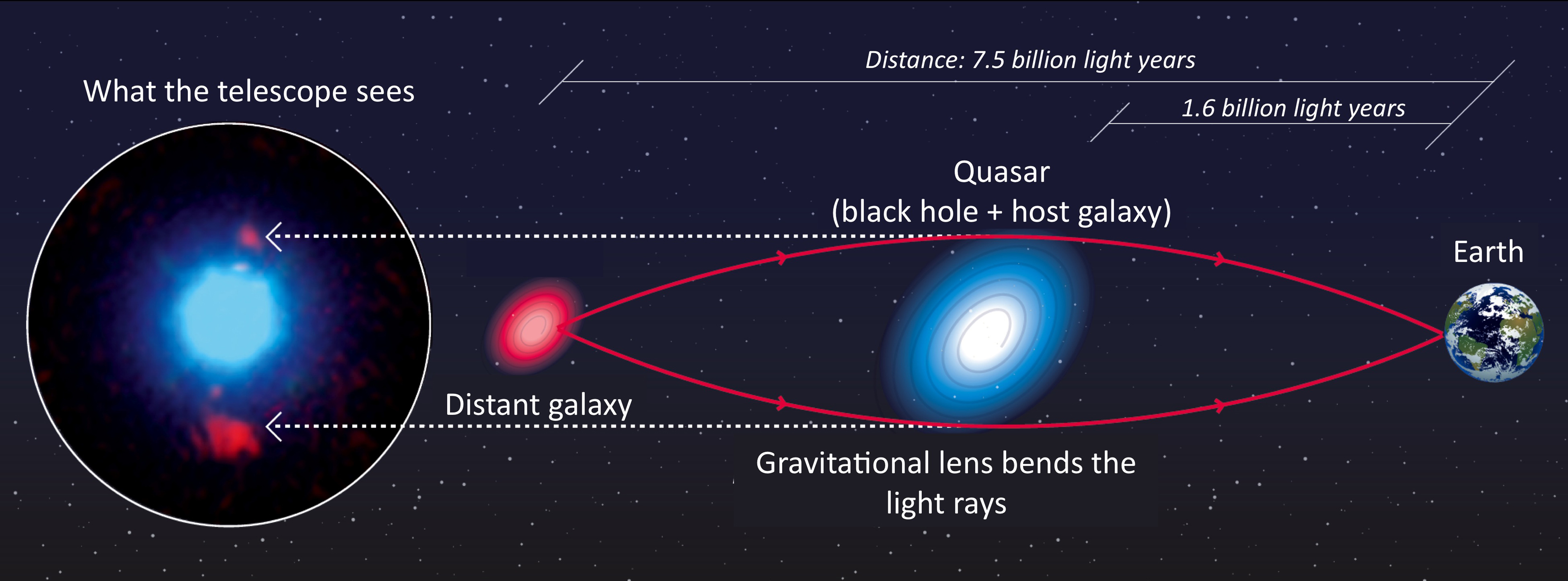
-**Credit: F. Courbin, S. G. Djorgovski, G. Meylan, et al., Caltech / EPFL / WMKO**
-https://www.cosmology.caltech.edu/~george/qsolens/
-
-As an observer, we don't see the source's true appearance (e.g. the red round blob of light). We only observe its
-light after it has been deflected and lensed by the foreground galaxies (e.g. as the two distinct red multiple images
- in the image on the left). We also observe the emission of the foreground galaxy (in blue).
-
-You can read more about gravitational lensing as the following link:
-
-https://en.wikipedia.org/wiki/Gravitational_lens
-
-__PyAutoLens__
-
-Strong gravitational lensing is the original science case that sparked the development of **PyAutoFit**, which is
-a spin off of our astronomy software **PyAutoLens** `https://github.com/Jammy2211/PyAutoLens`.
-
-We'll use **PyAutoLens** to illustrate how the tools we developed with **PyAutoFit** allowed us to
-ensure **PyAutoLens**'s model fitting tools were extensible, easy to maintain and enabled intuitive model composition.
-
-__Multi-Level Models__
-
-Strong lensing is a great case study for using **PyAutoFit**, due to the multi-component nature of how one composes
-a strong lens model. A strong lens model consists of light and mass models of each galaxy in the lens system, where
-each galaxy is a model in itself. The galaxies are combined into one overall "lens model", which in later tutorials
-we will show may also have a Cosmological model.
-
-This example project uses **PyAutoFit** to compose and fit models of a strong lens, in particular highlighting
-**PyAutoFits** multi-level model composition.
-
-__Strong Lens Modeling__
-
-The models are fitted to Hubble Space Telescope imaging of a real strong lens system and will allow us to come up
-with a description of how light is deflected on its path through the Universe.
-
-This project consists of two example scripts / notebooks:
-
- 1) `example_1_intro`: An introduction to strong lensing, and the various parts of the project's source code that are
- used to represent a strong lens galaxy.
-
- 2) `example_2_multi_level_model`: Using **PyAutoFit** to model a strong lens, with a strong emphasis on the
- multi-level model API.
-
-__This Example__
-
-This introduction primarily focuses on what strong lensing is, how we define the individual model-components and fit
-a strong lens model to data. It does not make much use of **PyAutoFit**, but it does provide a clear understanding of
-the model so that **PyAutoFit**'s use in example 2 is clear.
-
-Note that import `import src as cosmo`. The package `src` contains all the code we need for this example Cosmology
-use case, and can be thought of as the source-code you would write to perform model-fitting via **PyAutoFit** for your
-problem of interest.
-
-The module `src/__init__.py` performs a series of imports that are used throughout this lecture to provide convenient
-access to different parts of the source code.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import src as cosmo
-import matplotlib.pyplot as plt
-import numpy as np
-from scipy import signal
-from os import path
-
-"""
-__Plot__
-
-We will plot a lot of arrays of 2D data and grids of 2D coordinates in this example, so lets make a convenience
-functions.
-"""
-
-
-def plot_array(array, title=None, norm=None):
- plt.imshow(array, norm=norm)
- plt.colorbar()
- plt.title(title)
- plt.show()
- plt.close()
-
-
-def plot_grid(grid, title=None):
- plt.scatter(x=grid[:, :, 0], y=grid[:, :, 1], s=1)
- plt.title(title)
- plt.show()
- plt.close()
-
-
-"""
-__Data__
-
-First, lets load and plot Hubble Space Telescope imaging data of the strong gravitational lens called SDSSJ2303+1422,
-where this data includes:
-
- 1) The image of the strong lens, which is the data we'll fit.
- 2) The noise in every pixel of this image, which will be used when evaluating the log likelihood.
-
-__Masking__
-
-When fitting 2D imaging data, it is common to apply a mask which removes regions of the image that are not relevant to
-the model fitting.
-
-For example, when fitting the strong lens, we remove the edges of the image where the lens and source galaxy's light is
-not visible.
-
-In the strong lens image and noise map below, you can see this has already been performed, with the edge regions
-blank.
-"""
-dataset_path = path.join("projects", "cosmology", "dataset")
-
-data = np.load(file=path.join(dataset_path, "data.npy"))
-plot_array(array=data, title="Image of Strong Lens SDSSJ2303+1422")
-
-noise_map = np.load(file=path.join(dataset_path, "noise_map.npy"))
-plot_array(array=noise_map, title="Noise Map of Strong Lens SDSSJ2303+1422")
-
-"""
-In the image of the strong lens two distinct objects can be seen:
-
- 1) A central blob of light, corresponding to the foreground lens galaxy whose mass is responsible for deflecting light.
- 2) Two faint arcs of light in the bakcground, which is the lensed background source.
-
-__PSF__
-
-Another component of imaging data is the Point Spread Function (PSF), which describes how the light of the galaxies
-are blurred when they enter the Huble Space Telescope's.
-
-This is because diffraction occurs when the light enters HST's optics, causing the light to smear out. The PSF is
-a two dimensional array that describes this blurring via a 2D convolution kernel.
-
-When fitting the data below and in the `log_likelihood_function`, you'll see that the PSF is used when creating the
-model data. This is an example of how an `Analysis` class may be extended to include additional steps in the model
-fitting procedure.
-"""
-psf = np.load(file=path.join(dataset_path, "psf.npy"))
-plot_array(array=psf, title="Point Spread Function of Strong Lens SDSSJ2303+1422")
-
-
-"""
-__Grid__
-
-To perform strong lensing, we need a grid of (x,y) coordinates which we map throughout the Universe as if their path
-is deflected.
-
-For this, we create a simple 2D grid of coordinates below where the origin is (0.0, 0.0) and the size of
-a pixel is 0.05, which corresponds to the resolution of our image `data`.
-
-This grid only contains (y,x) coordinates within the cricular mask that was applied to the data, as we only need to
-perform ray-tracing within this region.
-"""
-grid = np.load(file=path.join(dataset_path, "grid.npy"))
-
-plot_grid(
- grid=grid,
- title="Cartesian grid of (x,y) coordinates aligned with strong lens dataset",
-)
-
-"""
-__Light Profiles__
-
-Our model of a strong lens must include a description of the light of each galaxy, which we call a "light profile".
-In the source-code of this example project, specifically the module `src/light_profiles.py` you will see there
-are two light profile classes named `LightDeVaucouleurs` and `LightExponential`.
-
-These Python classes are the model components we will use to represent each galaxy's light and they behave analogous
-to the `Gaussian` class seen in other tutorials. The input parameters of their `__init__` constructor (e.g. `centre`,
-`axis_ratio`, `angle`) are their model parameters that may be fitted for by a non-linear search.
-
-These classes also contain functions which create an image from the light profile if an input grid of (x,y) 2D
-coordinates are input, which we use below to create an image of a light profile.
-"""
-light_profile = cosmo.lp.LightExponential(
- centre=(0.01, 0.01), axis_ratio=0.7, angle=45.0, intensity=1.0, effective_radius=2.0
-)
-light_image = light_profile.image_from_grid(grid=grid)
-
-plot_array(array=light_image, title="Image of an Exponential light profile.")
-
-"""
-__Mass Profiles__
-
-Our model also includes the mass of the foreground lens galaxy, called a 'mass profile'. In the source-code of the
-example project, specifically the module `src/mass_profiles.py` you will see there is a mass profile class named
-`MassIsothermal`. Like the light profile, this will be a model-component **PyAutoFit** fits via a non-linear search.
-
-The class also contains functions which create the "deflections angles", which describe the angles by which light is
-deflected when it passes the mass of the foreground lens galaxy. These are subtracted from the (y,x) grid above to
-determine the original coordinates of the source galaxy before lensing.
-
-A higher mass galaxy, which bends light more, will have higher values of the deflection angles plotted below:
-"""
-mass_profile = cosmo.mp.MassIsothermal(
- centre=(0.01, 0.01), axis_ratio=0.7, angle=45.0, mass=0.5
-)
-mass_deflections = mass_profile.deflections_from_grid(grid=grid)
-
-plot_array(
- array=mass_deflections[:, :, 0],
- title="X-component of the deflection angles of a Isothermal mass profile.",
-)
-plot_array(
- array=mass_deflections[:, :, 1],
- title="Y-component of the deflection angles of a Isothermal mass profile.",
-)
-
-"""
-__Ray Tracing__
-
-The deflection angles describe how our (x,y) grid of coordinates are deflected by the mass of the foreground galaxy.
-
-We can therefore ray-trace the grid aligned with SDSSJ2303+1422 using the mass profile above and plot a grid of
-coordinates in the reference frame of before their light is gravitationally lensed:
-"""
-traced_grid = grid - mass_deflections
-
-plot_grid(grid=traced_grid, title="Cartesian grid of (x,y) traced coordinates.")
-
-"""
-By inputting this traced grid of (x,y) coordinates into our light profile, we can create an image of the galaxy as if
-it were gravitationally lensed by the mass profile.
-"""
-traced_light_image = light_profile.image_from_grid(grid=traced_grid)
-
-plot_array(
- array=traced_light_image,
- title="Image of a gravitationally lensed Exponential light profile.",
-)
-
-"""
-__Galaxy__
-
-In the `src/galaxy.py` module we define the `Galaxy` class, which is a collection of light and mass profiles at an
-input redshift. For strong lens modeling, we have to use `Galaxy` objects, as the redshifts define how ray-tracing is
-performed.
-
-We create two instances of the `Galaxy` class, representing the lens and source galaxies in a strong lens system.
-"""
-light_profile = cosmo.lp.LightDeVaucouleurs(
- centre=(0.01, 0.01), axis_ratio=0.9, angle=45.0, intensity=0.1, effective_radius=1.0
-)
-mass_profile = cosmo.mp.MassIsothermal(
- centre=(0.01, 0.01), axis_ratio=0.7, angle=45.0, mass=0.8
-)
-lens_galaxy = cosmo.Galaxy(
- redshift=0.5, light_profile_list=[light_profile], mass_profile_list=[mass_profile]
-)
-
-light_profile = cosmo.lp.LightExponential(
- centre=(0.1, 0.1), axis_ratio=0.5, angle=80.0, intensity=1.0, effective_radius=5.0
-)
-source_galaxy = cosmo.Galaxy(
- redshift=0.5, light_profile_list=[light_profile], mass_profile_list=[mass_profile]
-)
-
-"""
-A galaxy's image is the sum of its light profile images, and its deflection angles are the sum of its mass profile
-deflection angles.
-
-To illustrate this, lets plot the lens galaxy's light profile image:
-"""
-galaxy_image = lens_galaxy.image_from_grid(grid=grid)
-
-plot_array(array=galaxy_image, title="Image of the Lens Galaxy.")
-
-"""
-__Data Fitting__
-
-We can create an overall image of the strong lens by:
-
- 1) Creating an image of the lens galaxy.
- 2) Computing the deflection angles of the lens galaxy.
- 3) Ray-tracing light to the source galaxy reference frame and using its light profile to make its image.
-"""
-lens_image = lens_galaxy.image_from_grid(grid=grid)
-lens_deflections = lens_galaxy.deflections_from_grid(grid=grid)
-
-traced_grid = grid - lens_deflections
-
-source_image = source_galaxy.image_from_grid(grid=traced_grid)
-
-# The grid has zeros at its edges, which produce nans in the model image.
-# These lead to an ill-defined log likelihood, so we set them to zero.
-overall_image = np.nan_to_num(overall_image)
-
-plot_array(array=overall_image, title="Image of the overall Strong Lens System.")
-
-"""
-__Model Data__
-
-To produce the `model_data`, we now convolution the overall image with the Point Spread Function (PSF) of our
-observations. This blurs the image to simulate the telescope optics and pixelization used to observe the image.
-"""
-model_data = signal.convolve2d(overall_image, psf, mode="same")
-
-
-plot_array(array=model_data, title="Image of the overall Strong Lens System.")
-
-"""
-By subtracting this model image from the data, we can create a 2D residual map. This is equivalent to the residual maps
-we made in the 1D Gaussian examples, except for 2D imaging data.
-
-Clearly, the random lens model we used in this example does not provide a good fit to SDSSJ2303+1422.
-"""
-residual_map = data - model_data
-
-plot_array(array=residual_map, title="Residual Map of fit to SDSSJ2303+1422")
-
-"""
-Just like we did for the 1D `Gaussian` fitting examples, we can use the noise-map to compute the normalized residuals
-and chi-squared map of the lens model.
-"""
-# The circular masking introduces zeros at the edge of the noise-map,
-# which can lead to divide-by-zero errors.
-# We set these values to 1.0e8, to ensure they do not contribute to the log likelihood.
-noise_map_fit = noise_map
-noise_map_fit[noise_map == 0] = 1.0e8
-
-normalized_residual_map = residual_map / noise_map_fit
-
-chi_squared_map = (normalized_residual_map) ** 2.0
-
-plot_array(
- array=normalized_residual_map,
- title="Normalized Residual Map of fit to SDSSJ2303+1422",
-)
-plot_array(array=chi_squared_map, title="Chi Squared Map of fit to SDSSJ2303+1422")
-
-"""
-Finally, we can compute the `log_likelihood` of this lens model, which we will use in the next example to fit the
-lens model to data with a non-linear search.
-"""
-chi_squared = np.sum(chi_squared_map)
-noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
-
-log_likelihood = -0.5 * (chi_squared + noise_normalization)
-
-print(log_likelihood)
-
-"""
-__Wrap Up__
-
-In this example, we introduced the astrophysical phenomena of strong gravitational lensing, and gave an overview of how
-one can create a model for a strong lens system and fit it to imaging data.
-
-We ended by defining the log likelihood of the model-fit, which will form the `log_likelihood_function` of the
-`Analysis` class we use in the next example, which fits this strong lens using **PyAutoFit**.
-
-There is one thing you should think about, how would we translate the above classes (e.g. `LightExponential`,
-`MassIsothermal` and `Galaxy`) using the **PyAutoFit** `Model` and `Collection` objects? The `Galaxy` class contained
-instances of the light and mass profile classes, meaning the standard use of the `Model` and `Collection` objects could
-not handle this.
-
-This is where multi-level models come in, as will be shown in the next example!
-"""
+"""
+Project: Cosmology
+==================
+
+This project uses the astrophysical phenomena of Strong Gravitational Lensing to illustrate multi-level model
+composition and fitting with **PyAutoFit**.
+
+A strong gravitational lens is a system where two (or more) galaxies align perfectly down our line of sight from Earth
+such that the foreground galaxy's mass deflects the light of a background source galaxy(s).
+
+When the alignment is just right and the lens is massive enough, the background source galaxy appears multiple
+times. The schematic below shows such a system, where light-rays from the source are deflected around the lens galaxy
+to the observer following multiple distinct paths.
+
+
+**Credit: F. Courbin, S. G. Djorgovski, G. Meylan, et al., Caltech / EPFL / WMKO**
+https://www.cosmology.caltech.edu/~george/qsolens/
+
+As an observer, we don't see the source's true appearance (e.g. the red round blob of light). We only observe its
+light after it has been deflected and lensed by the foreground galaxies (e.g. as the two distinct red multiple images
+ in the image on the left). We also observe the emission of the foreground galaxy (in blue).
+
+You can read more about gravitational lensing as the following link:
+
+https://en.wikipedia.org/wiki/Gravitational_lens
+
+__PyAutoLens__
+
+Strong gravitational lensing is the original science case that sparked the development of **PyAutoFit**, which is
+a spin off of our astronomy software **PyAutoLens** `https://github.com/Jammy2211/PyAutoLens`.
+
+We'll use **PyAutoLens** to illustrate how the tools we developed with **PyAutoFit** allowed us to
+ensure **PyAutoLens**'s model fitting tools were extensible, easy to maintain and enabled intuitive model composition.
+
+__Multi-Level Models__
+
+Strong lensing is a great case study for using **PyAutoFit**, due to the multi-component nature of how one composes
+a strong lens model. A strong lens model consists of light and mass models of each galaxy in the lens system, where
+each galaxy is a model in itself. The galaxies are combined into one overall "lens model", which in later tutorials
+we will show may also have a Cosmological model.
+
+This example project uses **PyAutoFit** to compose and fit models of a strong lens, in particular highlighting
+**PyAutoFits** multi-level model composition.
+
+__Strong Lens Modeling__
+
+The models are fitted to Hubble Space Telescope imaging of a real strong lens system and will allow us to come up
+with a description of how light is deflected on its path through the Universe.
+
+This project consists of two example scripts / notebooks:
+
+ 1) `example_1_intro`: An introduction to strong lensing, and the various parts of the project's source code that are
+ used to represent a strong lens galaxy.
+
+ 2) `example_2_multi_level_model`: Using **PyAutoFit** to model a strong lens, with a strong emphasis on the
+ multi-level model API.
+
+__This Example__
+
+This introduction primarily focuses on what strong lensing is, how we define the individual model-components and fit
+a strong lens model to data. It does not make much use of **PyAutoFit**, but it does provide a clear understanding of
+the model so that **PyAutoFit**'s use in example 2 is clear.
+
+Note that import `import src as cosmo`. The package `src` contains all the code we need for this example Cosmology
+use case, and can be thought of as the source-code you would write to perform model-fitting via **PyAutoFit** for your
+problem of interest.
+
+The module `src/__init__.py` performs a series of imports that are used throughout this lecture to provide convenient
+access to different parts of the source code.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import src as cosmo
+import matplotlib.pyplot as plt
+import numpy as np
+from scipy import signal
+from os import path
+
+"""
+__Plot__
+
+We will plot a lot of arrays of 2D data and grids of 2D coordinates in this example, so lets make a convenience
+functions.
+"""
+
+
+def plot_array(array, title=None, norm=None):
+ plt.imshow(array, norm=norm)
+ plt.colorbar()
+ plt.title(title)
+ plt.show()
+ plt.close()
+
+
+def plot_grid(grid, title=None):
+ plt.scatter(x=grid[:, :, 0], y=grid[:, :, 1], s=1)
+ plt.title(title)
+ plt.show()
+ plt.close()
+
+
+"""
+__Data__
+
+First, lets load and plot Hubble Space Telescope imaging data of the strong gravitational lens called SDSSJ2303+1422,
+where this data includes:
+
+ 1) The image of the strong lens, which is the data we'll fit.
+ 2) The noise in every pixel of this image, which will be used when evaluating the log likelihood.
+
+__Masking__
+
+When fitting 2D imaging data, it is common to apply a mask which removes regions of the image that are not relevant to
+the model fitting.
+
+For example, when fitting the strong lens, we remove the edges of the image where the lens and source galaxy's light is
+not visible.
+
+In the strong lens image and noise map below, you can see this has already been performed, with the edge regions
+blank.
+"""
+dataset_path = path.join("projects", "cosmology", "dataset")
+
+data = np.load(file=path.join(dataset_path, "data.npy"))
+plot_array(array=data, title="Image of Strong Lens SDSSJ2303+1422")
+
+noise_map = np.load(file=path.join(dataset_path, "noise_map.npy"))
+plot_array(array=noise_map, title="Noise Map of Strong Lens SDSSJ2303+1422")
+
+"""
+In the image of the strong lens two distinct objects can be seen:
+
+ 1) A central blob of light, corresponding to the foreground lens galaxy whose mass is responsible for deflecting light.
+ 2) Two faint arcs of light in the bakcground, which is the lensed background source.
+
+__PSF__
+
+Another component of imaging data is the Point Spread Function (PSF), which describes how the light of the galaxies
+are blurred when they enter the Huble Space Telescope's.
+
+This is because diffraction occurs when the light enters HST's optics, causing the light to smear out. The PSF is
+a two dimensional array that describes this blurring via a 2D convolution kernel.
+
+When fitting the data below and in the `log_likelihood_function`, you'll see that the PSF is used when creating the
+model data. This is an example of how an `Analysis` class may be extended to include additional steps in the model
+fitting procedure.
+"""
+psf = np.load(file=path.join(dataset_path, "psf.npy"))
+plot_array(array=psf, title="Point Spread Function of Strong Lens SDSSJ2303+1422")
+
+
+"""
+__Grid__
+
+To perform strong lensing, we need a grid of (x,y) coordinates which we map throughout the Universe as if their path
+is deflected.
+
+For this, we create a simple 2D grid of coordinates below where the origin is (0.0, 0.0) and the size of
+a pixel is 0.05, which corresponds to the resolution of our image `data`.
+
+This grid only contains (y,x) coordinates within the cricular mask that was applied to the data, as we only need to
+perform ray-tracing within this region.
+"""
+grid = np.load(file=path.join(dataset_path, "grid.npy"))
+
+plot_grid(
+ grid=grid,
+ title="Cartesian grid of (x,y) coordinates aligned with strong lens dataset",
+)
+
+"""
+__Light Profiles__
+
+Our model of a strong lens must include a description of the light of each galaxy, which we call a "light profile".
+In the source-code of this example project, specifically the module `src/light_profiles.py` you will see there
+are two light profile classes named `LightDeVaucouleurs` and `LightExponential`.
+
+These Python classes are the model components we will use to represent each galaxy's light and they behave analogous
+to the `Gaussian` class seen in other tutorials. The input parameters of their `__init__` constructor (e.g. `centre`,
+`axis_ratio`, `angle`) are their model parameters that may be fitted for by a non-linear search.
+
+These classes also contain functions which create an image from the light profile if an input grid of (x,y) 2D
+coordinates are input, which we use below to create an image of a light profile.
+"""
+light_profile = cosmo.lp.LightExponential(
+ centre=(0.01, 0.01), axis_ratio=0.7, angle=45.0, intensity=1.0, effective_radius=2.0
+)
+light_image = light_profile.image_from_grid(grid=grid)
+
+plot_array(array=light_image, title="Image of an Exponential light profile.")
+
+"""
+__Mass Profiles__
+
+Our model also includes the mass of the foreground lens galaxy, called a 'mass profile'. In the source-code of the
+example project, specifically the module `src/mass_profiles.py` you will see there is a mass profile class named
+`MassIsothermal`. Like the light profile, this will be a model-component **PyAutoFit** fits via a non-linear search.
+
+The class also contains functions which create the "deflections angles", which describe the angles by which light is
+deflected when it passes the mass of the foreground lens galaxy. These are subtracted from the (y,x) grid above to
+determine the original coordinates of the source galaxy before lensing.
+
+A higher mass galaxy, which bends light more, will have higher values of the deflection angles plotted below:
+"""
+mass_profile = cosmo.mp.MassIsothermal(
+ centre=(0.01, 0.01), axis_ratio=0.7, angle=45.0, mass=0.5
+)
+mass_deflections = mass_profile.deflections_from_grid(grid=grid)
+
+plot_array(
+ array=mass_deflections[:, :, 0],
+ title="X-component of the deflection angles of a Isothermal mass profile.",
+)
+plot_array(
+ array=mass_deflections[:, :, 1],
+ title="Y-component of the deflection angles of a Isothermal mass profile.",
+)
+
+"""
+__Ray Tracing__
+
+The deflection angles describe how our (x,y) grid of coordinates are deflected by the mass of the foreground galaxy.
+
+We can therefore ray-trace the grid aligned with SDSSJ2303+1422 using the mass profile above and plot a grid of
+coordinates in the reference frame of before their light is gravitationally lensed:
+"""
+traced_grid = grid - mass_deflections
+
+plot_grid(grid=traced_grid, title="Cartesian grid of (x,y) traced coordinates.")
+
+"""
+By inputting this traced grid of (x,y) coordinates into our light profile, we can create an image of the galaxy as if
+it were gravitationally lensed by the mass profile.
+"""
+traced_light_image = light_profile.image_from_grid(grid=traced_grid)
+
+plot_array(
+ array=traced_light_image,
+ title="Image of a gravitationally lensed Exponential light profile.",
+)
+
+"""
+__Galaxy__
+
+In the `src/galaxy.py` module we define the `Galaxy` class, which is a collection of light and mass profiles at an
+input redshift. For strong lens modeling, we have to use `Galaxy` objects, as the redshifts define how ray-tracing is
+performed.
+
+We create two instances of the `Galaxy` class, representing the lens and source galaxies in a strong lens system.
+"""
+light_profile = cosmo.lp.LightDeVaucouleurs(
+ centre=(0.01, 0.01), axis_ratio=0.9, angle=45.0, intensity=0.1, effective_radius=1.0
+)
+mass_profile = cosmo.mp.MassIsothermal(
+ centre=(0.01, 0.01), axis_ratio=0.7, angle=45.0, mass=0.8
+)
+lens_galaxy = cosmo.Galaxy(
+ redshift=0.5, light_profile_list=[light_profile], mass_profile_list=[mass_profile]
+)
+
+light_profile = cosmo.lp.LightExponential(
+ centre=(0.1, 0.1), axis_ratio=0.5, angle=80.0, intensity=1.0, effective_radius=5.0
+)
+source_galaxy = cosmo.Galaxy(
+ redshift=0.5, light_profile_list=[light_profile], mass_profile_list=[mass_profile]
+)
+
+"""
+A galaxy's image is the sum of its light profile images, and its deflection angles are the sum of its mass profile
+deflection angles.
+
+To illustrate this, lets plot the lens galaxy's light profile image:
+"""
+galaxy_image = lens_galaxy.image_from_grid(grid=grid)
+
+plot_array(array=galaxy_image, title="Image of the Lens Galaxy.")
+
+"""
+__Data Fitting__
+
+We can create an overall image of the strong lens by:
+
+ 1) Creating an image of the lens galaxy.
+ 2) Computing the deflection angles of the lens galaxy.
+ 3) Ray-tracing light to the source galaxy reference frame and using its light profile to make its image.
+"""
+lens_image = lens_galaxy.image_from_grid(grid=grid)
+lens_deflections = lens_galaxy.deflections_from_grid(grid=grid)
+
+traced_grid = grid - lens_deflections
+
+source_image = source_galaxy.image_from_grid(grid=traced_grid)
+
+# The grid has zeros at its edges, which produce nans in the model image.
+# These lead to an ill-defined log likelihood, so we set them to zero.
+overall_image = np.nan_to_num(overall_image)
+
+plot_array(array=overall_image, title="Image of the overall Strong Lens System.")
+
+"""
+__Model Data__
+
+To produce the `model_data`, we now convolution the overall image with the Point Spread Function (PSF) of our
+observations. This blurs the image to simulate the telescope optics and pixelization used to observe the image.
+"""
+model_data = signal.convolve2d(overall_image, psf, mode="same")
+
+
+plot_array(array=model_data, title="Image of the overall Strong Lens System.")
+
+"""
+By subtracting this model image from the data, we can create a 2D residual map. This is equivalent to the residual maps
+we made in the 1D Gaussian examples, except for 2D imaging data.
+
+Clearly, the random lens model we used in this example does not provide a good fit to SDSSJ2303+1422.
+"""
+residual_map = data - model_data
+
+plot_array(array=residual_map, title="Residual Map of fit to SDSSJ2303+1422")
+
+"""
+Just like we did for the 1D `Gaussian` fitting examples, we can use the noise-map to compute the normalized residuals
+and chi-squared map of the lens model.
+"""
+# The circular masking introduces zeros at the edge of the noise-map,
+# which can lead to divide-by-zero errors.
+# We set these values to 1.0e8, to ensure they do not contribute to the log likelihood.
+noise_map_fit = noise_map
+noise_map_fit[noise_map == 0] = 1.0e8
+
+normalized_residual_map = residual_map / noise_map_fit
+
+chi_squared_map = (normalized_residual_map) ** 2.0
+
+plot_array(
+ array=normalized_residual_map,
+ title="Normalized Residual Map of fit to SDSSJ2303+1422",
+)
+plot_array(array=chi_squared_map, title="Chi Squared Map of fit to SDSSJ2303+1422")
+
+"""
+Finally, we can compute the `log_likelihood` of this lens model, which we will use in the next example to fit the
+lens model to data with a non-linear search.
+"""
+chi_squared = np.sum(chi_squared_map)
+noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
+
+log_likelihood = -0.5 * (chi_squared + noise_normalization)
+
+print(log_likelihood)
+
+"""
+__Wrap Up__
+
+In this example, we introduced the astrophysical phenomena of strong gravitational lensing, and gave an overview of how
+one can create a model for a strong lens system and fit it to imaging data.
+
+We ended by defining the log likelihood of the model-fit, which will form the `log_likelihood_function` of the
+`Analysis` class we use in the next example, which fits this strong lens using **PyAutoFit**.
+
+There is one thing you should think about, how would we translate the above classes (e.g. `LightExponential`,
+`MassIsothermal` and `Galaxy`) using the **PyAutoFit** `Model` and `Collection` objects? The `Galaxy` class contained
+instances of the light and mass profile classes, meaning the standard use of the `Model` and `Collection` objects could
+not handle this.
+
+This is where multi-level models come in, as will be shown in the next example!
+"""
diff --git a/projects/cosmology/example_2_multi_level_model.py b/projects/cosmology/example_2_multi_level_model.py
index e19f75f5..63c2eea5 100644
--- a/projects/cosmology/example_2_multi_level_model.py
+++ b/projects/cosmology/example_2_multi_level_model.py
@@ -1,366 +1,362 @@
-"""
-Project: Cosmology
-==================
-
-This project uses the astrophysical phenomena of Strong Gravitational Lensing to illustrate basic and advanced model
-composition and fitting with **PyAutoFit**. The first tutorial described what a strong gravitational lens is and how
-we build and fit a model of one.
-
-In this example, we use **PyAutoFit**'s multi-level models to compose a strong lens model consisting of a lens and
-source galaxy, and fit it to the data on SDSSJ2303+1422.
-
-__Config Path__
-
-We first set up the path to this projects config files, which is located at `autofit_workspace/projects/cosmology/config`.
-
-This includes the default priors for the lens model, check it out!
-"""
-
-import os
-from os import path
-from autoconf import conf
-
-cwd = os.getcwd()
-config_path = path.join(cwd, "projects", "cosmology", "config")
-conf.instance.push(new_path=config_path)
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import autofit as af
-import src as cosmo
-import matplotlib.pyplot as plt
-import numpy as np
-
-"""
-__Plot__
-
-First, lets again define the plotting convenience functions we used in the previous example.
-"""
-
-
-def plot_array(array, title=None, norm=None):
- plt.imshow(array, norm=norm)
- plt.colorbar()
- plt.title(title)
- plt.show()
- plt.close()
-
-
-def plot_grid(grid, title=None):
- plt.scatter(x=grid[:, :, 0], y=grid[:, :, 1], s=1)
- plt.title(title)
- plt.show()
- plt.close()
-
-
-"""
-__Data__
-
-Now lets load and plot Hubble Space Telescope imaging data of the strong gravitational lens SDSSJ2303+1422.
-"""
-dataset_path = path.join("projects", "cosmology", "dataset")
-
-data = np.load(file=path.join(dataset_path, "data.npy"))
-plot_array(array=data, title="Image of Strong Lens SDSSJ2303+1422")
-
-noise_map = np.load(file=path.join(dataset_path, "noise_map.npy"))
-plot_array(array=noise_map, title="Noise Map of Strong Lens SDSSJ2303+1422")
-
-psf = np.load(file=path.join(dataset_path, "psf.npy"))
-plot_array(array=psf, title="Point Spread Function of Strong Lens SDSSJ2303+1422")
-
-grid = np.load(file=path.join(dataset_path, "grid.npy"))
-
-plot_grid(
- grid=grid,
- title="Cartesian grid of (x,y) coordinates aligned with strong lens dataset",
-)
-
-"""
-__Multi-level Model__
-
-In the previous example, we saw that we can use instances of the light profiles, mass profiles and galaxy objects to
-perform strong lens ray-tracing calculations:
-"""
-light_profile = cosmo.lp.LightDeVaucouleurs(
- centre=(0.01, 0.01), axis_ratio=0.7, angle=45.0, intensity=1.0, effective_radius=2.0
-)
-mass_profile = cosmo.mp.MassIsothermal(
- centre=(0.01, 0.01), axis_ratio=0.7, angle=45.0, mass=0.5
-)
-galaxy = cosmo.Galaxy(
- redshift=0.5, light_profile_list=[light_profile], mass_profile_list=[mass_profile]
-)
-
-"""
-In this example, we want to perform a model-fit using a non-linear search, where the `Galaxy` is a `Model`, but it
-contains model subcomponents that are its individual light and mass profiles.
-
-Here is a pictoral representation of the model:
-
-
-
-__Model Composition__
-
-How do we compose a strong lens model where a `Galaxy` is a `Model`, but it contains the light and mass profiles
-as `Model` themselves?
-
-We use **PyAutoFit**'s multi-level model composition:
-"""
-lens = af.Model(
- cls=cosmo.Galaxy, # The overall model object uses this input.
- redshift=0.5,
- light_profile_list=[
- af.Model(cosmo.lp.LightDeVaucouleurs)
- ], # These will be subcomponents of the model.
- mass_profile_list=[
- af.Model(cosmo.mp.MassIsothermal)
- ], # These will be subcomponents of the model.
-)
-
-print(lens.info)
-
-"""
-Lets consider what is going on here:
-
- 1) We use a `Model` to create the overall model component. The `cls` input is the `Galaxy` class, therefore the
- overall model that is created is a `Galaxy`.
-
- 2) **PyAutoFit** next inspects whether the key word argument inputs to the `Model` match any of the `__init__`
- constructor arguments of the `Galaxy` class. This determine if these inputs are to be composed as model
- subcomponents of the overall `Galaxy` model.
-
- 3) **PyAutoFit** matches the `light_profile_list` and `mass_profile_list` inputs, noting they are passed as separate
- lists containing the `LightDeVaucouleurs` and `MassIsothermal` class. They are both created as subcomponents of
- the overall `Galaxy` model.
-
- 4) It also matches the `redshift` input, making it a fixed value of 0.5 for the model and not treating it as a
- free parameter.
-
-We can confirm this by printing the `total_free_parameters` of the lens, and noting it is 11 (6 parameters for
-the `LightDeVaucouleurs` and 5 for the `MassIsothermal`).
-"""
-print(lens.total_free_parameters)
-print(lens.light_profile_list[0].total_free_parameters)
-print(lens.mass_profile_list[0].total_free_parameters)
-
-"""
-The `lens` behaves exactly like the model-components we are used to previously. For example, we can unpack its
-individual parameters to customize the model, where below we:
-
- 1) Fix the light and mass profiles to the centre (0.0, 0.0).
- 2) Customize the prior on the light profile `axis_ratio`.
- 3) Fix the `axis_ratio` of the mass profile to 0.8.
-"""
-
-lens.light_profile_list[0].centre = (0.0, 0.0)
-lens.light_profile_list[0].axis_ratio = af.UniformPrior(
- lower_limit=0.7, upper_limit=0.9
-)
-lens.light_profile_list[0].angle = af.UniformPrior(lower_limit=0.0, upper_limit=180.0)
-lens.light_profile_list[0].intensity = af.LogUniformPrior(
- lower_limit=1e-4, upper_limit=1e4
-)
-lens.light_profile_list[0].effective_radius = af.UniformPrior(
- lower_limit=0.0, upper_limit=5.0
-)
-
-lens.mass_profile_list[0].centre = (0.0, 0.0)
-lens.mass_profile_list[0].axis_ratio = 0.8
-lens.mass_profile_list[0].angle = af.UniformPrior(lower_limit=0.0, upper_limit=180.0)
-lens.mass_profile_list[0].mass = af.UniformPrior(lower_limit=0.0, upper_limit=2.0)
-
-print(lens.info)
-
-"""
-__Alternative API__
-
-We can create the `Galaxy` model component with the exact same customization by creating each profile as a `Model` and
-passing these to the galaxy `Model`.
-"""
-light = af.Model(cosmo.lp.LightDeVaucouleurs)
-
-light.centre = af.UniformPrior(lower_limit=-0.05, upper_limit=0.05)
-light.axis_ratio = af.UniformPrior(lower_limit=0.7, upper_limit=0.9)
-light.angle = af.UniformPrior(lower_limit=0.0, upper_limit=180.0)
-light.intensity = af.LogUniformPrior(lower_limit=1e-4, upper_limit=1e4)
-light.effective_radius = af.UniformPrior(lower_limit=0.0, upper_limit=5.0)
-
-
-mass = af.Model(cosmo.mp.MassIsothermal)
-
-mass.centre = (0.0, 0.0)
-mass.axis_ratio = af.UniformPrior(lower_limit=0.7, upper_limit=1.0)
-mass.angle = af.UniformPrior(lower_limit=0.0, upper_limit=180.0)
-mass.mass = af.UniformPrior(lower_limit=0.0, upper_limit=4.0)
-
-lens = af.Model(
- cosmo.Galaxy, redshift=0.5, light_profile_list=[light], mass_profile_list=[mass]
-)
-
-print(lens.info)
-
-"""
-We can now create a model of our source galaxy using the same API.
-"""
-light = af.Model(cosmo.lp.LightExponential)
-
-light.centre.centre_0 = af.GaussianPrior(mean=0.0, sigma=0.3)
-light.centre.centre_1 = af.GaussianPrior(mean=0.0, sigma=0.3)
-light.axis_ratio = af.UniformPrior(lower_limit=0.7, upper_limit=1.0)
-light.angle = af.UniformPrior(lower_limit=0.0, upper_limit=180.0)
-light.intensity = af.LogUniformPrior(lower_limit=1e-4, upper_limit=1e4)
-light.effective_radius = af.UniformPrior(lower_limit=0.0, upper_limit=1.0)
-
-source = af.Model(cosmo.Galaxy, redshift=1.0, light_profile_list=[light])
-
-print(source.info)
-
-"""
-We can now create our overall strong lens model, using a `Collection` in the same way we have seen previously.
-"""
-model = af.Collection(galaxies=af.Collection(lens=lens, source=source))
-
-print(model.info)
-
-"""
-The model contains both galaxies in the strong lens, alongside all of their light and mass profiles.
-
-For every iteration of the non-linear search **PyAutoFit** generates an instance of this model, where all of the
-`LightDeVaucouleurs`, `MassIsothermal` and `Galaxy` parameters of the are determined via their priors.
-
-An example instance is show below:
-"""
-instance = model.instance_from_prior_medians()
-
-print("Strong Lens Model Instance:")
-print("Lens Galaxy = ", instance.galaxies.lens)
-print("Lens Galaxy Light = ", instance.galaxies.lens.profile_list)
-print("Lens Galaxy Light Centre = ", instance.galaxies.lens.profile_list[0].centre)
-print("Lens Galaxy Mass Centre = ", instance.galaxies.lens.mass_profile_list[0].centre)
-print("Source Galaxy = ", instance.galaxies.source)
-
-"""
-We have successfully composed a multi-level model, which we can fit via a non-linear search.
-
-At this point, you should check out the `Analysis` class of this example project, in the
-module `projects/cosmology/src/analysis.py`. This class serves the same purpose that we have seen in the Gaussian 1D
-examples, with the `log_likelihood_function` implementing the calculation we showed in the first tutorial.
-
-The `path_prefix1 and `name` inputs below sepciify the path and folder where the results of the model-fit are stored
-in the output folder `autolens_workspace/output`. Results for this tutorial are writtent to hard-disk, due to the
-longer run-times of the model-fit.
-"""
-
-search = af.DynestyStatic(
- path_prefix=path.join("projects", "cosmology"),
- name="multi_level",
- nlive=50,
- iterations_per_full_update=2500,
-)
-
-analysis = cosmo.Analysis(data=data, noise_map=noise_map, psf=psf, grid=grid)
-
-"""
-If you comment out the code below, you will perform a lens model fit using the model and analysis class for
-this project. However, this model-fit is slow to run, and it isn't paramount that you run it yourself.
-
-The animation below shows a slide-show of the lens modeling procedure. Many lens models are fitted to the data over
-and over, gradually improving the quality of the fit to the data and looking more and more like the observed image.
-
-
-"""
-
-result = search.fit(model=model, analysis=analysis)
-
-"""
-__Extensibility__
-
-This example project highlights how multi-level models can make certain model-fitting problem fully extensible. For
-example:
-
- 1) A `Galaxy` class can be created using any combination of light and mass profiles, because it can wrap their
- `image_from_grid` and `deflections_from_grid` methods as the sum of the individual profiles.
-
- 2) The overall strong lens model can contain any number of `Galaxy`'s, as their methods are used
- to implement the lensing calculations in the `Analysis` class and `log_likelihood_function`.
-
-For problems of this nature, we can design and write code in a way that fully utilizes **PyAutoFit**'s multi-level
-modeling features to compose and fits models of arbitrary complexity and dimensionality.
-
-__Galaxy Clusters__
-
-To illustrate this further, consider the following dataset which is called a "strong lens galaxy cluster":
-
-
-
-For this strong lens, there are many tens of strong lens galaxies as well as multiple background source galaxies.
-
-However, despite it being a significantly more complex system than the single-galaxy strong lens we modeled above,
-our use of multi-level models ensures that we can model such datasets without any additional code development, for
-example:
-
-The lensing calculations in the source code `Analysis` object did not properly account for multiple galaxies
-(called multi-plane ray tracing). This would need to be updated to properly model a galaxy cluster, but this
-tutorial shows how a model can be composed for such a system.
-"""
-lens_0 = af.Model(
- cosmo.Galaxy,
- redshift=0.5,
- light_profile_list=[cosmo.lp.LightDeVaucouleurs],
- mass_profile_list=[cosmo.mp.MassIsothermal],
-)
-
-lens_1 = af.Model(
- cosmo.Galaxy,
- redshift=0.5,
- light_profile_list=[cosmo.lp.LightDeVaucouleurs],
- mass_profile_list=[cosmo.mp.MassIsothermal],
-)
-
-source_0 = af.Model(
- cosmo.Galaxy, redshift=1.0, light_profile_list=[af.Model(cosmo.lp.LightExponential)]
-)
-
-# ... repeat for desired model complexity ...
-
-model = af.Collection(
- galaxies=af.Collection(
- lens_0=lens_0,
- lens_1=lens_1,
- source_0=source_0,
- # ... repeat for desired model complexity ...
- )
-)
-
-print(model.info)
-
-"""
-Here is a pictoral representation of a strong lens cluster as a multi-level model:
-
-
-
-__Wrap Up__
-
-Strong gravitational lensing is a great example of a problem that can be approached using multi-level models.
-
-At the core of this is how there are many different models one could imagine defining which describe the light or mass
-of a galaxy. However, all of these models must derive the same fundamental property in order to fit the data, for
-example the image of a light profile or the deflection angles of the mass profile.
-
-The multi-level nature of strong lensing is not unique, and is commonly found in my Astronomy problems and the
-scientific literature in general. For example Astronomy problems:
-
- - Studies of galaxy structure, which represent the surface brightness distributions of galaxies as sums of Sersic
- profiles (or other parametric equations) to quantify whether they are bulge-like or disk-like.
-
- - Studies of galaxy dynamics, which represent the mass distribution of galaxies as sums of profiles like the Isothermal
- profile.
-
- - Studies of the activate galactic nuclei (AGN) of galaxies, where the different components of the AGN are represented
- as different model components.
-"""
+"""
+Project: Cosmology
+==================
+
+This project uses the astrophysical phenomena of Strong Gravitational Lensing to illustrate basic and advanced model
+composition and fitting with **PyAutoFit**. The first tutorial described what a strong gravitational lens is and how
+we build and fit a model of one.
+
+In this example, we use **PyAutoFit**'s multi-level models to compose a strong lens model consisting of a lens and
+source galaxy, and fit it to the data on SDSSJ2303+1422.
+
+__Config Path__
+
+We first set up the path to this projects config files, which is located at `autofit_workspace/projects/cosmology/config`.
+
+This includes the default priors for the lens model, check it out!
+"""
+
+import os
+from os import path
+from autoconf import conf
+
+cwd = os.getcwd()
+config_path = path.join(cwd, "projects", "cosmology", "config")
+conf.instance.push(new_path=config_path)
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import autofit as af
+import src as cosmo
+import matplotlib.pyplot as plt
+import numpy as np
+
+"""
+__Plot__
+
+First, lets again define the plotting convenience functions we used in the previous example.
+"""
+
+
+def plot_array(array, title=None, norm=None):
+ plt.imshow(array, norm=norm)
+ plt.colorbar()
+ plt.title(title)
+ plt.show()
+ plt.close()
+
+
+def plot_grid(grid, title=None):
+ plt.scatter(x=grid[:, :, 0], y=grid[:, :, 1], s=1)
+ plt.title(title)
+ plt.show()
+ plt.close()
+
+
+"""
+__Data__
+
+Now lets load and plot Hubble Space Telescope imaging data of the strong gravitational lens SDSSJ2303+1422.
+"""
+dataset_path = path.join("projects", "cosmology", "dataset")
+
+data = np.load(file=path.join(dataset_path, "data.npy"))
+plot_array(array=data, title="Image of Strong Lens SDSSJ2303+1422")
+
+noise_map = np.load(file=path.join(dataset_path, "noise_map.npy"))
+plot_array(array=noise_map, title="Noise Map of Strong Lens SDSSJ2303+1422")
+
+psf = np.load(file=path.join(dataset_path, "psf.npy"))
+plot_array(array=psf, title="Point Spread Function of Strong Lens SDSSJ2303+1422")
+
+grid = np.load(file=path.join(dataset_path, "grid.npy"))
+
+plot_grid(
+ grid=grid,
+ title="Cartesian grid of (x,y) coordinates aligned with strong lens dataset",
+)
+
+"""
+__Multi-level Model__
+
+In the previous example, we saw that we can use instances of the light profiles, mass profiles and galaxy objects to
+perform strong lens ray-tracing calculations:
+"""
+light_profile = cosmo.lp.LightDeVaucouleurs(
+ centre=(0.01, 0.01), axis_ratio=0.7, angle=45.0, intensity=1.0, effective_radius=2.0
+)
+mass_profile = cosmo.mp.MassIsothermal(
+ centre=(0.01, 0.01), axis_ratio=0.7, angle=45.0, mass=0.5
+)
+galaxy = cosmo.Galaxy(
+ redshift=0.5, light_profile_list=[light_profile], mass_profile_list=[mass_profile]
+)
+
+"""
+In this example, we want to perform a model-fit using a non-linear search, where the `Galaxy` is a `Model`, but it
+contains model subcomponents that are its individual light and mass profiles.
+
+Here is a pictoral representation of the model:
+
+
+
+__Model Composition__
+
+How do we compose a strong lens model where a `Galaxy` is a `Model`, but it contains the light and mass profiles
+as `Model` themselves?
+
+We use **PyAutoFit**'s multi-level model composition:
+"""
+lens = af.Model(
+ cls=cosmo.Galaxy, # The overall model object uses this input.
+ redshift=0.5,
+ light_profile_list=[
+ af.Model(cosmo.lp.LightDeVaucouleurs)
+ ], # These will be subcomponents of the model.
+ mass_profile_list=[
+ af.Model(cosmo.mp.MassIsothermal)
+ ], # These will be subcomponents of the model.
+)
+
+print(lens.info)
+
+"""
+Lets consider what is going on here:
+
+ 1) We use a `Model` to create the overall model component. The `cls` input is the `Galaxy` class, therefore the
+ overall model that is created is a `Galaxy`.
+
+ 2) **PyAutoFit** next inspects whether the key word argument inputs to the `Model` match any of the `__init__`
+ constructor arguments of the `Galaxy` class. This determine if these inputs are to be composed as model
+ subcomponents of the overall `Galaxy` model.
+
+ 3) **PyAutoFit** matches the `light_profile_list` and `mass_profile_list` inputs, noting they are passed as separate
+ lists containing the `LightDeVaucouleurs` and `MassIsothermal` class. They are both created as subcomponents of
+ the overall `Galaxy` model.
+
+ 4) It also matches the `redshift` input, making it a fixed value of 0.5 for the model and not treating it as a
+ free parameter.
+
+We can confirm this by printing the `total_free_parameters` of the lens, and noting it is 11 (6 parameters for
+the `LightDeVaucouleurs` and 5 for the `MassIsothermal`).
+"""
+print(lens.total_free_parameters)
+print(lens.light_profile_list[0].total_free_parameters)
+print(lens.mass_profile_list[0].total_free_parameters)
+
+"""
+The `lens` behaves exactly like the model-components we are used to previously. For example, we can unpack its
+individual parameters to customize the model, where below we:
+
+ 1) Fix the light and mass profiles to the centre (0.0, 0.0).
+ 2) Customize the prior on the light profile `axis_ratio`.
+ 3) Fix the `axis_ratio` of the mass profile to 0.8.
+"""
+
+lens.light_profile_list[0].centre = (0.0, 0.0)
+lens.light_profile_list[0].axis_ratio = af.UniformPrior(
+ lower_limit=0.7, upper_limit=0.9
+)
+lens.light_profile_list[0].angle = af.UniformPrior(lower_limit=0.0, upper_limit=180.0)
+lens.light_profile_list[0].intensity = af.LogUniformPrior(
+ lower_limit=1e-4, upper_limit=1e4
+)
+lens.light_profile_list[0].effective_radius = af.UniformPrior(
+ lower_limit=0.0, upper_limit=5.0
+)
+
+lens.mass_profile_list[0].centre = (0.0, 0.0)
+lens.mass_profile_list[0].axis_ratio = 0.8
+lens.mass_profile_list[0].angle = af.UniformPrior(lower_limit=0.0, upper_limit=180.0)
+lens.mass_profile_list[0].mass = af.UniformPrior(lower_limit=0.0, upper_limit=2.0)
+
+print(lens.info)
+
+"""
+__Alternative API__
+
+We can create the `Galaxy` model component with the exact same customization by creating each profile as a `Model` and
+passing these to the galaxy `Model`.
+"""
+light = af.Model(cosmo.lp.LightDeVaucouleurs)
+
+light.centre = af.UniformPrior(lower_limit=-0.05, upper_limit=0.05)
+light.axis_ratio = af.UniformPrior(lower_limit=0.7, upper_limit=0.9)
+light.angle = af.UniformPrior(lower_limit=0.0, upper_limit=180.0)
+light.intensity = af.LogUniformPrior(lower_limit=1e-4, upper_limit=1e4)
+light.effective_radius = af.UniformPrior(lower_limit=0.0, upper_limit=5.0)
+
+
+mass = af.Model(cosmo.mp.MassIsothermal)
+
+mass.centre = (0.0, 0.0)
+mass.axis_ratio = af.UniformPrior(lower_limit=0.7, upper_limit=1.0)
+mass.angle = af.UniformPrior(lower_limit=0.0, upper_limit=180.0)
+mass.mass = af.UniformPrior(lower_limit=0.0, upper_limit=4.0)
+
+lens = af.Model(
+ cosmo.Galaxy, redshift=0.5, light_profile_list=[light], mass_profile_list=[mass]
+)
+
+print(lens.info)
+
+"""
+We can now create a model of our source galaxy using the same API.
+"""
+light = af.Model(cosmo.lp.LightExponential)
+
+light.centre.centre_0 = af.GaussianPrior(mean=0.0, sigma=0.3)
+light.centre.centre_1 = af.GaussianPrior(mean=0.0, sigma=0.3)
+light.axis_ratio = af.UniformPrior(lower_limit=0.7, upper_limit=1.0)
+light.angle = af.UniformPrior(lower_limit=0.0, upper_limit=180.0)
+light.intensity = af.LogUniformPrior(lower_limit=1e-4, upper_limit=1e4)
+light.effective_radius = af.UniformPrior(lower_limit=0.0, upper_limit=1.0)
+
+source = af.Model(cosmo.Galaxy, redshift=1.0, light_profile_list=[light])
+
+print(source.info)
+
+"""
+We can now create our overall strong lens model, using a `Collection` in the same way we have seen previously.
+"""
+model = af.Collection(galaxies=af.Collection(lens=lens, source=source))
+
+print(model.info)
+
+"""
+The model contains both galaxies in the strong lens, alongside all of their light and mass profiles.
+
+For every iteration of the non-linear search **PyAutoFit** generates an instance of this model, where all of the
+`LightDeVaucouleurs`, `MassIsothermal` and `Galaxy` parameters of the are determined via their priors.
+
+An example instance is show below:
+"""
+instance = model.instance_from_prior_medians()
+
+print("Strong Lens Model Instance:")
+print("Lens Galaxy = ", instance.galaxies.lens)
+print("Lens Galaxy Light = ", instance.galaxies.lens.profile_list)
+print("Lens Galaxy Light Centre = ", instance.galaxies.lens.profile_list[0].centre)
+print("Lens Galaxy Mass Centre = ", instance.galaxies.lens.mass_profile_list[0].centre)
+print("Source Galaxy = ", instance.galaxies.source)
+
+"""
+We have successfully composed a multi-level model, which we can fit via a non-linear search.
+
+At this point, you should check out the `Analysis` class of this example project, in the
+module `projects/cosmology/src/analysis.py`. This class serves the same purpose that we have seen in the Gaussian 1D
+examples, with the `log_likelihood_function` implementing the calculation we showed in the first tutorial.
+
+The `path_prefix1 and `name` inputs below sepciify the path and folder where the results of the model-fit are stored
+in the output folder `autolens_workspace/output`. Results for this tutorial are writtent to hard-disk, due to the
+longer run-times of the model-fit.
+"""
+
+search = af.DynestyStatic(
+ path_prefix=path.join("projects", "cosmology"),
+ name="multi_level",
+ nlive=50,
+ iterations_per_full_update=2500,
+)
+
+analysis = cosmo.Analysis(data=data, noise_map=noise_map, psf=psf, grid=grid)
+
+"""
+If you comment out the code below, you will perform a lens model fit using the model and analysis class for
+this project. However, this model-fit is slow to run, and it isn't paramount that you run it yourself.
+
+The animation below shows a slide-show of the lens modeling procedure. Many lens models are fitted to the data over
+and over, gradually improving the quality of the fit to the data and looking more and more like the observed image.
+
+
+"""
+
+result = search.fit(model=model, analysis=analysis)
+
+"""
+__Extensibility__
+
+This example project highlights how multi-level models can make certain model-fitting problem fully extensible. For
+example:
+
+ 1) A `Galaxy` class can be created using any combination of light and mass profiles, because it can wrap their
+ `image_from_grid` and `deflections_from_grid` methods as the sum of the individual profiles.
+
+ 2) The overall strong lens model can contain any number of `Galaxy`'s, as their methods are used
+ to implement the lensing calculations in the `Analysis` class and `log_likelihood_function`.
+
+For problems of this nature, we can design and write code in a way that fully utilizes **PyAutoFit**'s multi-level
+modeling features to compose and fits models of arbitrary complexity and dimensionality.
+
+__Galaxy Clusters__
+
+To illustrate this further, consider the following dataset which is called a "strong lens galaxy cluster":
+
+
+
+For this strong lens, there are many tens of strong lens galaxies as well as multiple background source galaxies.
+
+However, despite it being a significantly more complex system than the single-galaxy strong lens we modeled above,
+our use of multi-level models ensures that we can model such datasets without any additional code development, for
+example:
+
+The lensing calculations in the source code `Analysis` object did not properly account for multiple galaxies
+(called multi-plane ray tracing). This would need to be updated to properly model a galaxy cluster, but this
+tutorial shows how a model can be composed for such a system.
+"""
+lens_0 = af.Model(
+ cosmo.Galaxy,
+ redshift=0.5,
+ light_profile_list=[cosmo.lp.LightDeVaucouleurs],
+ mass_profile_list=[cosmo.mp.MassIsothermal],
+)
+
+lens_1 = af.Model(
+ cosmo.Galaxy,
+ redshift=0.5,
+ light_profile_list=[cosmo.lp.LightDeVaucouleurs],
+ mass_profile_list=[cosmo.mp.MassIsothermal],
+)
+
+source_0 = af.Model(
+ cosmo.Galaxy, redshift=1.0, light_profile_list=[af.Model(cosmo.lp.LightExponential)]
+)
+
+# ... repeat for desired model complexity ...
+
+model = af.Collection(
+ galaxies=af.Collection(
+ lens_0=lens_0,
+ lens_1=lens_1,
+ source_0=source_0,
+ # ... repeat for desired model complexity ...
+ )
+)
+
+print(model.info)
+
+"""
+Here is a pictoral representation of a strong lens cluster as a multi-level model:
+
+
+
+__Wrap Up__
+
+Strong gravitational lensing is a great example of a problem that can be approached using multi-level models.
+
+At the core of this is how there are many different models one could imagine defining which describe the light or mass
+of a galaxy. However, all of these models must derive the same fundamental property in order to fit the data, for
+example the image of a light profile or the deflection angles of the mass profile.
+
+The multi-level nature of strong lensing is not unique, and is commonly found in my Astronomy problems and the
+scientific literature in general. For example Astronomy problems:
+
+ - Studies of galaxy structure, which represent the surface brightness distributions of galaxies as sums of Sersic
+ profiles (or other parametric equations) to quantify whether they are bulge-like or disk-like.
+
+ - Studies of galaxy dynamics, which represent the mass distribution of galaxies as sums of profiles like the Isothermal
+ profile.
+
+ - Studies of the activate galactic nuclei (AGN) of galaxies, where the different components of the AGN are represented
+ as different model components.
+"""
diff --git a/run_scripts.sh b/run_scripts.sh
index 6ac5076d..5deb5c50 100644
--- a/run_scripts.sh
+++ b/run_scripts.sh
@@ -13,6 +13,9 @@ NO_RUN_YAML="$SCRIPT_DIR/../PyAutoBuild/autobuild/config/no_run.yaml"
PROJECT_KEY="autofit"
export PYAUTOFIT_TEST_MODE=1
+export PYAUTO_WORKSPACE_SMALL_DATASETS=1
+export PYAUTO_DISABLE_CRITICAL_CAUSTICS=1
+export PYAUTO_FAST_PLOTS=1
# ---------------------------------------------------------------------------
# Build ordered script list: within each directory, start_here.py runs first,
diff --git a/scripts/cookbooks/analysis.py b/scripts/cookbooks/analysis.py
index 184da5d2..91e1ff6a 100644
--- a/scripts/cookbooks/analysis.py
+++ b/scripts/cookbooks/analysis.py
@@ -1,721 +1,717 @@
-"""
-Cookbook: Analysis
-==================
-
-The `Analysis` class is the interface between the data and model, whereby a `log_likelihood_function` is defined
-and called by the non-linear search to fit the model.
-
-This cookbook provides an overview of how to use and extend `Analysis` objects in **PyAutoFit**.
-
-__Contents__
-
- - Example: A simple example of an analysis class which can be adapted for you use-case.
- - Customization: Customizing an analysis class with different data inputs and editing the `log_likelihood_function`.
- - Visualization: Using a `visualize` method so that model-specific visuals are output to hard-disk.
- - Custom Result: Return a custom Result object with methods specific to your model fitting problem.
- - Latent Variables: Adding a `compute_latent_variables` method to the analysis to output latent variables to hard-disk.
- - Custom Output: Add methods which output model-specific results to hard-disk in the `files` folder (e.g. as .json
- files) to aid in the interpretation of results.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import json
-import numpy as np
-from os import path
-from typing import Optional, Tuple
-
-import autofit as af
-
-"""
-__Example__
-
-An example simple `Analysis` class, to remind ourselves of the basic structure and inputs.
-
-This can be adapted for your use case.
-"""
-
-
-class Analysis(af.Analysis):
- def __init__(self, data: np.ndarray, noise_map: np.ndarray):
- """
- The `Analysis` class acts as an interface between the data and model in **PyAutoFit**.
-
- Its `log_likelihood_function` defines how the model is fitted to the data and it is called many times by
- the non-linear search fitting algorithm.
-
- In this example the `Analysis` `__init__` constructor only contains the `data` and `noise-map`, but it can be
- easily extended to include other quantities.
-
- Parameters
- ----------
- data
- A 1D numpy array containing the data (e.g. a noisy 1D signal) fitted in the workspace examples.
- noise_map
- A 1D numpy array containing the noise values of the data, used for computing the goodness of fit
- metric, the log likelihood.
- """
- super().__init__()
-
- self.data = data
- self.noise_map = noise_map
-
- def log_likelihood_function(self, instance) -> float:
- """
- Returns the log likelihood of a fit of a 1D Gaussian to the dataset.
-
- The data is fitted using an `instance` of the `Gaussian` class where its `model_data_from`
- is called in order to create a model data representation of the Gaussian that is fitted to the data.
- """
-
- xvalues = np.arange(self.data.shape[0])
-
- model_data = instance.model_data_from(xvalues=xvalues)
-
- residual_map = self.data - model_data
- chi_squared_map = (residual_map / self.noise_map) ** 2.0
- chi_squared = sum(chi_squared_map)
- noise_normalization = np.sum(np.log(2 * np.pi * self.noise_map**2.0))
- log_likelihood = -0.5 * (chi_squared + noise_normalization)
-
- return log_likelihood
-
-
-"""
-An instance of the analysis class is created as follows.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-analysis = Analysis(data=data, noise_map=noise_map)
-
-"""
-__Customization__
-
-The `Analysis` class can be fully customized to be suitable for your model-fit.
-
-For example, additional inputs can be included in the `__init__` constructor and used in the `log_likelihood_function`.
-if they are required for your `log_likelihood_function` to work.
-
-The example below includes three additional inputs:
-
- - Instead of inputting a `noise_map`, a `noise_covariance_matrix` is input, which means that corrrlated noise is
- accounted for in the `log_likelihood_function`.
-
- - A `mask` is input which masks the data such that certain data points are omitted from the log likelihood
-
- - A `kernel` is input which can account for certain blurring operations during data acquisition.
-"""
-
-
-class Analysis(af.Analysis):
- def __init__(
- self,
- data: np.ndarray,
- noise_covariance_matrix: np.ndarray,
- mask: np.ndarray,
- kernel: np.ndarray,
- ):
- """
- The `Analysis` class which has had its inputs edited for a different model-fit.
-
- Parameters
- ----------
- data
- A 1D numpy array containing the data (e.g. a noisy 1D signal) fitted in the workspace examples.
- noise_covariance_matrix
- A 2D numpy array containing the noise values and their covariances for the data, used for computing the
- goodness of fit whilst accounting for correlated noise.
- mask
- A 1D numpy array containing a mask, where `True` values mean a data point is masked and is omitted from
- the log likelihood.
- kernel
- A 1D numpy array containing the blurring kernel of the data, used for creating the model data.
- """
- super().__init__()
-
- self.data = data
- self.noise_covariance_matrix = noise_covariance_matrix
- self.mask = mask
- self.kernel = kernel
-
- def log_likelihood_function(self, instance) -> float:
- """
- The `log_likelihood_function` now has access to the `noise_covariance_matrix`, `mask` and `kernel`
- input above.
- """
- print(self.noise_covariance_matrix)
- print(self.mask)
- print(self.kernel)
-
- """
- We do not provide a specific example of how to use these inputs in the `log_likelihood_function` as they are
- specific to your model fitting problem.
-
- The key point is that any inputs required to compute the log likelihood can be passed into the `__init__`
- constructor of the `Analysis` class and used in the `log_likelihood_function`.
- """
-
- log_likelihood = None
-
- return log_likelihood
-
-
-"""
-An instance of the analysis class is created as follows.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-
-noise_covariance_matrix = np.ones(shape=(data.shape[0], data.shape[0]))
-mask = np.full(fill_value=False, shape=data.shape)
-kernel = np.full(fill_value=1.0, shape=data.shape)
-
-analysis = Analysis(
- data=data, noise_covariance_matrix=noise_covariance_matrix, mask=mask, kernel=kernel
-)
-
-"""
-__Visualization__
-
-If a `name` is input into a non-linear search, all results are output to hard-disk in a folder.
-
-By overwriting the `Visualizer` object of an `Analysis` class with a custom `Visualizer` class, custom results of the
-model-fit can be visualized during the model-fit.
-
-The `Visualizer` below has the methods `visualize_before_fit` and `visualize`, which perform model specific
-visualization will also be output into an `image` folder, for example as `.png` files.
-
-This uses the maximum log likelihood model of the model-fit inferred so far.
-
-Visualization of the results of the search, such as the corner plot of what is called the "Probability Density
-Function", are also automatically output during the model-fit on the fly.
-"""
-
-
-class Visualizer(af.Visualizer):
- @staticmethod
- def visualize_before_fit(
- analysis, paths: af.DirectoryPaths, model: af.AbstractPriorModel
- ):
- """
- Before a model-fit, the `visualize_before_fit` method is called to perform visualization.
-
- The function receives as input an instance of the `Analysis` class which is being used to perform the fit,
- which is used to perform the visualization (e.g. it contains the data and noise map which are plotted).
-
- This can output visualization of quantities which do not change during the model-fit, for example the
- data and noise-map.
-
- The `paths` object contains the path to the folder where the visualization should be output, which is determined
- by the non-linear search `name` and other inputs.
- """
-
- import matplotlib.pyplot as plt
-
- xvalues = np.arange(analysis.data.shape[0])
-
- plt.errorbar(
- x=xvalues,
- y=analysis.data,
- yerr=analysis.noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
- )
- plt.title("Maximum Likelihood Fit")
- plt.xlabel("x value of profile")
- plt.ylabel("Profile Normalization")
- plt.savefig(path.join(paths.image_path, f"data.png"))
- plt.clf()
-
- @staticmethod
- def visualize(analysis, paths: af.DirectoryPaths, instance, during_analysis):
- """
- During a model-fit, the `visualize` method is called throughout the non-linear search.
-
- The function receives as input an instance of the `Analysis` class which is being used to perform the fit,
- which is used to perform the visualization (e.g. it generates the model data which is plotted).
-
- The `instance` passed into the visualize method is maximum log likelihood solution obtained by the model-fit
- so far and it can be used to provide on-the-fly images showing how the model-fit is going.
-
- The `paths` object contains the path to the folder where the visualization should be output, which is determined
- by the non-linear search `name` and other inputs.
- """
- xvalues = np.arange(analysis.data.shape[0])
-
- model_data = instance.model_data_from(xvalues=xvalues)
- residual_map = analysis.data - model_data
-
- """
- The visualizer now outputs images of the best-fit results to hard-disk (checkout `visualizer.py`).
- """
- import matplotlib.pyplot as plt
-
- plt.errorbar(
- x=xvalues,
- y=analysis.data,
- yerr=analysis.noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
- )
- plt.plot(xvalues, model_data, color="r")
- plt.title("Maximum Likelihood Fit")
- plt.xlabel("x value of profile")
- plt.ylabel("Profile Normalization")
- plt.savefig(path.join(paths.image_path, f"model_fit.png"))
- plt.clf()
-
- plt.errorbar(
- x=xvalues,
- y=residual_map,
- yerr=analysis.noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
- )
- plt.title("Residuals of Maximum Likelihood Fit")
- plt.xlabel("x value of profile")
- plt.ylabel("Residual")
- plt.savefig(path.join(paths.image_path, f"model_fit.png"))
- plt.clf()
-
-
-"""
-The `Analysis` class is defined following the same API as before, but now with its `Visualizer` class attribute
-overwritten with the `Visualizer` class above.
-"""
-
-
-class Analysis(af.Analysis):
- """
- This over-write means the `Visualizer` class is used for visualization throughout the model-fit.
-
- This `VisualizerExample` object is in the `autofit.example.visualize` module and is used to customize the
- plots output during the model-fit.
-
- It has been extended with visualize methods that output visuals specific to the fitting of `1D` data.
- """
-
- Visualizer = Visualizer
-
- def __init__(self, data, noise_map):
- """
- An Analysis class which illustrates visualization.
- """
- super().__init__()
-
- self.data = data
- self.noise_map = noise_map
-
- def log_likelihood_function(self, instance):
- """
- The `log_likelihood_function` is identical to the example above
- """
- xvalues = np.arange(self.data.shape[0])
-
- model_data = instance.model_data_from(xvalues=xvalues)
- residual_map = self.data - model_data
- chi_squared_map = (residual_map / self.noise_map) ** 2.0
- chi_squared = sum(chi_squared_map)
- noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
- log_likelihood = -0.5 * (chi_squared + noise_normalization)
-
- return log_likelihood
-
-
-"""
-__Custom Result__
-
-The `Result` object is returned by a non-linear search after running the following code:
-
-`result = search.fit(model=model, analysis=analysis)`
-
-The result can be can be customized to include additional information about the model-fit that is specific to your
-model-fitting problem.
-
-For example, for fitting 1D profiles, the `Result` could include the maximum log likelihood model 1D data:
-
-`print(result.max_log_likelihood_model_data_1d)`
-
-In other examples, this quantity has been manually computed after the model-fit has completed.
-
-The custom result API allows us to do this. First, we define a custom `Result` class, which includes the property
-`max_log_likelihood_model_data_1d`.
-"""
-
-
-class ResultExample(af.Result):
- @property
- def max_log_likelihood_model_data_1d(self) -> np.ndarray:
- """
- Returns the maximum log likelihood model's 1D model data.
-
- This is an example of how we can pass the `Analysis` class a custom `Result` object and extend this result
- object with new properties that are specific to the model-fit we are performing.
- """
- xvalues = np.arange(self.analysis.data.shape[0])
-
- return self.instance.model_data_from(xvalues=xvalues)
-
-
-"""
-The custom result has access to the analysis class, meaning that we can use any of its methods or properties to
-compute custom result properties.
-
-To make it so that the `ResultExample` object above is returned by the search we overwrite the `Result` class attribute
-of the `Analysis` and define a `make_result` object describing what we want it to contain:
-"""
-
-
-class Analysis(af.Analysis):
- """
- This overwrite means the `ResultExample` class is returned after the model-fit.
- """
-
- Result = ResultExample
-
- def __init__(self, data, noise_map):
- """
- An Analysis class which illustrates custom results.
- """
- super().__init__()
-
- self.data = data
- self.noise_map = noise_map
-
- def log_likelihood_function(self, instance):
- """
- The `log_likelihood_function` is identical to the example above
- """
- xvalues = np.arange(self.data.shape[0])
-
- model_data = instance.model_data_from(xvalues=xvalues)
- residual_map = self.data - model_data
- chi_squared_map = (residual_map / self.noise_map) ** 2.0
- chi_squared = sum(chi_squared_map)
- noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
- log_likelihood = -0.5 * (chi_squared + noise_normalization)
-
- return log_likelihood
-
- def make_result(
- self,
- samples_summary: af.SamplesSummary,
- paths: af.AbstractPaths,
- samples: Optional[af.SamplesPDF] = None,
- search_internal: Optional[object] = None,
- analysis: Optional[object] = None,
- ) -> Result:
- """
- Returns the `Result` of the non-linear search after it is completed.
-
- The result type is defined as a class variable in the `Analysis` class (see top of code under the python code
- `class Analysis(af.Analysis)`.
-
- The result can be manually overwritten by a user to return a user-defined result object, which can be extended
- with additional methods and attribute specific to the model-fit.
-
- This example class does example this, whereby the analysis result has been overwritten with the `ResultExample`
- class, which contains a property `max_log_likelihood_model_data_1d` that returns the model data of the
- best-fit model. This API means you can customize your result object to include whatever attributes you want
- and therefore make a result object specific to your model-fit and model-fitting problem.
-
- The `Result` object you return can be customized to include:
-
- - The samples summary, which contains the maximum log likelihood instance and median PDF model.
-
- - The paths of the search, which are used for loading the samples and search internal below when a search
- is resumed.
-
- - The samples of the non-linear search (e.g. MCMC chains) also stored in `samples.csv`.
-
- - The non-linear search used for the fit in its internal representation, which is used for resuming a search
- and making bespoke visualization using the search's internal results.
-
- - The analysis used to fit the model (default disabled to save memory, but option may be useful for certain
- projects).
-
- Parameters
- ----------
- samples_summary
- The summary of the samples of the non-linear search, which include the maximum log likelihood instance and
- median PDF model.
- paths
- An object describing the paths for saving data (e.g. hard-disk directories or entries in sqlite database).
- samples
- The samples of the non-linear search, for example the chains of an MCMC run.
- search_internal
- The internal representation of the non-linear search used to perform the model-fit.
- analysis
- The analysis used to fit the model.
-
- Returns
- -------
- Result
- The result of the non-linear search, which is defined as a class variable in the `Analysis` class.
- """
- return self.Result(
- samples_summary=samples_summary,
- paths=paths,
- samples=samples,
- search_internal=search_internal,
- analysis=self,
- )
-
-
-"""
-For the sake of brevity, we do not run the code below, but the following code would work:
-
-`result = search.fit(model=model, analysis=analysis)`
-`print(result.max_log_likelihood_model_data_1d)`
-
-__Latent Variables__
-
-A latent variable is not a model parameter but can be derived from the model. Its value and errors may be of interest
-and aid in the interpretation of a model-fit.
-
-For example, for the simple 1D Gaussian example, it could be the full-width half maximum (FWHM) of the Gaussian.
-This is not included in the model but can be easily derived from the Gaussian's sigma value.
-
-By overwriting the Analysis class's `compute_latent_variables` method we can manually specify latent variables that
-are calculated. If the search has a `name`, these are output to a `latent.csv` file, which mirrors
-the `samples.csv` file.
-
-There may also be a `latent.results` and `latent_summary.json` files output. The `output.yaml` config file contains
-settings customizing what files are output and how often.
-
-This function takes as input the `parameters`, not the `instance`, because it means the function supports JAX.jit
-and thus if JAX is being used can be fully accelerated. The `instance` is created immediately inside the function.
-"""
-
-
-class Analysis(af.Analysis):
-
- LATENT_KEYS = ["gaussian.fwhm"]
-
- def __init__(self, data, noise_map):
- """
- An Analysis class which illustrates latent variables.
- """
- super().__init__()
-
- self.data = data
- self.noise_map = noise_map
-
- def log_likelihood_function(self, instance):
- """
- The `log_likelihood_function` is identical to the example above
- """
- xvalues = np.arange(self.data.shape[0])
-
- model_data = instance.model_data_from(xvalues=xvalues)
- residual_map = self.data - model_data
- chi_squared_map = (residual_map / self.noise_map) ** 2.0
- chi_squared = sum(chi_squared_map)
- noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
- log_likelihood = -0.5 * (chi_squared + noise_normalization)
-
- return log_likelihood
-
- def compute_latent_variables(self, parameters, model) -> Tuple:
- """
- A latent variable is not a model parameter but can be derived from the model. Its value and errors may be
- of interest and aid in the interpretation of a model-fit.
-
- For example, for the simple 1D Gaussian example, it could be the full-width half maximum (FWHM) of the
- Gaussian. This is not included in the model but can be easily derived from the Gaussian's sigma value.
-
- By overwriting this method we can manually specify latent variables that are calculated and output to
- a `latent.csv` file, which mirrors the `samples.csv` file.
-
- In the example below, the `latent.csv` file will contain one column with the FWHM of every Gausian model
- sampled by the non-linear search.
-
- This function is called at the end of search, following one of two schemes depending on the settings in
- `output.yaml`:
-
- 1) Call for every search sample, which produces a complete `latent/samples.csv` which mirrors the normal
- `samples.csv` file but takes a long time to compute.
-
- 2) Call only for N random draws from the posterior inferred at the end of the search, which only produces a
- `latent/latent_summary.json` file with the median and 1 and 3 sigma errors of the latent variables but is
- fast to compute.
-
- Parameters
- ----------
- instance
- The instances of the model which the latent variable is derived from.
-
- Returns
- -------
- A dictionary mapping every latent variable name to its value.
- """
-
- instance = model.instance_from_vector(vector=parameters)
-
- return (instance.fwhm,)
-
-
-"""
-Outputting latent variables manually after a fit is complete is simple, just call
-the `analysis.compute_latent_variables()` function.
-
-For many use cases, the best set up disables autofit latent variable output during a fit via the `output.yaml`
-file and perform it manually after completing a successful model-fit. This will save computational run time by not
-computing latent variables during a any model-fit which is unsuccessful.
-"""
-analysis = Analysis(data=data, noise_map=noise_map)
-
-# Commented out because we do not run the search in this cookbook
-
-# latent_samples = analysis.compute_latent_variables(samples=result.samples)
-
-"""
-Analysing and interpreting latent variables is described fully in the result cookbook.
-
-However, in brief, the `latent_samples` object is a `Samples` object and uses the same API as samples objects.
-"""
-# print(latent_samples.median_pdf().gaussian.fwhm)
-
-"""
-__Custom Output__
-
-When performing fits which output results to hard-disc, a `files` folder is created containing .json / .csv files of
-the model, samples, search, etc.
-
-These files are human readable and help one quickly inspect and interpret results.
-
-By extending an `Analysis` class with the methods `save_attributes` and `save_results`,
-custom files can be written to the `files` folder to further aid this inspection.
-
-These files can then also be loaded via the database, as described in the database cookbook.
-"""
-
-
-class Analysis(af.Analysis):
- def __init__(self, data: np.ndarray, noise_map: np.ndarray):
- """
- Standard Analysis class example used throughout PyAutoFit examples.
- """
- super().__init__()
-
- self.data = data
- self.noise_map = noise_map
-
- def log_likelihood_function(self, instance) -> float:
- """
- Standard log likelihood function used throughout PyAutoFit examples.
- """
-
- xvalues = np.arange(self.data.shape[0])
-
- model_data = instance.model_data_from(xvalues=xvalues)
-
- residual_map = self.data - model_data
- chi_squared_map = (residual_map / self.noise_map) ** 2.0
- chi_squared = sum(chi_squared_map)
- noise_normalization = np.sum(np.log(2 * np.pi * self.noise_map**2.0))
- log_likelihood = -0.5 * (chi_squared + noise_normalization)
-
- return log_likelihood
-
- def save_attributes(self, paths: af.DirectoryPaths):
- """
- Before the non-linear search begins, this routine saves attributes of the `Analysis` object to the `files`
- folder such that they can be loaded after the analysis using PyAutoFit's database and aggregator tools.
-
- For this analysis, it uses the `AnalysisDataset` object's method to output the following:
-
- - The dataset's data as a .json file.
- - The dataset's noise-map as a .json file.
-
- These are accessed using the aggregator via `agg.values("data")` and `agg.values("noise_map")`.
-
- Parameters
- ----------
- paths
- The paths object which manages all paths, e.g. where the non-linear search outputs are stored,
- visualization, and the pickled objects used by the aggregator output by this function.
- """
- # The path where data.json is saved, e.g. output/dataset_name/unique_id/files/data.json
-
- file_path = (path.join(paths._json_path, "data.json"),)
-
- with open(file_path, "w+") as f:
- json.dump(self.data, f, indent=4)
-
- # The path where noise_map.json is saved, e.g. output/noise_mapset_name/unique_id/files/noise_map.json
-
- file_path = (path.join(paths._json_path, "noise_map.json"),)
-
- with open(file_path, "w+") as f:
- json.dump(self.noise_map, f, indent=4)
-
- def save_results(self, paths: af.DirectoryPaths, result: af.Result):
- """
- At the end of a model-fit, this routine saves attributes of the `Analysis` object to the `files`
- folder such that they can be loaded after the analysis using PyAutoFit's database and aggregator tools.
-
- For this analysis it outputs the following:
-
- - The maximum log likelihood model data as a .json file.
-
- This is accessed using the aggregator via `agg.values("model_data")`.
-
- Parameters
- ----------
- paths
- The paths object which manages all paths, e.g. where the non-linear search outputs are stored,
- visualization and the pickled objects used by the aggregator output by this function.
- result
- The result of a model fit, including the non-linear search, samples and maximum likelihood model.
- """
- xvalues = np.arange(self.data.shape[0])
-
- instance = result.max_log_likelihood_instance
-
- model_data = instance.model_data_from(xvalues=xvalues)
-
- # The path where model_data.json is saved, e.g. output/dataset_name/unique_id/files/model_data.json
-
- file_path = (path.join(paths._json_path, "model_data.json"),)
-
- with open(file_path, "w+") as f:
- json.dump(model_data, f, indent=4)
-
-
-"""
-Finish.
-"""
+"""
+Cookbook: Analysis
+==================
+
+The `Analysis` class is the interface between the data and model, whereby a `log_likelihood_function` is defined
+and called by the non-linear search to fit the model.
+
+This cookbook provides an overview of how to use and extend `Analysis` objects in **PyAutoFit**.
+
+__Contents__
+
+ - Example: A simple example of an analysis class which can be adapted for you use-case.
+ - Customization: Customizing an analysis class with different data inputs and editing the `log_likelihood_function`.
+ - Visualization: Using a `visualize` method so that model-specific visuals are output to hard-disk.
+ - Custom Result: Return a custom Result object with methods specific to your model fitting problem.
+ - Latent Variables: Adding a `compute_latent_variables` method to the analysis to output latent variables to hard-disk.
+ - Custom Output: Add methods which output model-specific results to hard-disk in the `files` folder (e.g. as .json
+ files) to aid in the interpretation of results.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import json
+import numpy as np
+from os import path
+from typing import Optional, Tuple
+
+import autofit as af
+
+"""
+__Example__
+
+An example simple `Analysis` class, to remind ourselves of the basic structure and inputs.
+
+This can be adapted for your use case.
+"""
+
+
+class Analysis(af.Analysis):
+ def __init__(self, data: np.ndarray, noise_map: np.ndarray):
+ """
+ The `Analysis` class acts as an interface between the data and model in **PyAutoFit**.
+
+ Its `log_likelihood_function` defines how the model is fitted to the data and it is called many times by
+ the non-linear search fitting algorithm.
+
+ In this example the `Analysis` `__init__` constructor only contains the `data` and `noise-map`, but it can be
+ easily extended to include other quantities.
+
+ Parameters
+ ----------
+ data
+ A 1D numpy array containing the data (e.g. a noisy 1D signal) fitted in the workspace examples.
+ noise_map
+ A 1D numpy array containing the noise values of the data, used for computing the goodness of fit
+ metric, the log likelihood.
+ """
+ super().__init__()
+
+ self.data = data
+ self.noise_map = noise_map
+
+ def log_likelihood_function(self, instance) -> float:
+ """
+ Returns the log likelihood of a fit of a 1D Gaussian to the dataset.
+
+ The data is fitted using an `instance` of the `Gaussian` class where its `model_data_from`
+ is called in order to create a model data representation of the Gaussian that is fitted to the data.
+ """
+
+ xvalues = np.arange(self.data.shape[0])
+
+ model_data = instance.model_data_from(xvalues=xvalues)
+
+ residual_map = self.data - model_data
+ chi_squared_map = (residual_map / self.noise_map) ** 2.0
+ chi_squared = sum(chi_squared_map)
+ noise_normalization = np.sum(np.log(2 * np.pi * self.noise_map**2.0))
+ log_likelihood = -0.5 * (chi_squared + noise_normalization)
+
+ return log_likelihood
+
+
+"""
+An instance of the analysis class is created as follows.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+analysis = Analysis(data=data, noise_map=noise_map)
+
+"""
+__Customization__
+
+The `Analysis` class can be fully customized to be suitable for your model-fit.
+
+For example, additional inputs can be included in the `__init__` constructor and used in the `log_likelihood_function`.
+if they are required for your `log_likelihood_function` to work.
+
+The example below includes three additional inputs:
+
+ - Instead of inputting a `noise_map`, a `noise_covariance_matrix` is input, which means that corrrlated noise is
+ accounted for in the `log_likelihood_function`.
+
+ - A `mask` is input which masks the data such that certain data points are omitted from the log likelihood
+
+ - A `kernel` is input which can account for certain blurring operations during data acquisition.
+"""
+
+
+class Analysis(af.Analysis):
+ def __init__(
+ self,
+ data: np.ndarray,
+ noise_covariance_matrix: np.ndarray,
+ mask: np.ndarray,
+ kernel: np.ndarray,
+ ):
+ """
+ The `Analysis` class which has had its inputs edited for a different model-fit.
+
+ Parameters
+ ----------
+ data
+ A 1D numpy array containing the data (e.g. a noisy 1D signal) fitted in the workspace examples.
+ noise_covariance_matrix
+ A 2D numpy array containing the noise values and their covariances for the data, used for computing the
+ goodness of fit whilst accounting for correlated noise.
+ mask
+ A 1D numpy array containing a mask, where `True` values mean a data point is masked and is omitted from
+ the log likelihood.
+ kernel
+ A 1D numpy array containing the blurring kernel of the data, used for creating the model data.
+ """
+ super().__init__()
+
+ self.data = data
+ self.noise_covariance_matrix = noise_covariance_matrix
+ self.mask = mask
+ self.kernel = kernel
+
+ def log_likelihood_function(self, instance) -> float:
+ """
+ The `log_likelihood_function` now has access to the `noise_covariance_matrix`, `mask` and `kernel`
+ input above.
+ """
+ print(self.noise_covariance_matrix)
+ print(self.mask)
+ print(self.kernel)
+
+ """
+ We do not provide a specific example of how to use these inputs in the `log_likelihood_function` as they are
+ specific to your model fitting problem.
+
+ The key point is that any inputs required to compute the log likelihood can be passed into the `__init__`
+ constructor of the `Analysis` class and used in the `log_likelihood_function`.
+ """
+
+ log_likelihood = None
+
+ return log_likelihood
+
+
+"""
+An instance of the analysis class is created as follows.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+
+noise_covariance_matrix = np.ones(shape=(data.shape[0], data.shape[0]))
+mask = np.full(fill_value=False, shape=data.shape)
+kernel = np.full(fill_value=1.0, shape=data.shape)
+
+analysis = Analysis(
+ data=data, noise_covariance_matrix=noise_covariance_matrix, mask=mask, kernel=kernel
+)
+
+"""
+__Visualization__
+
+If a `name` is input into a non-linear search, all results are output to hard-disk in a folder.
+
+By overwriting the `Visualizer` object of an `Analysis` class with a custom `Visualizer` class, custom results of the
+model-fit can be visualized during the model-fit.
+
+The `Visualizer` below has the methods `visualize_before_fit` and `visualize`, which perform model specific
+visualization will also be output into an `image` folder, for example as `.png` files.
+
+This uses the maximum log likelihood model of the model-fit inferred so far.
+
+Visualization of the results of the search, such as the corner plot of what is called the "Probability Density
+Function", are also automatically output during the model-fit on the fly.
+"""
+
+
+class Visualizer(af.Visualizer):
+ @staticmethod
+ def visualize_before_fit(
+ analysis, paths: af.DirectoryPaths, model: af.AbstractPriorModel
+ ):
+ """
+ Before a model-fit, the `visualize_before_fit` method is called to perform visualization.
+
+ The function receives as input an instance of the `Analysis` class which is being used to perform the fit,
+ which is used to perform the visualization (e.g. it contains the data and noise map which are plotted).
+
+ This can output visualization of quantities which do not change during the model-fit, for example the
+ data and noise-map.
+
+ The `paths` object contains the path to the folder where the visualization should be output, which is determined
+ by the non-linear search `name` and other inputs.
+ """
+
+ import matplotlib.pyplot as plt
+
+ xvalues = np.arange(analysis.data.shape[0])
+
+ plt.errorbar(
+ x=xvalues,
+ y=analysis.data,
+ yerr=analysis.noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+ )
+ plt.title("Maximum Likelihood Fit")
+ plt.xlabel("x value of profile")
+ plt.ylabel("Profile Normalization")
+ plt.savefig(path.join(paths.image_path, f"data.png"))
+ plt.clf()
+
+ @staticmethod
+ def visualize(analysis, paths: af.DirectoryPaths, instance, during_analysis):
+ """
+ During a model-fit, the `visualize` method is called throughout the non-linear search.
+
+ The function receives as input an instance of the `Analysis` class which is being used to perform the fit,
+ which is used to perform the visualization (e.g. it generates the model data which is plotted).
+
+ The `instance` passed into the visualize method is maximum log likelihood solution obtained by the model-fit
+ so far and it can be used to provide on-the-fly images showing how the model-fit is going.
+
+ The `paths` object contains the path to the folder where the visualization should be output, which is determined
+ by the non-linear search `name` and other inputs.
+ """
+ xvalues = np.arange(analysis.data.shape[0])
+
+ model_data = instance.model_data_from(xvalues=xvalues)
+ residual_map = analysis.data - model_data
+
+ """
+ The visualizer now outputs images of the best-fit results to hard-disk (checkout `visualizer.py`).
+ """
+ import matplotlib.pyplot as plt
+
+ plt.errorbar(
+ x=xvalues,
+ y=analysis.data,
+ yerr=analysis.noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+ )
+ plt.plot(xvalues, model_data, color="r")
+ plt.title("Maximum Likelihood Fit")
+ plt.xlabel("x value of profile")
+ plt.ylabel("Profile Normalization")
+ plt.savefig(path.join(paths.image_path, f"model_fit.png"))
+ plt.clf()
+
+ plt.errorbar(
+ x=xvalues,
+ y=residual_map,
+ yerr=analysis.noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+ )
+ plt.title("Residuals of Maximum Likelihood Fit")
+ plt.xlabel("x value of profile")
+ plt.ylabel("Residual")
+ plt.savefig(path.join(paths.image_path, f"model_fit.png"))
+ plt.clf()
+
+
+"""
+The `Analysis` class is defined following the same API as before, but now with its `Visualizer` class attribute
+overwritten with the `Visualizer` class above.
+"""
+
+
+class Analysis(af.Analysis):
+ """
+ This over-write means the `Visualizer` class is used for visualization throughout the model-fit.
+
+ This `VisualizerExample` object is in the `autofit.example.visualize` module and is used to customize the
+ plots output during the model-fit.
+
+ It has been extended with visualize methods that output visuals specific to the fitting of `1D` data.
+ """
+
+ Visualizer = Visualizer
+
+ def __init__(self, data, noise_map):
+ """
+ An Analysis class which illustrates visualization.
+ """
+ super().__init__()
+
+ self.data = data
+ self.noise_map = noise_map
+
+ def log_likelihood_function(self, instance):
+ """
+ The `log_likelihood_function` is identical to the example above
+ """
+ xvalues = np.arange(self.data.shape[0])
+
+ model_data = instance.model_data_from(xvalues=xvalues)
+ residual_map = self.data - model_data
+ chi_squared_map = (residual_map / self.noise_map) ** 2.0
+ chi_squared = sum(chi_squared_map)
+ noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
+ log_likelihood = -0.5 * (chi_squared + noise_normalization)
+
+ return log_likelihood
+
+
+"""
+__Custom Result__
+
+The `Result` object is returned by a non-linear search after running the following code:
+
+`result = search.fit(model=model, analysis=analysis)`
+
+The result can be can be customized to include additional information about the model-fit that is specific to your
+model-fitting problem.
+
+For example, for fitting 1D profiles, the `Result` could include the maximum log likelihood model 1D data:
+
+`print(result.max_log_likelihood_model_data_1d)`
+
+In other examples, this quantity has been manually computed after the model-fit has completed.
+
+The custom result API allows us to do this. First, we define a custom `Result` class, which includes the property
+`max_log_likelihood_model_data_1d`.
+"""
+
+
+class ResultExample(af.Result):
+ @property
+ def max_log_likelihood_model_data_1d(self) -> np.ndarray:
+ """
+ Returns the maximum log likelihood model's 1D model data.
+
+ This is an example of how we can pass the `Analysis` class a custom `Result` object and extend this result
+ object with new properties that are specific to the model-fit we are performing.
+ """
+ xvalues = np.arange(self.analysis.data.shape[0])
+
+ return self.instance.model_data_from(xvalues=xvalues)
+
+
+"""
+The custom result has access to the analysis class, meaning that we can use any of its methods or properties to
+compute custom result properties.
+
+To make it so that the `ResultExample` object above is returned by the search we overwrite the `Result` class attribute
+of the `Analysis` and define a `make_result` object describing what we want it to contain:
+"""
+
+
+class Analysis(af.Analysis):
+ """
+ This overwrite means the `ResultExample` class is returned after the model-fit.
+ """
+
+ Result = ResultExample
+
+ def __init__(self, data, noise_map):
+ """
+ An Analysis class which illustrates custom results.
+ """
+ super().__init__()
+
+ self.data = data
+ self.noise_map = noise_map
+
+ def log_likelihood_function(self, instance):
+ """
+ The `log_likelihood_function` is identical to the example above
+ """
+ xvalues = np.arange(self.data.shape[0])
+
+ model_data = instance.model_data_from(xvalues=xvalues)
+ residual_map = self.data - model_data
+ chi_squared_map = (residual_map / self.noise_map) ** 2.0
+ chi_squared = sum(chi_squared_map)
+ noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
+ log_likelihood = -0.5 * (chi_squared + noise_normalization)
+
+ return log_likelihood
+
+ def make_result(
+ self,
+ samples_summary: af.SamplesSummary,
+ paths: af.AbstractPaths,
+ samples: Optional[af.SamplesPDF] = None,
+ search_internal: Optional[object] = None,
+ analysis: Optional[object] = None,
+ ) -> Result:
+ """
+ Returns the `Result` of the non-linear search after it is completed.
+
+ The result type is defined as a class variable in the `Analysis` class (see top of code under the python code
+ `class Analysis(af.Analysis)`.
+
+ The result can be manually overwritten by a user to return a user-defined result object, which can be extended
+ with additional methods and attribute specific to the model-fit.
+
+ This example class does example this, whereby the analysis result has been overwritten with the `ResultExample`
+ class, which contains a property `max_log_likelihood_model_data_1d` that returns the model data of the
+ best-fit model. This API means you can customize your result object to include whatever attributes you want
+ and therefore make a result object specific to your model-fit and model-fitting problem.
+
+ The `Result` object you return can be customized to include:
+
+ - The samples summary, which contains the maximum log likelihood instance and median PDF model.
+
+ - The paths of the search, which are used for loading the samples and search internal below when a search
+ is resumed.
+
+ - The samples of the non-linear search (e.g. MCMC chains) also stored in `samples.csv`.
+
+ - The non-linear search used for the fit in its internal representation, which is used for resuming a search
+ and making bespoke visualization using the search's internal results.
+
+ - The analysis used to fit the model (default disabled to save memory, but option may be useful for certain
+ projects).
+
+ Parameters
+ ----------
+ samples_summary
+ The summary of the samples of the non-linear search, which include the maximum log likelihood instance and
+ median PDF model.
+ paths
+ An object describing the paths for saving data (e.g. hard-disk directories or entries in sqlite database).
+ samples
+ The samples of the non-linear search, for example the chains of an MCMC run.
+ search_internal
+ The internal representation of the non-linear search used to perform the model-fit.
+ analysis
+ The analysis used to fit the model.
+
+ Returns
+ -------
+ Result
+ The result of the non-linear search, which is defined as a class variable in the `Analysis` class.
+ """
+ return self.Result(
+ samples_summary=samples_summary,
+ paths=paths,
+ samples=samples,
+ search_internal=search_internal,
+ analysis=self,
+ )
+
+
+"""
+For the sake of brevity, we do not run the code below, but the following code would work:
+
+`result = search.fit(model=model, analysis=analysis)`
+`print(result.max_log_likelihood_model_data_1d)`
+
+__Latent Variables__
+
+A latent variable is not a model parameter but can be derived from the model. Its value and errors may be of interest
+and aid in the interpretation of a model-fit.
+
+For example, for the simple 1D Gaussian example, it could be the full-width half maximum (FWHM) of the Gaussian.
+This is not included in the model but can be easily derived from the Gaussian's sigma value.
+
+By overwriting the Analysis class's `compute_latent_variables` method we can manually specify latent variables that
+are calculated. If the search has a `name`, these are output to a `latent.csv` file, which mirrors
+the `samples.csv` file.
+
+There may also be a `latent.results` and `latent_summary.json` files output. The `output.yaml` config file contains
+settings customizing what files are output and how often.
+
+This function takes as input the `parameters`, not the `instance`, because it means the function supports JAX.jit
+and thus if JAX is being used can be fully accelerated. The `instance` is created immediately inside the function.
+"""
+
+
+class Analysis(af.Analysis):
+
+ LATENT_KEYS = ["gaussian.fwhm"]
+
+ def __init__(self, data, noise_map):
+ """
+ An Analysis class which illustrates latent variables.
+ """
+ super().__init__()
+
+ self.data = data
+ self.noise_map = noise_map
+
+ def log_likelihood_function(self, instance):
+ """
+ The `log_likelihood_function` is identical to the example above
+ """
+ xvalues = np.arange(self.data.shape[0])
+
+ model_data = instance.model_data_from(xvalues=xvalues)
+ residual_map = self.data - model_data
+ chi_squared_map = (residual_map / self.noise_map) ** 2.0
+ chi_squared = sum(chi_squared_map)
+ noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
+ log_likelihood = -0.5 * (chi_squared + noise_normalization)
+
+ return log_likelihood
+
+ def compute_latent_variables(self, parameters, model) -> Tuple:
+ """
+ A latent variable is not a model parameter but can be derived from the model. Its value and errors may be
+ of interest and aid in the interpretation of a model-fit.
+
+ For example, for the simple 1D Gaussian example, it could be the full-width half maximum (FWHM) of the
+ Gaussian. This is not included in the model but can be easily derived from the Gaussian's sigma value.
+
+ By overwriting this method we can manually specify latent variables that are calculated and output to
+ a `latent.csv` file, which mirrors the `samples.csv` file.
+
+ In the example below, the `latent.csv` file will contain one column with the FWHM of every Gausian model
+ sampled by the non-linear search.
+
+ This function is called at the end of search, following one of two schemes depending on the settings in
+ `output.yaml`:
+
+ 1) Call for every search sample, which produces a complete `latent/samples.csv` which mirrors the normal
+ `samples.csv` file but takes a long time to compute.
+
+ 2) Call only for N random draws from the posterior inferred at the end of the search, which only produces a
+ `latent/latent_summary.json` file with the median and 1 and 3 sigma errors of the latent variables but is
+ fast to compute.
+
+ Parameters
+ ----------
+ instance
+ The instances of the model which the latent variable is derived from.
+
+ Returns
+ -------
+ A dictionary mapping every latent variable name to its value.
+ """
+
+ instance = model.instance_from_vector(vector=parameters)
+
+ return (instance.fwhm,)
+
+
+"""
+Outputting latent variables manually after a fit is complete is simple, just call
+the `analysis.compute_latent_variables()` function.
+
+For many use cases, the best set up disables autofit latent variable output during a fit via the `output.yaml`
+file and perform it manually after completing a successful model-fit. This will save computational run time by not
+computing latent variables during a any model-fit which is unsuccessful.
+"""
+analysis = Analysis(data=data, noise_map=noise_map)
+
+# Commented out because we do not run the search in this cookbook
+
+# latent_samples = analysis.compute_latent_variables(samples=result.samples)
+
+"""
+Analysing and interpreting latent variables is described fully in the result cookbook.
+
+However, in brief, the `latent_samples` object is a `Samples` object and uses the same API as samples objects.
+"""
+# print(latent_samples.median_pdf().gaussian.fwhm)
+
+"""
+__Custom Output__
+
+When performing fits which output results to hard-disc, a `files` folder is created containing .json / .csv files of
+the model, samples, search, etc.
+
+These files are human readable and help one quickly inspect and interpret results.
+
+By extending an `Analysis` class with the methods `save_attributes` and `save_results`,
+custom files can be written to the `files` folder to further aid this inspection.
+
+These files can then also be loaded via the database, as described in the database cookbook.
+"""
+
+
+class Analysis(af.Analysis):
+ def __init__(self, data: np.ndarray, noise_map: np.ndarray):
+ """
+ Standard Analysis class example used throughout PyAutoFit examples.
+ """
+ super().__init__()
+
+ self.data = data
+ self.noise_map = noise_map
+
+ def log_likelihood_function(self, instance) -> float:
+ """
+ Standard log likelihood function used throughout PyAutoFit examples.
+ """
+
+ xvalues = np.arange(self.data.shape[0])
+
+ model_data = instance.model_data_from(xvalues=xvalues)
+
+ residual_map = self.data - model_data
+ chi_squared_map = (residual_map / self.noise_map) ** 2.0
+ chi_squared = sum(chi_squared_map)
+ noise_normalization = np.sum(np.log(2 * np.pi * self.noise_map**2.0))
+ log_likelihood = -0.5 * (chi_squared + noise_normalization)
+
+ return log_likelihood
+
+ def save_attributes(self, paths: af.DirectoryPaths):
+ """
+ Before the non-linear search begins, this routine saves attributes of the `Analysis` object to the `files`
+ folder such that they can be loaded after the analysis using PyAutoFit's database and aggregator tools.
+
+ For this analysis, it uses the `AnalysisDataset` object's method to output the following:
+
+ - The dataset's data as a .json file.
+ - The dataset's noise-map as a .json file.
+
+ These are accessed using the aggregator via `agg.values("data")` and `agg.values("noise_map")`.
+
+ Parameters
+ ----------
+ paths
+ The paths object which manages all paths, e.g. where the non-linear search outputs are stored,
+ visualization, and the pickled objects used by the aggregator output by this function.
+ """
+ # The path where data.json is saved, e.g. output/dataset_name/unique_id/files/data.json
+
+ file_path = (path.join(paths._json_path, "data.json"),)
+
+ with open(file_path, "w+") as f:
+ json.dump(self.data, f, indent=4)
+
+ # The path where noise_map.json is saved, e.g. output/noise_mapset_name/unique_id/files/noise_map.json
+
+ file_path = (path.join(paths._json_path, "noise_map.json"),)
+
+ with open(file_path, "w+") as f:
+ json.dump(self.noise_map, f, indent=4)
+
+ def save_results(self, paths: af.DirectoryPaths, result: af.Result):
+ """
+ At the end of a model-fit, this routine saves attributes of the `Analysis` object to the `files`
+ folder such that they can be loaded after the analysis using PyAutoFit's database and aggregator tools.
+
+ For this analysis it outputs the following:
+
+ - The maximum log likelihood model data as a .json file.
+
+ This is accessed using the aggregator via `agg.values("model_data")`.
+
+ Parameters
+ ----------
+ paths
+ The paths object which manages all paths, e.g. where the non-linear search outputs are stored,
+ visualization and the pickled objects used by the aggregator output by this function.
+ result
+ The result of a model fit, including the non-linear search, samples and maximum likelihood model.
+ """
+ xvalues = np.arange(self.data.shape[0])
+
+ instance = result.max_log_likelihood_instance
+
+ model_data = instance.model_data_from(xvalues=xvalues)
+
+ # The path where model_data.json is saved, e.g. output/dataset_name/unique_id/files/model_data.json
+
+ file_path = (path.join(paths._json_path, "model_data.json"),)
+
+ with open(file_path, "w+") as f:
+ json.dump(model_data, f, indent=4)
+
+
+"""
+Finish.
+"""
diff --git a/scripts/cookbooks/configs.py b/scripts/cookbooks/configs.py
index 040a7daf..228fb7cf 100644
--- a/scripts/cookbooks/configs.py
+++ b/scripts/cookbooks/configs.py
@@ -1,303 +1,299 @@
-"""
-Cookbook: Config
-================
-
-Configuration files are used to control the behaviour model components in **PyAutoFit**, which perform the
-following tasks:
-
- - Specify the default priors of model components, so that a user does not have to manually specify priors every time
- they create a model.
-
- - Specify labels of every parameter, which are used for plotting and visualizing results.
-
-This cookbook illustrates how to create configuration files for your own model components, so that they can be used
-with **PyAutoFit**.
-
-__Contents__
-
- - No Config Behaviour: An example of what happens when a model component does not have a config file.
- - Templates: A template config file for specifying default model component priors.
- - Modules: Writing prior config files based on the Python module the model component Python class is contained in.
- - Labels: Config files which specify the labels of model component parameters for visualization.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import numpy as np
-from os import path
-
-import autofit as af
-
-"""
-__No Config Behaviour__
-
-The examples seen so far have used `Gaussian` and `Exponential` model components, which have configuration files in
-the `autofit_workspace/config/priors` folder which define their priors and labels.
-
-If a model component does not have a configuration file and we try to use it in a fit, **PyAutoFit** will raise an
-error.
-
-Lets illustrate this by setting up the usual Gaussian object, but naming it `GaussianNoConfig` so that it does
-not have a config file.
-"""
-
-
-class GaussianNoConfig:
- def __init__(
- self,
- centre=0.0, # <- PyAutoFit recognises these constructor arguments
- normalization=0.1, # <- are the Gaussian`s model parameters.
- sigma=0.01,
- ):
- """
- Represents a 1D `Gaussian` profile, which does not have a config file set up.
- """
- self.centre = centre
- self.normalization = normalization
- self.sigma = sigma
-
- def model_data_from(self, xvalues: np.ndarray) -> np.ndarray:
- """
- The usual method that returns the 1D data of the `Gaussian` profile.
- """
- transformed_xvalues = xvalues - self.centre
-
- return np.multiply(
- np.divide(self.normalization, self.sigma * np.sqrt(2.0 * np.pi)),
- np.exp(-0.5 * np.square(np.divide(transformed_xvalues, self.sigma))),
- )
-
-
-"""
-When we try make this a `Model` and fit it, **PyAutoFit** raises an error, as it does not know where the priors
-of the `GaussianNoConfig` are located.
-
-I have commented the Python code out below so the script can run, but feel free to uncomment it and run it to see the
-error.
-"""
-model = af.Model(GaussianNoConfig)
-
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-search = af.DynestyStatic()
-
-# result = search.fit(model=model, analysis=analysis)
-
-"""
-In all other examples, the fits runs because the priors have been defined in one of two ways:
-
- - They were manually input in the example script.
- - They were loaded via config files "behind the scenes".
-
-Checkout the folder `autofit_workspace/config/priors`, where .yaml files defining the priors of the `Gaussian` and
-`Exponential` model components are located. These are the config files that **PyAutoFit** loads in the background
-in order to setup the default priors of these model components.
-
-If we do not manually override priors, these are the priors that will be used by default when a model-fit is performed.
-
-__Templates__
-
-For your model-fitting task, you therefore should set up a config file for every model component you defining its
-default priors.
-
-Next, inspect the `TemplateObject.yaml` priors configuration file in `autofit_workspace/config/priors`.
-
-You should see the following text:
-
- parameter0:
- type: Uniform
- lower_limit: 0.0
- upper_limit: 1.0
- parameter1:
- type: TruncatedGaussian
- mean: 0.0
- sigma: 0.1
- lower_limit: 0.0
- upper_limit: inf
- parameter2:
- type: Uniform
- lower_limit: 0.0
- upper_limit: 10.0
-
-This specifies the default priors on two parameters, named `parameter0` and `parameter1`.
-
-The `type` is the type of prior assumed by **PyAutoFit** by default for its corresponding parameter, where in this
-example:
-
-- `parameter0` is given a `UniformPrior` with limits between 0.0 and 1.0.
-- `parameter1` a `TruncatedGaussianPrior` with mean 0.0 and sigma 1.0.
-- `parameter2` is given a `UniformPrior` with limits between 0.0 and 10.0.
-
-The `lower_limit` and `upper_limit` of a `TruncatedGaussianPrior` define the boundaries of what parameter values are
-physically allowed. If a model-component is given a value outside these limits during model-fitting the model is
-instantly resampled and discarded.
-
-We can easily adapt this template for any model component, for example the `GaussianNoConfig`.
-
-First, copy and paste the `TemplateObject.yaml` file to create a new file called `GaussianNoConfig.yaml`.
-
-The name of the class is matched to the name of the configuration file, therefore it is a requirement that the
-configuration file is named `GaussianNoConfig.yaml` so that **PyAutoFit** can associate it with the `GaussianNoConfig`
-Python class.
-
-Now perform the follow changes to the `.yaml` file:
-
- - Rename `parameter0` to `centre` and updates its uniform prior to be from a `lower_limit` of 0.0 and
- an `upper_limit` of 100.0.
- - Rename `parameter1` to `normalization`.
- - Rename `parameter2` to `sigma`.
-
-The `.yaml` file should read as follows:
-
- centre:
- type: Uniform
- lower_limit: 0.0
- upper_limit: 100.0
- normalization:
- type: TruncatedGaussian
- mean: 0.0
- sigma: 0.1
- lower_limit: 0.0
- upper_limit: inf
- sigma:
- type: Uniform
- lower_limit: 0.0
- upper_limit: 10.0
-
-We should now be able to make a `Model` of the `GaussianNoConfig` class and fit it, without manually specifying
-the priors.
-
-You may need to reset your Jupyter notebook's kernel for the changes to the `.yaml` file to take effect.
-"""
-model = af.Model(GaussianNoConfig)
-
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-search = af.DynestyStatic()
-
-# result = search.fit(model=model, analysis=analysis)
-
-"""
-__Modules__
-
-For larger projects, it may not be ideal to have to write a .yaml file for every Python class which acts as a model
-component.
-
-We instead would prefer them to be in their own dedicated Python module.
-
-Suppose the `Gaussian` and `Exponential` model components were contained in a module named `profiles.py` in your
-project's source code.
-
-You could then write a priors .yaml config file following the format given in the example config file
-`autofit_workspace/config/priors/profiles.yaml`, noting that there is a paring between the module name
-(`profiles.py`) and the name of the `.yaml` file (`profiles.yaml`).
-
-The file `autofit_workspace/config/priors/template_module.yaml` provides the tempolate for module based prior
-configs and reads as follows:
-
-ModelComponent0:
- parameter0:
- type: Uniform
- lower_limit: 0.0
- upper_limit: 1.0
- parameter1:
- type: LogUniform
- lower_limit: 1.0e-06
- upper_limit: 1000000.0
- parameter2:
- type: Uniform
- lower_limit: 0.0
- upper_limit: 25.0
-ModelComponent1:
- parameter0:
- type: Uniform
- lower_limit: 0.0
- upper_limit: 1.0
- parameter1:
- type: LogUniform
- lower_limit: 1.0e-06
- upper_limit: 1000000.0
- parameter2:
- type: Uniform
- lower_limit: 0.0
- upper_limit: 1.0
-
-This looks very similar to `TemplateObject`, the only differences are:
-
- - It now contains the model-component class name in the configuration file, e.g. `ModelComponent0`, `ModelComponent1`.
- - It includes multiple model-components, whereas `TemplateObject.yaml` corresponded to only one model component.
-
-Labels
-------
-
-There is an optional configs which associate model parameters with labels:
-
-`autofit_workspace/config/notation.yaml`
-
-It includes a `label` section which pairs every parameter with a label, which is used when visualizing results
-(e.g. these labels are used when creating a corner plot).
-
-label:
- label:
- sigma: \sigma
- centre: x
- normalization: norm
- parameter0: a
- parameter1: b
- parameter2: c
- rate: \lambda
-
-It also contains a `superscript` section which pairs every model-component label with a superscript, so that
-models with the same parameter names (e.g. `centre` can be distinguished).
-
-label:
- superscript:
- Exponential: e
- Gaussian: g
- ModelComponent0: M0
- ModelComponent1: M1
-
-The `label_format` section sets Python formatting options for every parameter, controlling how they display in
-the `model.results` file.
-label_format:
- format:
- sigma: '{:.2f}'
- centre: '{:.2f}'
- normalization: '{:.2f}'
- parameter0: '{:.2f}'
- parameter1: '{:.2f}'
- parameter2: '{:.2f}'
- rate: '{:.2f}'
-"""
+"""
+Cookbook: Config
+================
+
+Configuration files are used to control the behaviour model components in **PyAutoFit**, which perform the
+following tasks:
+
+ - Specify the default priors of model components, so that a user does not have to manually specify priors every time
+ they create a model.
+
+ - Specify labels of every parameter, which are used for plotting and visualizing results.
+
+This cookbook illustrates how to create configuration files for your own model components, so that they can be used
+with **PyAutoFit**.
+
+__Contents__
+
+ - No Config Behaviour: An example of what happens when a model component does not have a config file.
+ - Templates: A template config file for specifying default model component priors.
+ - Modules: Writing prior config files based on the Python module the model component Python class is contained in.
+ - Labels: Config files which specify the labels of model component parameters for visualization.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import numpy as np
+from os import path
+
+import autofit as af
+
+"""
+__No Config Behaviour__
+
+The examples seen so far have used `Gaussian` and `Exponential` model components, which have configuration files in
+the `autofit_workspace/config/priors` folder which define their priors and labels.
+
+If a model component does not have a configuration file and we try to use it in a fit, **PyAutoFit** will raise an
+error.
+
+Lets illustrate this by setting up the usual Gaussian object, but naming it `GaussianNoConfig` so that it does
+not have a config file.
+"""
+
+
+class GaussianNoConfig:
+ def __init__(
+ self,
+ centre=0.0, # <- PyAutoFit recognises these constructor arguments
+ normalization=0.1, # <- are the Gaussian`s model parameters.
+ sigma=0.01,
+ ):
+ """
+ Represents a 1D `Gaussian` profile, which does not have a config file set up.
+ """
+ self.centre = centre
+ self.normalization = normalization
+ self.sigma = sigma
+
+ def model_data_from(self, xvalues: np.ndarray) -> np.ndarray:
+ """
+ The usual method that returns the 1D data of the `Gaussian` profile.
+ """
+ transformed_xvalues = xvalues - self.centre
+
+ return np.multiply(
+ np.divide(self.normalization, self.sigma * np.sqrt(2.0 * np.pi)),
+ np.exp(-0.5 * np.square(np.divide(transformed_xvalues, self.sigma))),
+ )
+
+
+"""
+When we try make this a `Model` and fit it, **PyAutoFit** raises an error, as it does not know where the priors
+of the `GaussianNoConfig` are located.
+
+I have commented the Python code out below so the script can run, but feel free to uncomment it and run it to see the
+error.
+"""
+model = af.Model(GaussianNoConfig)
+
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+search = af.DynestyStatic()
+
+# result = search.fit(model=model, analysis=analysis)
+
+"""
+In all other examples, the fits runs because the priors have been defined in one of two ways:
+
+ - They were manually input in the example script.
+ - They were loaded via config files "behind the scenes".
+
+Checkout the folder `autofit_workspace/config/priors`, where .yaml files defining the priors of the `Gaussian` and
+`Exponential` model components are located. These are the config files that **PyAutoFit** loads in the background
+in order to setup the default priors of these model components.
+
+If we do not manually override priors, these are the priors that will be used by default when a model-fit is performed.
+
+__Templates__
+
+For your model-fitting task, you therefore should set up a config file for every model component you defining its
+default priors.
+
+Next, inspect the `TemplateObject.yaml` priors configuration file in `autofit_workspace/config/priors`.
+
+You should see the following text:
+
+ parameter0:
+ type: Uniform
+ lower_limit: 0.0
+ upper_limit: 1.0
+ parameter1:
+ type: TruncatedGaussian
+ mean: 0.0
+ sigma: 0.1
+ lower_limit: 0.0
+ upper_limit: inf
+ parameter2:
+ type: Uniform
+ lower_limit: 0.0
+ upper_limit: 10.0
+
+This specifies the default priors on two parameters, named `parameter0` and `parameter1`.
+
+The `type` is the type of prior assumed by **PyAutoFit** by default for its corresponding parameter, where in this
+example:
+
+- `parameter0` is given a `UniformPrior` with limits between 0.0 and 1.0.
+- `parameter1` a `TruncatedGaussianPrior` with mean 0.0 and sigma 1.0.
+- `parameter2` is given a `UniformPrior` with limits between 0.0 and 10.0.
+
+The `lower_limit` and `upper_limit` of a `TruncatedGaussianPrior` define the boundaries of what parameter values are
+physically allowed. If a model-component is given a value outside these limits during model-fitting the model is
+instantly resampled and discarded.
+
+We can easily adapt this template for any model component, for example the `GaussianNoConfig`.
+
+First, copy and paste the `TemplateObject.yaml` file to create a new file called `GaussianNoConfig.yaml`.
+
+The name of the class is matched to the name of the configuration file, therefore it is a requirement that the
+configuration file is named `GaussianNoConfig.yaml` so that **PyAutoFit** can associate it with the `GaussianNoConfig`
+Python class.
+
+Now perform the follow changes to the `.yaml` file:
+
+ - Rename `parameter0` to `centre` and updates its uniform prior to be from a `lower_limit` of 0.0 and
+ an `upper_limit` of 100.0.
+ - Rename `parameter1` to `normalization`.
+ - Rename `parameter2` to `sigma`.
+
+The `.yaml` file should read as follows:
+
+ centre:
+ type: Uniform
+ lower_limit: 0.0
+ upper_limit: 100.0
+ normalization:
+ type: TruncatedGaussian
+ mean: 0.0
+ sigma: 0.1
+ lower_limit: 0.0
+ upper_limit: inf
+ sigma:
+ type: Uniform
+ lower_limit: 0.0
+ upper_limit: 10.0
+
+We should now be able to make a `Model` of the `GaussianNoConfig` class and fit it, without manually specifying
+the priors.
+
+You may need to reset your Jupyter notebook's kernel for the changes to the `.yaml` file to take effect.
+"""
+model = af.Model(GaussianNoConfig)
+
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+search = af.DynestyStatic()
+
+# result = search.fit(model=model, analysis=analysis)
+
+"""
+__Modules__
+
+For larger projects, it may not be ideal to have to write a .yaml file for every Python class which acts as a model
+component.
+
+We instead would prefer them to be in their own dedicated Python module.
+
+Suppose the `Gaussian` and `Exponential` model components were contained in a module named `profiles.py` in your
+project's source code.
+
+You could then write a priors .yaml config file following the format given in the example config file
+`autofit_workspace/config/priors/profiles.yaml`, noting that there is a paring between the module name
+(`profiles.py`) and the name of the `.yaml` file (`profiles.yaml`).
+
+The file `autofit_workspace/config/priors/template_module.yaml` provides the tempolate for module based prior
+configs and reads as follows:
+
+ModelComponent0:
+ parameter0:
+ type: Uniform
+ lower_limit: 0.0
+ upper_limit: 1.0
+ parameter1:
+ type: LogUniform
+ lower_limit: 1.0e-06
+ upper_limit: 1000000.0
+ parameter2:
+ type: Uniform
+ lower_limit: 0.0
+ upper_limit: 25.0
+ModelComponent1:
+ parameter0:
+ type: Uniform
+ lower_limit: 0.0
+ upper_limit: 1.0
+ parameter1:
+ type: LogUniform
+ lower_limit: 1.0e-06
+ upper_limit: 1000000.0
+ parameter2:
+ type: Uniform
+ lower_limit: 0.0
+ upper_limit: 1.0
+
+This looks very similar to `TemplateObject`, the only differences are:
+
+ - It now contains the model-component class name in the configuration file, e.g. `ModelComponent0`, `ModelComponent1`.
+ - It includes multiple model-components, whereas `TemplateObject.yaml` corresponded to only one model component.
+
+Labels
+------
+
+There is an optional configs which associate model parameters with labels:
+
+`autofit_workspace/config/notation.yaml`
+
+It includes a `label` section which pairs every parameter with a label, which is used when visualizing results
+(e.g. these labels are used when creating a corner plot).
+
+label:
+ label:
+ sigma: \sigma
+ centre: x
+ normalization: norm
+ parameter0: a
+ parameter1: b
+ parameter2: c
+ rate: \lambda
+
+It also contains a `superscript` section which pairs every model-component label with a superscript, so that
+models with the same parameter names (e.g. `centre` can be distinguished).
+
+label:
+ superscript:
+ Exponential: e
+ Gaussian: g
+ ModelComponent0: M0
+ ModelComponent1: M1
+
+The `label_format` section sets Python formatting options for every parameter, controlling how they display in
+the `model.results` file.
+label_format:
+ format:
+ sigma: '{:.2f}'
+ centre: '{:.2f}'
+ normalization: '{:.2f}'
+ parameter0: '{:.2f}'
+ parameter1: '{:.2f}'
+ parameter2: '{:.2f}'
+ rate: '{:.2f}'
+"""
diff --git a/scripts/cookbooks/model.py b/scripts/cookbooks/model.py
index 84cbd118..77335db2 100644
--- a/scripts/cookbooks/model.py
+++ b/scripts/cookbooks/model.py
@@ -1,787 +1,783 @@
-"""
-Cookbook: Models
-================
-
-Model composition is the process of defining a probabilistic model as a collection of model components, which are
-ultimate fitted to a dataset via a non-linear search.
-
-This cookbook provides an overview of basic model composition tools.
-
-__Contents__
-
-**Models:**
-
-If first describes how to use the `af.Model` object to define models with a single model component from single
-Python classes, with the following sections:
-
- - Python Class Template: The template of a model component written as a Python class.
- - Model Composition (Model): Creating a model via `af.Model()`.
- - Priors (Model): How the default priors of a model are set and how to customize them.
- - Instances (Model): Creating an instance of a model via input parameters.
- - Model Customization (Model): Customizing a model (e.g. fixing parameters or linking them to one another).
- - Tuple Parameters (Model): Defining model components with parameters that are tuples.
- - Json Output (Model): Output a model in human readable text via a .json file and loading it back again.
-
-**Collections:**
-
-It then describes how to use the `af.Collection` object to define models with many model components from multiple
-Python classes, with the following sections:
-
- - Model Composition (Collection): Creating a model via `af.Collection()`.
- - Priors (Collection): How the default priors of a collection are set and how to customize them.
- - Instances (Collection): Create an instance of a collection via input parameters.
- - Model Customization (Collection): Customize a collection (e.g. fixing parameters or linking them to one another).
- - Json Output (Collection): Output a collection in human readable text via a .json file and loading it back again.
- - Extensible Models (Collection): Using collections to extend models with new model components, including the use
- of Python dictionaries and lists.
-
-**Arrays:**
-
-The cookbook next describes using NumPy arrays via tbe `af.Array` object to compose models, where each entry of the
-array is a free parameters, therefore offering maximum flexibility with the number of free parameter. This has
-the following sections:
-
- - Model Composition (af.Array): Composing models using NumPy arrays and `af.Array`().
- - Prior Customization (af.Array): How to customize the priors of a numpy array model.
- - Instances (af.Array): Create an instance of a numpy array model via input parameters.
- - Model Customization (af.Array): Customize a numpy array model (e.g. fixing parameters or linking them to one another).
- - Json Output (af.Array): Output a numpy array model in human readable text via a .json file and loading it back again.
- - Extensible Models (af.Array): Using numpy arrays to compose models with a flexible number of parameters.
- - Wrap Up: A summary of model composition in PyAutoFit.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import json
-import os
-from os import path
-from typing import Tuple
-
-import autofit as af
-
-"""
-__Python Class Template__
-
-A model component is written as a Python class using the following format:
-
- - The name of the class is the name of the model component, in this case, “Gaussian”.
-
- - The input arguments of the constructor are the parameters of the mode (here `centre`, `normalization` and `sigma`).
-
- - The default values and typing of the input arguments tell PyAutoFit whether a parameter is a single-valued float or
- a multi-valued tuple.
-
-We define a 1D Gaussian model component to illustrate model composition in PyAutoFit.
-"""
-
-
-class Gaussian:
- def __init__(
- self,
- centre: float = 30.0, # <- **PyAutoFit** recognises these constructor arguments
- normalization: float = 1.0, # <- are the Gaussian``s model parameters.
- sigma: float = 5.0,
- ):
- self.centre = centre
- self.normalization = normalization
- self.sigma = sigma
-
-
-"""
-__Model Composition (Model)__
-
-We can instantiate a Python class as a model component using `af.Model()`.
-"""
-model = af.Model(Gaussian)
-
-"""
-The model has 3 free parameters, corresponding to the 3 parameters defined above (`centre`, `normalization`
-and `sigma`).
-
-Each parameter has a prior associated with it, meaning they are fitted for if the model is passed to a non-linear
-search.
-"""
-print(f"Model Total Free Parameters = {model.total_free_parameters}")
-
-"""
-If we print the `info` attribute of the model we get information on all of the parameters and their priors.
-"""
-print(model.info)
-
-"""
-__Priors (Model)__
-
-The model has a set of default priors, which have been loaded from a config file in the PyAutoFit workspace.
-
-The config cookbook describes how to setup config files in order to produce custom priors, which means that you do not
-need to manually specify priors in your Python code every time you compose a model.
-
-If you do not setup config files, all priors must be manually specified before you fit the model, as shown below.
-"""
-model = af.Model(Gaussian)
-model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.normalization = af.LogUniformPrior(lower_limit=1e-4, upper_limit=1e4)
-model.sigma = af.TruncatedGaussianPrior(
- mean=0.0, sigma=1.0, lower_limit=0.0, upper_limit=1e5
-)
-
-"""
-__Instances (Model)__
-
-Instances of the model components above (created via `af.Model`) can be created, where an input `vector` of
-parameters is mapped to create an instance of the Python class of the model.
-
-We first need to know the order of parameters in the model, so we know how to define the input `vector`. This
-information is contained in the models `paths` attribute.
-"""
-print(model.paths)
-
-"""
-We create an `instance` of the `Gaussian` class via the model where `centre=30.0`, `normalization=2.0` and `sigma=3.0`.
-"""
-instance = model.instance_from_vector(vector=[30.0, 2.0, 3.0])
-
-print("\nModel Instance:")
-print(instance)
-
-print("\nInstance Parameters:\n")
-print("centre = ", instance.centre)
-print("normalization = ", instance.normalization)
-print("sigma = ", instance.sigma)
-
-"""
-We can create an `instance` by inputting unit values (e.g. between 0.0 and 1.0) which are mapped to the input values
-via the priors.
-
-The inputs of 0.5 below are mapped as follows:
-
- - `centre`: goes to 0.5 because this is the midpoint of a `UniformPrior` with `lower_limit=0.0` and `upper_limit=1.0`.
-
- - `normalization` goes to 1.0 because this is the midpoint of the `LogUniformPrior`' with `lower_limit=1e-4`
- and `upper_limit=1e4`, corresponding to log10 space.
-
- - `sigma`: goes to 0.0 because this is the `mean` of the `TruncatedGaussianPrior`.
-"""
-instance = model.instance_from_unit_vector(unit_vector=[0.5, 0.5, 0.5])
-
-print("\nModel Instance:")
-print(instance)
-
-print("\nInstance Parameters:\n")
-print("centre = ", instance.centre)
-print("normalization = ", instance.normalization)
-print("sigma = ", instance.sigma)
-
-"""
-We can create instances of the `Gaussian` using the median value of the prior of every parameter.
-"""
-instance = model.instance_from_prior_medians()
-
-print("\nInstance Parameters:\n")
-print("centre = ", instance.centre)
-print("normalization = ", instance.normalization)
-print("sigma = ", instance.sigma)
-
-"""
-We can create a random instance, where the random values are unit values drawn between 0.0 and 1.0.
-
-This means the parameter values of this instance are randomly drawn from the priors.
-"""
-model = af.Model(Gaussian)
-instance = model.random_instance()
-
-"""
-__Model Customization (Model)__
-
-We can fix a free parameter to a specific value (reducing the dimensionality of parameter space by 1):
-"""
-model = af.Model(Gaussian)
-model.centre = 0.0
-
-print(
- f"\n Model Total Free Parameters After Fixing Centre = {model.total_free_parameters}"
-)
-
-"""
-We can link two parameters together such they always assume the same value (reducing the dimensionality of
-parameter space by 1):
-"""
-model.centre = model.normalization
-
-print(
- f"\n Model Total Free Parameters After Linking Parameters = {model.total_free_parameters}"
-)
-
-"""
-Offsets between linked parameters or with certain values are possible:
-"""
-model.centre = model.normalization + model.sigma
-
-print(
- f"Model Total Free Parameters After Linking Parameters = {model.total_free_parameters}"
-)
-
-"""
-Assertions remove regions of parameter space (but do not reduce the dimensionality of parameter space):
-"""
-model.add_assertion(model.sigma > 5.0)
-model.add_assertion(model.centre > model.normalization)
-
-"""
-The customized model can be inspected by printing its `info` attribute.
-"""
-print(model.info)
-
-"""
-The overwriting of priors shown above can be achieved via the following alternative API:
-"""
-model = af.Model(
- Gaussian,
- centre=af.UniformPrior(lower_limit=0.0, upper_limit=1.0),
- normalization=af.LogUniformPrior(lower_limit=1e-4, upper_limit=1e4),
- sigma=af.GaussianPrior(mean=0.0, sigma=1.0),
-)
-
-"""
-This API can also be used for fixing a parameter to a certain value:
-"""
-model = af.Model(Gaussian, centre=0.0)
-
-
-"""
-__Tuple Parameters__
-
-The `Gaussian` model component above only has parameters that are single-valued floats.
-
-Parameters can also be tuples, which is useful for defining model components where certain parameters are naturally
-grouped together.
-
-For example, we can define a 2D Gaussian with a center that has two coordinates and therefore free parameters, (x, y),
-using a tuple.
-"""
-
-
-class Gaussian2D:
- def __init__(
- self,
- centre: Tuple[float, float] = (0.0, 0.0),
- normalization: float = 0.1,
- sigma: float = 1.0,
- ):
- self.centre = centre
- self.normalization = normalization
- self.sigma = sigma
-
-
-"""
-The model's `total_free_parameters` attribute now includes 4 free parameters, as the tuple `centre` parameter accounts
-for 2 free parameters.
-"""
-model = af.Model(Gaussian2D)
-
-print(f"Model Total Free Parameters = {model.total_free_parameters}")
-
-"""
-This information is again displayed in the `info` attribute:
-"""
-print("\nInfo:")
-print(model.info)
-
-"""
-Here are examples of how model customization can be applied to a model with tuple parameters:
-"""
-model = af.Model(Gaussian2D)
-model.centre = (0.0, 0.0)
-
-model.centre_0 = model.normalization
-
-model.centre_1 = model.normalization + model.sigma
-
-model.add_assertion(model.centre_0 > model.normalization)
-
-"""
-__Json Outputs (Model)__
-
-A model has a `dict` attribute, which expresses all information about the model as a Python dictionary.
-
-By printing this dictionary we can therefore get a concise summary of the model.
-"""
-model = af.Model(Gaussian)
-
-print(model.dict())
-
-"""
-The dictionary representation printed above can be saved to hard disk as a `.json` file.
-
-This means we can save any **PyAutoFit** model to hard-disk in a human readable format.
-
-Checkout the file `autofit_workspace/*/cookbooks/jsons/collection.json` to see the model written as a .json.
-"""
-model_path = path.join("scripts", "cookbooks", "jsons")
-
-os.makedirs(model_path, exist_ok=True)
-
-model_file = path.join(model_path, "model.json")
-
-with open(model_file, "w+") as f:
- json.dump(model.dict(), f, indent=4)
-
-"""
-We can load the model from its `.json` file, meaning that one can easily save a model to hard disk and load it
-elsewhere.
-"""
-model = af.Model.from_json(file=model_file)
-
-print(model.info)
-
-"""
-__Model Composition (Collection)__
-
-To illustrate `Collection` objects we define a second model component, representing a `Exponential` profile.
-"""
-
-
-class Exponential:
- def __init__(
- self,
- centre=0.0, # <- PyAutoFit recognises these constructor arguments are the model
- normalization=0.1, # <- parameters of the Exponential.
- rate=0.01,
- ):
- self.centre = centre
- self.normalization = normalization
- self.rate = rate
-
-
-"""
-To instantiate multiple Python classes into a combined model component we combine the `af.Collection()` and `af.Model()`
-objects.
-
-By passing the key word arguments `gaussian` and `exponential` below, these are used as the names of the attributes of
-instances created using this model (which is illustrated clearly below).
-"""
-model = af.Collection(gaussian=af.Model(Gaussian), exponential=af.Model(Exponential))
-
-"""
-We can check the model has a `total_free_parameters` of 6, meaning the 3 parameters defined
-above (`centre`, `normalization`, `sigma` and `rate`) for both the `Gaussian` and `Exponential` classes all have
-priors associated with them .
-
-This also means each parameter is fitted for if we fitted the model to data via a non-linear search.
-"""
-print(f"Model Total Free Parameters = {model.total_free_parameters}")
-
-"""
-Printing the `info` attribute of the model gives us information on all of the parameters.
-"""
-print(model.info)
-
-"""
-__Priors (Collection)__
-
-The model has a set of default priors, which have been loaded from a config file in the PyAutoFit workspace.
-
-The configs cookbook describes how to setup config files in order to produce custom priors, which means that you do not
-need to manually specify priors in your Python code every time you compose a model.
-
-If you do not setup config files, all priors must be manually specified before you fit the model, as shown below.
-"""
-model.gaussian.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.gaussian.normalization = af.UniformPrior(lower_limit=0.0, upper_limit=1e2)
-model.gaussian.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
-model.exponential.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.exponential.normalization = af.UniformPrior(lower_limit=0.0, upper_limit=1e2)
-model.exponential.rate = af.UniformPrior(lower_limit=0.0, upper_limit=10.0)
-
-"""
-When creating a model via a `Collection`, there is no need to actually pass the python classes as an `af.Model()`
-because **PyAutoFit** implicitly assumes they are to be created as a `Model()`.
-
-This enables more concise code, whereby the following code:
-"""
-model = af.Collection(gaussian=af.Model(Gaussian), exponential=af.Model(Exponential))
-
-"""
-Can instead be written as:
-"""
-model = af.Collection(gaussian=Gaussian, exponential=Exponential)
-
-"""
-__Instances (Collection)__
-
-We can create an instance of collection containing both the `Gaussian` and `Exponential` classes using this model.
-
-We create an `instance` where:
-
-- The `Gaussian` class has `centre=30.0`, `normalization=2.0` and `sigma=3.0`.
-- The `Exponential` class has `centre=60.0`, `normalization=4.0` and `rate=1.0``.
-"""
-instance = model.instance_from_vector(vector=[30.0, 2.0, 3.0, 60.0, 4.0, 1.0])
-
-"""
-Because we passed the key word arguments `gaussian` and `exponential` above, these are the names of the attributes of
-instances created using this model (e.g. this is why we write `instance.gaussian`):
-"""
-
-print("\nModel Instance:")
-print(instance)
-
-print("\nInstance Parameters:\n")
-print("centre (Gaussian) = ", instance.gaussian.centre)
-print("normalization (Gaussian) = ", instance.gaussian.normalization)
-print("sigma (Gaussian) = ", instance.gaussian.sigma)
-print("centre (Exponential) = ", instance.exponential.centre)
-print("normalization (Exponential) = ", instance.exponential.normalization)
-print("rate (Exponential) = ", instance.exponential.rate)
-
-"""
-Alternatively, the instance's variables can also be accessed as a list, whereby instead of using attribute names
-(e.g. `gaussian_0`) we input the list index.
-
-Note that the order of the instance model components is determined from the order the components are input into the
-`Collection`.
-
-For example, for the line `af.Collection(gaussian=gaussian, exponential=exponential)`, the first entry in the list
-is the gaussian because it is the first input to the `Collection`.
-"""
-print("centre (Gaussian) = ", instance[0].centre)
-print("normalization (Gaussian) = ", instance[0].normalization)
-print("sigma (Gaussian) = ", instance[0].sigma)
-print("centre (Gaussian) = ", instance[1].centre)
-print("normalization (Gaussian) = ", instance[1].normalization)
-print("rate (Exponential) = ", instance[1].rate)
-
-"""
-__Model Customization (Collection)__
-
-By setting up each Model first the model can be customized using either of the API’s shown above:
-"""
-gaussian = af.Model(Gaussian)
-gaussian.normalization = 1.0
-gaussian.sigma = af.GaussianPrior(mean=0.0, sigma=1.0)
-
-exponential = af.Model(Exponential)
-exponential.centre = 50.0
-exponential.add_assertion(exponential.rate > 5.0)
-
-model = af.Collection(gaussian=gaussian, exponential=exponential)
-
-print(model.info)
-
-"""
-Below is an alternative API that can be used to create the same model as above.
-
-Which API is used is up to the user and which they find most intuitive.
-"""
-gaussian = af.Model(
- Gaussian, normalization=1.0, sigma=af.GaussianPrior(mean=0.0, sigma=1.0)
-)
-exponential = af.Model(Exponential, centre=50.0)
-exponential.add_assertion(exponential.rate > 5.0)
-
-model = af.Collection(gaussian=gaussian, exponential=exponential)
-
-print(model.info)
-
-"""
-After creating the model as a `Collection` we can customize it afterwards:
-"""
-model = af.Collection(gaussian=Gaussian, exponential=Exponential)
-
-model.gaussian.normalization = 1.0
-model.gaussian.sigma = af.GaussianPrior(mean=0.0, sigma=1.0)
-
-model.exponential.centre = 50.0
-model.exponential.add_assertion(exponential.rate > 5.0)
-
-print(model.info)
-
-"""
-__JSon Outputs (Collection)__
-
-A `Collection` has a `dict` attribute, which express all information about the model as a Python dictionary.
-
-By printing this dictionary we can therefore get a concise summary of the model.
-"""
-print(model.dict())
-
-"""
-Python dictionaries can easily be saved to hard disk as a `.json` file.
-
-This means we can save any **PyAutoFit** model to hard-disk.
-
-Checkout the file `autofit_workspace/*/model/jsons/model.json` to see the model written as a .json.
-"""
-model_path = path.join("scripts", "model", "jsons")
-
-os.makedirs(model_path, exist_ok=True)
-
-model_file = path.join(model_path, "collection.json")
-
-with open(model_file, "w+") as f:
- json.dump(model.dict(), f, indent=4)
-
-"""
-We can load the model from its `.json` file, meaning that one can easily save a model to hard disk and load it
-elsewhere.
-"""
-model = af.Model.from_json(file=model_file)
-
-print(f"\n Model via Json Prior Count = {model.prior_count}")
-
-
-"""
-__Extensible Models (Collection)__
-
-There is no limit to the number of components we can use to set up a model via a `Collection`.
-"""
-model = af.Collection(
- gaussian_0=Gaussian,
- gaussian_1=Gaussian,
- exponential_0=Exponential,
- exponential_1=Exponential,
- exponential_2=Exponential,
-)
-
-print(model.info)
-
-"""
-A model can be created via `af.Collection()` where a dictionary of `af.Model()` objects are passed to it.
-
-The two models created below are identical - one uses the API detailed above whereas the second uses a dictionary.
-"""
-model = af.Collection(gaussian_0=Gaussian, gaussian_1=Gaussian)
-print(model.info)
-
-model_dict = {"gaussian_0": Gaussian, "gaussian_1": Gaussian}
-model = af.Collection(**model_dict)
-print(model.info)
-
-"""
-The keys of the dictionary passed to the model (e.g. `gaussian_0` and `gaussian_1` above) are used to create the
-names of the attributes of instances of the model.
-"""
-instance = model.instance_from_vector(vector=[1.0, 2.0, 3.0, 4.0, 5.0, 6.0])
-
-print("\nModel Instance:")
-print(instance)
-
-print("\nInstance Parameters:\n")
-print("centre (Gaussian) = ", instance.gaussian_0.centre)
-print("normalization (Gaussian) = ", instance.gaussian_0.normalization)
-print("sigma (Gaussian) = ", instance.gaussian_0.sigma)
-print("centre (Gaussian) = ", instance.gaussian_1.centre)
-print("normalization (Gaussian) = ", instance.gaussian_1.normalization)
-print("sigma (Gaussian) = ", instance.gaussian_1.sigma)
-
-"""
-A list of model components can also be passed to an `af.Collection` to create a model:
-"""
-model = af.Collection([Gaussian, Gaussian])
-
-print(model.info)
-
-"""
-When a list is used, there is no string with which to name the model components (e.g. we do not input `gaussian_0`
-and `gaussian_1` anywhere.
-
-The `instance` therefore can only be accessed via list indexing.
-"""
-instance = model.instance_from_vector(vector=[1.0, 2.0, 3.0, 4.0, 5.0, 6.0])
-
-print("\nModel Instance:")
-print(instance)
-
-print("\nInstance Parameters:\n")
-print("centre (Gaussian) = ", instance[0].centre)
-print("normalization (Gaussian) = ", instance[0].normalization)
-print("sigma (Gaussian) = ", instance[0].sigma)
-print("centre (Gaussian) = ", instance[1].centre)
-print("normalization (Gaussian) = ", instance[1].normalization)
-print("sigma (Gaussian) = ", instance[1].sigma)
-
-"""
-__Model Composition (af.Array)__
-
-Models can be composed using NumPy arrays, where each element of the array is a free parameter.
-
-This offers a lot more flexibility than using `Model` and `Collection` objects, as the number of parameters in the
-model is chosen on initialization via the input of the `shape` attribute.
-
-For many use cases, this flexibility is key to ensuring model composition is as easy as possible, for example when
-a part of the model being fitted is a matrix of parameters which may change shape depending on the dataset being
-fitted.
-
-To compose models using NumPy arrays, we use the `af.Array` object.
-"""
-model = af.Array(
- shape=(2, 2),
- prior=af.GaussianPrior(mean=0.0, sigma=1.0),
-)
-
-"""
-Each element of the array is a free parameter, which for `shape=(2,2)` means the model has 4 free parameters.
-"""
-print(f"Model Total Free Parameters = {model.total_free_parameters}")
-
-"""
-The `info` attribute of the model gives information on all of the parameters and their priors.
-"""
-print(model.info)
-
-"""
-__Prior Customization (af.Array)__
-
-The prior of every parameter in the array is set via the `prior` input above.
-
-NumPy array models do not currently support default priors via config files, so all priors must be manually specified.
-
-The prior of every parameter in the array can be customized by normal NumPy array indexing:
-"""
-model = af.Array(shape=(2, 2), prior=af.GaussianPrior(mean=0.0, sigma=1.0))
-
-model[0, 0] = af.UniformPrior(lower_limit=0.0, upper_limit=1.0)
-model[0, 1] = af.LogUniformPrior(lower_limit=1e-4, upper_limit=1e4)
-model[1, 0] = af.GaussianPrior(mean=0.0, sigma=2.0)
-
-"""
-The `info` attribute shows the customized priors.
-"""
-print(model.info)
-
-"""
-__Instances (af.Array)__
-
-Instances of numpy array model components can be created, where an input `vector` of parameters is mapped to create
-an instance of the Python class of the model.
-
-If the priors of the numpy array are not customized, ordering of parameters goes from element [0,0] to [0,1] to [1,0],
-as shown by the `paths` attribute.
-"""
-model = af.Array(
- shape=(2, 2),
- prior=af.GaussianPrior(mean=0.0, sigma=1.0),
-)
-
-print(model.paths)
-
-"""
-An instance can then be created by passing a vector of parameters to the model via the `instance_from_vector` method.
-
-The `instance` created is a NumPy array, where each element is the value passed in the vector.
-"""
-instance = model.instance_from_vector(vector=[0.0, 1.0, 2.0, 3.0])
-
-print("\nModel Instance:")
-print(instance)
-
-"""
-Prior customization changes the order of the parameters, therefore if you customize the priors of the numpy
-array you must check the ordering of the parameters in the `paths` attribute before passing a vector to
-the `instance_from_vector`
-"""
-model[0, 0] = af.UniformPrior(lower_limit=0.0, upper_limit=1.0)
-model[0, 1] = af.LogUniformPrior(lower_limit=1e-4, upper_limit=1e4)
-model[1, 0] = af.GaussianPrior(mean=0.0, sigma=2.0)
-
-print(model.paths)
-
-instance = model.instance_from_vector(vector=[0.0, 1.0, 2.0, 3.0])
-
-print("\nModel Instance:")
-print(instance)
-
-"""
-__Model Customization (af.Array)__
-
-The model customization API for numpy array models is the same as for `af.Model` and `af.Collection` objects.
-"""
-model = af.Array(
- shape=(2, 2),
- prior=af.GaussianPrior(mean=0.0, sigma=1.0),
-)
-
-model[0, 0] = 50.0
-model[0, 1] = model[1, 0]
-model.add_assertion(model[1, 1] > 0.0)
-
-print(model.info)
-
-"""
-__JSon Outputs (af.Array)__
-
-An `Array` has a `dict` attribute, which express all information about the model as a Python dictionary.
-
-By printing this dictionary we can therefore get a concise summary of the model.
-"""
-model = af.Array(
- shape=(2, 2),
- prior=af.GaussianPrior(mean=0.0, sigma=1.0),
-)
-
-print(model.dict())
-
-"""
-Python dictionaries can easily be saved to hard disk as a `.json` file.
-
-This means we can save any **PyAutoFit** model to hard-disk.
-
-Checkout the file `autofit_workspace/*/model/jsons/array.json` to see the model written as a .json.
-"""
-model_path = path.join("scripts", "model", "jsons")
-
-os.makedirs(model_path, exist_ok=True)
-
-model_file = path.join(model_path, "array.json")
-
-with open(model_file, "w+") as f:
- json.dump(model.dict(), f, indent=4)
-
-"""
-We can load the model from its `.json` file, meaning that one can easily save a model to hard disk and load it
-elsewhere.
-"""
-model = af.Array.from_json(file=model_file)
-
-print(f"\n Model via Json Prior Count = {model.prior_count}")
-
-"""
-__Extensible Models (af.Array)__
-
-For `Model` objects, the number of parameters is fixed to those listed in the input Python class when the model is
-created.
-
-For `Collection` objects, the use of dictionaries and lists allows for the number of parameters to be extended, but it
-was still tied to the input Python classes when the model was created.
-
-For `Array` objects, the number of parameters is fully customizable, you choose the shape of the array and therefore
-the number of parameters in the model when you create it.
-
-This makes `Array` objects the most extensible and flexible way to compose models.
-
-You can also combine `Array` objects with `Collection` objects to create models with a mix of fixed and extensible
-parameters.
-"""
-model = af.Collection(
- gaussian=Gaussian,
- array=af.Array(shape=(3, 2), prior=af.GaussianPrior(mean=0.0, sigma=1.0)),
-)
-
-model.gaussian.sigma = 2.0
-model.array[0, 0] = 1.0
-
-print(model.info)
-
-"""
-__Wrap Up__
-
-This cookbook shows how to compose models consisting of multiple components using the `af.Model()`
-and `af.Collection()` object.
-
-Advanced model composition uses multi-level models, which compose models from hierarchies of Python classes. This is
-described in the multi-level model cookbook.
-"""
+"""
+Cookbook: Models
+================
+
+Model composition is the process of defining a probabilistic model as a collection of model components, which are
+ultimate fitted to a dataset via a non-linear search.
+
+This cookbook provides an overview of basic model composition tools.
+
+__Contents__
+
+**Models:**
+
+If first describes how to use the `af.Model` object to define models with a single model component from single
+Python classes, with the following sections:
+
+ - Python Class Template: The template of a model component written as a Python class.
+ - Model Composition (Model): Creating a model via `af.Model()`.
+ - Priors (Model): How the default priors of a model are set and how to customize them.
+ - Instances (Model): Creating an instance of a model via input parameters.
+ - Model Customization (Model): Customizing a model (e.g. fixing parameters or linking them to one another).
+ - Tuple Parameters (Model): Defining model components with parameters that are tuples.
+ - Json Output (Model): Output a model in human readable text via a .json file and loading it back again.
+
+**Collections:**
+
+It then describes how to use the `af.Collection` object to define models with many model components from multiple
+Python classes, with the following sections:
+
+ - Model Composition (Collection): Creating a model via `af.Collection()`.
+ - Priors (Collection): How the default priors of a collection are set and how to customize them.
+ - Instances (Collection): Create an instance of a collection via input parameters.
+ - Model Customization (Collection): Customize a collection (e.g. fixing parameters or linking them to one another).
+ - Json Output (Collection): Output a collection in human readable text via a .json file and loading it back again.
+ - Extensible Models (Collection): Using collections to extend models with new model components, including the use
+ of Python dictionaries and lists.
+
+**Arrays:**
+
+The cookbook next describes using NumPy arrays via tbe `af.Array` object to compose models, where each entry of the
+array is a free parameters, therefore offering maximum flexibility with the number of free parameter. This has
+the following sections:
+
+ - Model Composition (af.Array): Composing models using NumPy arrays and `af.Array`().
+ - Prior Customization (af.Array): How to customize the priors of a numpy array model.
+ - Instances (af.Array): Create an instance of a numpy array model via input parameters.
+ - Model Customization (af.Array): Customize a numpy array model (e.g. fixing parameters or linking them to one another).
+ - Json Output (af.Array): Output a numpy array model in human readable text via a .json file and loading it back again.
+ - Extensible Models (af.Array): Using numpy arrays to compose models with a flexible number of parameters.
+ - Wrap Up: A summary of model composition in PyAutoFit.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import json
+import os
+from os import path
+from typing import Tuple
+
+import autofit as af
+
+"""
+__Python Class Template__
+
+A model component is written as a Python class using the following format:
+
+ - The name of the class is the name of the model component, in this case, “Gaussian”.
+
+ - The input arguments of the constructor are the parameters of the mode (here `centre`, `normalization` and `sigma`).
+
+ - The default values and typing of the input arguments tell PyAutoFit whether a parameter is a single-valued float or
+ a multi-valued tuple.
+
+We define a 1D Gaussian model component to illustrate model composition in PyAutoFit.
+"""
+
+
+class Gaussian:
+ def __init__(
+ self,
+ centre: float = 30.0, # <- **PyAutoFit** recognises these constructor arguments
+ normalization: float = 1.0, # <- are the Gaussian``s model parameters.
+ sigma: float = 5.0,
+ ):
+ self.centre = centre
+ self.normalization = normalization
+ self.sigma = sigma
+
+
+"""
+__Model Composition (Model)__
+
+We can instantiate a Python class as a model component using `af.Model()`.
+"""
+model = af.Model(Gaussian)
+
+"""
+The model has 3 free parameters, corresponding to the 3 parameters defined above (`centre`, `normalization`
+and `sigma`).
+
+Each parameter has a prior associated with it, meaning they are fitted for if the model is passed to a non-linear
+search.
+"""
+print(f"Model Total Free Parameters = {model.total_free_parameters}")
+
+"""
+If we print the `info` attribute of the model we get information on all of the parameters and their priors.
+"""
+print(model.info)
+
+"""
+__Priors (Model)__
+
+The model has a set of default priors, which have been loaded from a config file in the PyAutoFit workspace.
+
+The config cookbook describes how to setup config files in order to produce custom priors, which means that you do not
+need to manually specify priors in your Python code every time you compose a model.
+
+If you do not setup config files, all priors must be manually specified before you fit the model, as shown below.
+"""
+model = af.Model(Gaussian)
+model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.normalization = af.LogUniformPrior(lower_limit=1e-4, upper_limit=1e4)
+model.sigma = af.TruncatedGaussianPrior(
+ mean=0.0, sigma=1.0, lower_limit=0.0, upper_limit=1e5
+)
+
+"""
+__Instances (Model)__
+
+Instances of the model components above (created via `af.Model`) can be created, where an input `vector` of
+parameters is mapped to create an instance of the Python class of the model.
+
+We first need to know the order of parameters in the model, so we know how to define the input `vector`. This
+information is contained in the models `paths` attribute.
+"""
+print(model.paths)
+
+"""
+We create an `instance` of the `Gaussian` class via the model where `centre=30.0`, `normalization=2.0` and `sigma=3.0`.
+"""
+instance = model.instance_from_vector(vector=[30.0, 2.0, 3.0])
+
+print("\nModel Instance:")
+print(instance)
+
+print("\nInstance Parameters:\n")
+print("centre = ", instance.centre)
+print("normalization = ", instance.normalization)
+print("sigma = ", instance.sigma)
+
+"""
+We can create an `instance` by inputting unit values (e.g. between 0.0 and 1.0) which are mapped to the input values
+via the priors.
+
+The inputs of 0.5 below are mapped as follows:
+
+ - `centre`: goes to 0.5 because this is the midpoint of a `UniformPrior` with `lower_limit=0.0` and `upper_limit=1.0`.
+
+ - `normalization` goes to 1.0 because this is the midpoint of the `LogUniformPrior`' with `lower_limit=1e-4`
+ and `upper_limit=1e4`, corresponding to log10 space.
+
+ - `sigma`: goes to 0.0 because this is the `mean` of the `TruncatedGaussianPrior`.
+"""
+instance = model.instance_from_unit_vector(unit_vector=[0.5, 0.5, 0.5])
+
+print("\nModel Instance:")
+print(instance)
+
+print("\nInstance Parameters:\n")
+print("centre = ", instance.centre)
+print("normalization = ", instance.normalization)
+print("sigma = ", instance.sigma)
+
+"""
+We can create instances of the `Gaussian` using the median value of the prior of every parameter.
+"""
+instance = model.instance_from_prior_medians()
+
+print("\nInstance Parameters:\n")
+print("centre = ", instance.centre)
+print("normalization = ", instance.normalization)
+print("sigma = ", instance.sigma)
+
+"""
+We can create a random instance, where the random values are unit values drawn between 0.0 and 1.0.
+
+This means the parameter values of this instance are randomly drawn from the priors.
+"""
+model = af.Model(Gaussian)
+instance = model.random_instance()
+
+"""
+__Model Customization (Model)__
+
+We can fix a free parameter to a specific value (reducing the dimensionality of parameter space by 1):
+"""
+model = af.Model(Gaussian)
+model.centre = 0.0
+
+print(
+ f"\n Model Total Free Parameters After Fixing Centre = {model.total_free_parameters}"
+)
+
+"""
+We can link two parameters together such they always assume the same value (reducing the dimensionality of
+parameter space by 1):
+"""
+model.centre = model.normalization
+
+print(
+ f"\n Model Total Free Parameters After Linking Parameters = {model.total_free_parameters}"
+)
+
+"""
+Offsets between linked parameters or with certain values are possible:
+"""
+model.centre = model.normalization + model.sigma
+
+print(
+ f"Model Total Free Parameters After Linking Parameters = {model.total_free_parameters}"
+)
+
+"""
+Assertions remove regions of parameter space (but do not reduce the dimensionality of parameter space):
+"""
+model.add_assertion(model.sigma > 5.0)
+model.add_assertion(model.centre > model.normalization)
+
+"""
+The customized model can be inspected by printing its `info` attribute.
+"""
+print(model.info)
+
+"""
+The overwriting of priors shown above can be achieved via the following alternative API:
+"""
+model = af.Model(
+ Gaussian,
+ centre=af.UniformPrior(lower_limit=0.0, upper_limit=1.0),
+ normalization=af.LogUniformPrior(lower_limit=1e-4, upper_limit=1e4),
+ sigma=af.GaussianPrior(mean=0.0, sigma=1.0),
+)
+
+"""
+This API can also be used for fixing a parameter to a certain value:
+"""
+model = af.Model(Gaussian, centre=0.0)
+
+
+"""
+__Tuple Parameters__
+
+The `Gaussian` model component above only has parameters that are single-valued floats.
+
+Parameters can also be tuples, which is useful for defining model components where certain parameters are naturally
+grouped together.
+
+For example, we can define a 2D Gaussian with a center that has two coordinates and therefore free parameters, (x, y),
+using a tuple.
+"""
+
+
+class Gaussian2D:
+ def __init__(
+ self,
+ centre: Tuple[float, float] = (0.0, 0.0),
+ normalization: float = 0.1,
+ sigma: float = 1.0,
+ ):
+ self.centre = centre
+ self.normalization = normalization
+ self.sigma = sigma
+
+
+"""
+The model's `total_free_parameters` attribute now includes 4 free parameters, as the tuple `centre` parameter accounts
+for 2 free parameters.
+"""
+model = af.Model(Gaussian2D)
+
+print(f"Model Total Free Parameters = {model.total_free_parameters}")
+
+"""
+This information is again displayed in the `info` attribute:
+"""
+print("\nInfo:")
+print(model.info)
+
+"""
+Here are examples of how model customization can be applied to a model with tuple parameters:
+"""
+model = af.Model(Gaussian2D)
+model.centre = (0.0, 0.0)
+
+model.centre_0 = model.normalization
+
+model.centre_1 = model.normalization + model.sigma
+
+model.add_assertion(model.centre_0 > model.normalization)
+
+"""
+__Json Outputs (Model)__
+
+A model has a `dict` attribute, which expresses all information about the model as a Python dictionary.
+
+By printing this dictionary we can therefore get a concise summary of the model.
+"""
+model = af.Model(Gaussian)
+
+print(model.dict())
+
+"""
+The dictionary representation printed above can be saved to hard disk as a `.json` file.
+
+This means we can save any **PyAutoFit** model to hard-disk in a human readable format.
+
+Checkout the file `autofit_workspace/*/cookbooks/jsons/collection.json` to see the model written as a .json.
+"""
+model_path = path.join("scripts", "cookbooks", "jsons")
+
+os.makedirs(model_path, exist_ok=True)
+
+model_file = path.join(model_path, "model.json")
+
+with open(model_file, "w+") as f:
+ json.dump(model.dict(), f, indent=4)
+
+"""
+We can load the model from its `.json` file, meaning that one can easily save a model to hard disk and load it
+elsewhere.
+"""
+model = af.Model.from_json(file=model_file)
+
+print(model.info)
+
+"""
+__Model Composition (Collection)__
+
+To illustrate `Collection` objects we define a second model component, representing a `Exponential` profile.
+"""
+
+
+class Exponential:
+ def __init__(
+ self,
+ centre=0.0, # <- PyAutoFit recognises these constructor arguments are the model
+ normalization=0.1, # <- parameters of the Exponential.
+ rate=0.01,
+ ):
+ self.centre = centre
+ self.normalization = normalization
+ self.rate = rate
+
+
+"""
+To instantiate multiple Python classes into a combined model component we combine the `af.Collection()` and `af.Model()`
+objects.
+
+By passing the key word arguments `gaussian` and `exponential` below, these are used as the names of the attributes of
+instances created using this model (which is illustrated clearly below).
+"""
+model = af.Collection(gaussian=af.Model(Gaussian), exponential=af.Model(Exponential))
+
+"""
+We can check the model has a `total_free_parameters` of 6, meaning the 3 parameters defined
+above (`centre`, `normalization`, `sigma` and `rate`) for both the `Gaussian` and `Exponential` classes all have
+priors associated with them .
+
+This also means each parameter is fitted for if we fitted the model to data via a non-linear search.
+"""
+print(f"Model Total Free Parameters = {model.total_free_parameters}")
+
+"""
+Printing the `info` attribute of the model gives us information on all of the parameters.
+"""
+print(model.info)
+
+"""
+__Priors (Collection)__
+
+The model has a set of default priors, which have been loaded from a config file in the PyAutoFit workspace.
+
+The configs cookbook describes how to setup config files in order to produce custom priors, which means that you do not
+need to manually specify priors in your Python code every time you compose a model.
+
+If you do not setup config files, all priors must be manually specified before you fit the model, as shown below.
+"""
+model.gaussian.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.gaussian.normalization = af.UniformPrior(lower_limit=0.0, upper_limit=1e2)
+model.gaussian.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
+model.exponential.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.exponential.normalization = af.UniformPrior(lower_limit=0.0, upper_limit=1e2)
+model.exponential.rate = af.UniformPrior(lower_limit=0.0, upper_limit=10.0)
+
+"""
+When creating a model via a `Collection`, there is no need to actually pass the python classes as an `af.Model()`
+because **PyAutoFit** implicitly assumes they are to be created as a `Model()`.
+
+This enables more concise code, whereby the following code:
+"""
+model = af.Collection(gaussian=af.Model(Gaussian), exponential=af.Model(Exponential))
+
+"""
+Can instead be written as:
+"""
+model = af.Collection(gaussian=Gaussian, exponential=Exponential)
+
+"""
+__Instances (Collection)__
+
+We can create an instance of collection containing both the `Gaussian` and `Exponential` classes using this model.
+
+We create an `instance` where:
+
+- The `Gaussian` class has `centre=30.0`, `normalization=2.0` and `sigma=3.0`.
+- The `Exponential` class has `centre=60.0`, `normalization=4.0` and `rate=1.0``.
+"""
+instance = model.instance_from_vector(vector=[30.0, 2.0, 3.0, 60.0, 4.0, 1.0])
+
+"""
+Because we passed the key word arguments `gaussian` and `exponential` above, these are the names of the attributes of
+instances created using this model (e.g. this is why we write `instance.gaussian`):
+"""
+
+print("\nModel Instance:")
+print(instance)
+
+print("\nInstance Parameters:\n")
+print("centre (Gaussian) = ", instance.gaussian.centre)
+print("normalization (Gaussian) = ", instance.gaussian.normalization)
+print("sigma (Gaussian) = ", instance.gaussian.sigma)
+print("centre (Exponential) = ", instance.exponential.centre)
+print("normalization (Exponential) = ", instance.exponential.normalization)
+print("rate (Exponential) = ", instance.exponential.rate)
+
+"""
+Alternatively, the instance's variables can also be accessed as a list, whereby instead of using attribute names
+(e.g. `gaussian_0`) we input the list index.
+
+Note that the order of the instance model components is determined from the order the components are input into the
+`Collection`.
+
+For example, for the line `af.Collection(gaussian=gaussian, exponential=exponential)`, the first entry in the list
+is the gaussian because it is the first input to the `Collection`.
+"""
+print("centre (Gaussian) = ", instance[0].centre)
+print("normalization (Gaussian) = ", instance[0].normalization)
+print("sigma (Gaussian) = ", instance[0].sigma)
+print("centre (Gaussian) = ", instance[1].centre)
+print("normalization (Gaussian) = ", instance[1].normalization)
+print("rate (Exponential) = ", instance[1].rate)
+
+"""
+__Model Customization (Collection)__
+
+By setting up each Model first the model can be customized using either of the API’s shown above:
+"""
+gaussian = af.Model(Gaussian)
+gaussian.normalization = 1.0
+gaussian.sigma = af.GaussianPrior(mean=0.0, sigma=1.0)
+
+exponential = af.Model(Exponential)
+exponential.centre = 50.0
+exponential.add_assertion(exponential.rate > 5.0)
+
+model = af.Collection(gaussian=gaussian, exponential=exponential)
+
+print(model.info)
+
+"""
+Below is an alternative API that can be used to create the same model as above.
+
+Which API is used is up to the user and which they find most intuitive.
+"""
+gaussian = af.Model(
+ Gaussian, normalization=1.0, sigma=af.GaussianPrior(mean=0.0, sigma=1.0)
+)
+exponential = af.Model(Exponential, centre=50.0)
+exponential.add_assertion(exponential.rate > 5.0)
+
+model = af.Collection(gaussian=gaussian, exponential=exponential)
+
+print(model.info)
+
+"""
+After creating the model as a `Collection` we can customize it afterwards:
+"""
+model = af.Collection(gaussian=Gaussian, exponential=Exponential)
+
+model.gaussian.normalization = 1.0
+model.gaussian.sigma = af.GaussianPrior(mean=0.0, sigma=1.0)
+
+model.exponential.centre = 50.0
+model.exponential.add_assertion(exponential.rate > 5.0)
+
+print(model.info)
+
+"""
+__JSon Outputs (Collection)__
+
+A `Collection` has a `dict` attribute, which express all information about the model as a Python dictionary.
+
+By printing this dictionary we can therefore get a concise summary of the model.
+"""
+print(model.dict())
+
+"""
+Python dictionaries can easily be saved to hard disk as a `.json` file.
+
+This means we can save any **PyAutoFit** model to hard-disk.
+
+Checkout the file `autofit_workspace/*/model/jsons/model.json` to see the model written as a .json.
+"""
+model_path = path.join("scripts", "model", "jsons")
+
+os.makedirs(model_path, exist_ok=True)
+
+model_file = path.join(model_path, "collection.json")
+
+with open(model_file, "w+") as f:
+ json.dump(model.dict(), f, indent=4)
+
+"""
+We can load the model from its `.json` file, meaning that one can easily save a model to hard disk and load it
+elsewhere.
+"""
+model = af.Model.from_json(file=model_file)
+
+print(f"\n Model via Json Prior Count = {model.prior_count}")
+
+
+"""
+__Extensible Models (Collection)__
+
+There is no limit to the number of components we can use to set up a model via a `Collection`.
+"""
+model = af.Collection(
+ gaussian_0=Gaussian,
+ gaussian_1=Gaussian,
+ exponential_0=Exponential,
+ exponential_1=Exponential,
+ exponential_2=Exponential,
+)
+
+print(model.info)
+
+"""
+A model can be created via `af.Collection()` where a dictionary of `af.Model()` objects are passed to it.
+
+The two models created below are identical - one uses the API detailed above whereas the second uses a dictionary.
+"""
+model = af.Collection(gaussian_0=Gaussian, gaussian_1=Gaussian)
+print(model.info)
+
+model_dict = {"gaussian_0": Gaussian, "gaussian_1": Gaussian}
+model = af.Collection(**model_dict)
+print(model.info)
+
+"""
+The keys of the dictionary passed to the model (e.g. `gaussian_0` and `gaussian_1` above) are used to create the
+names of the attributes of instances of the model.
+"""
+instance = model.instance_from_vector(vector=[1.0, 2.0, 3.0, 4.0, 5.0, 6.0])
+
+print("\nModel Instance:")
+print(instance)
+
+print("\nInstance Parameters:\n")
+print("centre (Gaussian) = ", instance.gaussian_0.centre)
+print("normalization (Gaussian) = ", instance.gaussian_0.normalization)
+print("sigma (Gaussian) = ", instance.gaussian_0.sigma)
+print("centre (Gaussian) = ", instance.gaussian_1.centre)
+print("normalization (Gaussian) = ", instance.gaussian_1.normalization)
+print("sigma (Gaussian) = ", instance.gaussian_1.sigma)
+
+"""
+A list of model components can also be passed to an `af.Collection` to create a model:
+"""
+model = af.Collection([Gaussian, Gaussian])
+
+print(model.info)
+
+"""
+When a list is used, there is no string with which to name the model components (e.g. we do not input `gaussian_0`
+and `gaussian_1` anywhere.
+
+The `instance` therefore can only be accessed via list indexing.
+"""
+instance = model.instance_from_vector(vector=[1.0, 2.0, 3.0, 4.0, 5.0, 6.0])
+
+print("\nModel Instance:")
+print(instance)
+
+print("\nInstance Parameters:\n")
+print("centre (Gaussian) = ", instance[0].centre)
+print("normalization (Gaussian) = ", instance[0].normalization)
+print("sigma (Gaussian) = ", instance[0].sigma)
+print("centre (Gaussian) = ", instance[1].centre)
+print("normalization (Gaussian) = ", instance[1].normalization)
+print("sigma (Gaussian) = ", instance[1].sigma)
+
+"""
+__Model Composition (af.Array)__
+
+Models can be composed using NumPy arrays, where each element of the array is a free parameter.
+
+This offers a lot more flexibility than using `Model` and `Collection` objects, as the number of parameters in the
+model is chosen on initialization via the input of the `shape` attribute.
+
+For many use cases, this flexibility is key to ensuring model composition is as easy as possible, for example when
+a part of the model being fitted is a matrix of parameters which may change shape depending on the dataset being
+fitted.
+
+To compose models using NumPy arrays, we use the `af.Array` object.
+"""
+model = af.Array(
+ shape=(2, 2),
+ prior=af.GaussianPrior(mean=0.0, sigma=1.0),
+)
+
+"""
+Each element of the array is a free parameter, which for `shape=(2,2)` means the model has 4 free parameters.
+"""
+print(f"Model Total Free Parameters = {model.total_free_parameters}")
+
+"""
+The `info` attribute of the model gives information on all of the parameters and their priors.
+"""
+print(model.info)
+
+"""
+__Prior Customization (af.Array)__
+
+The prior of every parameter in the array is set via the `prior` input above.
+
+NumPy array models do not currently support default priors via config files, so all priors must be manually specified.
+
+The prior of every parameter in the array can be customized by normal NumPy array indexing:
+"""
+model = af.Array(shape=(2, 2), prior=af.GaussianPrior(mean=0.0, sigma=1.0))
+
+model[0, 0] = af.UniformPrior(lower_limit=0.0, upper_limit=1.0)
+model[0, 1] = af.LogUniformPrior(lower_limit=1e-4, upper_limit=1e4)
+model[1, 0] = af.GaussianPrior(mean=0.0, sigma=2.0)
+
+"""
+The `info` attribute shows the customized priors.
+"""
+print(model.info)
+
+"""
+__Instances (af.Array)__
+
+Instances of numpy array model components can be created, where an input `vector` of parameters is mapped to create
+an instance of the Python class of the model.
+
+If the priors of the numpy array are not customized, ordering of parameters goes from element [0,0] to [0,1] to [1,0],
+as shown by the `paths` attribute.
+"""
+model = af.Array(
+ shape=(2, 2),
+ prior=af.GaussianPrior(mean=0.0, sigma=1.0),
+)
+
+print(model.paths)
+
+"""
+An instance can then be created by passing a vector of parameters to the model via the `instance_from_vector` method.
+
+The `instance` created is a NumPy array, where each element is the value passed in the vector.
+"""
+instance = model.instance_from_vector(vector=[0.0, 1.0, 2.0, 3.0])
+
+print("\nModel Instance:")
+print(instance)
+
+"""
+Prior customization changes the order of the parameters, therefore if you customize the priors of the numpy
+array you must check the ordering of the parameters in the `paths` attribute before passing a vector to
+the `instance_from_vector`
+"""
+model[0, 0] = af.UniformPrior(lower_limit=0.0, upper_limit=1.0)
+model[0, 1] = af.LogUniformPrior(lower_limit=1e-4, upper_limit=1e4)
+model[1, 0] = af.GaussianPrior(mean=0.0, sigma=2.0)
+
+print(model.paths)
+
+instance = model.instance_from_vector(vector=[0.0, 1.0, 2.0, 3.0])
+
+print("\nModel Instance:")
+print(instance)
+
+"""
+__Model Customization (af.Array)__
+
+The model customization API for numpy array models is the same as for `af.Model` and `af.Collection` objects.
+"""
+model = af.Array(
+ shape=(2, 2),
+ prior=af.GaussianPrior(mean=0.0, sigma=1.0),
+)
+
+model[0, 0] = 50.0
+model[0, 1] = model[1, 0]
+model.add_assertion(model[1, 1] > 0.0)
+
+print(model.info)
+
+"""
+__JSon Outputs (af.Array)__
+
+An `Array` has a `dict` attribute, which express all information about the model as a Python dictionary.
+
+By printing this dictionary we can therefore get a concise summary of the model.
+"""
+model = af.Array(
+ shape=(2, 2),
+ prior=af.GaussianPrior(mean=0.0, sigma=1.0),
+)
+
+print(model.dict())
+
+"""
+Python dictionaries can easily be saved to hard disk as a `.json` file.
+
+This means we can save any **PyAutoFit** model to hard-disk.
+
+Checkout the file `autofit_workspace/*/model/jsons/array.json` to see the model written as a .json.
+"""
+model_path = path.join("scripts", "model", "jsons")
+
+os.makedirs(model_path, exist_ok=True)
+
+model_file = path.join(model_path, "array.json")
+
+with open(model_file, "w+") as f:
+ json.dump(model.dict(), f, indent=4)
+
+"""
+We can load the model from its `.json` file, meaning that one can easily save a model to hard disk and load it
+elsewhere.
+"""
+model = af.Array.from_json(file=model_file)
+
+print(f"\n Model via Json Prior Count = {model.prior_count}")
+
+"""
+__Extensible Models (af.Array)__
+
+For `Model` objects, the number of parameters is fixed to those listed in the input Python class when the model is
+created.
+
+For `Collection` objects, the use of dictionaries and lists allows for the number of parameters to be extended, but it
+was still tied to the input Python classes when the model was created.
+
+For `Array` objects, the number of parameters is fully customizable, you choose the shape of the array and therefore
+the number of parameters in the model when you create it.
+
+This makes `Array` objects the most extensible and flexible way to compose models.
+
+You can also combine `Array` objects with `Collection` objects to create models with a mix of fixed and extensible
+parameters.
+"""
+model = af.Collection(
+ gaussian=Gaussian,
+ array=af.Array(shape=(3, 2), prior=af.GaussianPrior(mean=0.0, sigma=1.0)),
+)
+
+model.gaussian.sigma = 2.0
+model.array[0, 0] = 1.0
+
+print(model.info)
+
+"""
+__Wrap Up__
+
+This cookbook shows how to compose models consisting of multiple components using the `af.Model()`
+and `af.Collection()` object.
+
+Advanced model composition uses multi-level models, which compose models from hierarchies of Python classes. This is
+described in the multi-level model cookbook.
+"""
diff --git a/scripts/cookbooks/model_internal.py b/scripts/cookbooks/model_internal.py
index 06f4e8b0..3b13c317 100644
--- a/scripts/cookbooks/model_internal.py
+++ b/scripts/cookbooks/model_internal.py
@@ -25,11 +25,7 @@
- Serialization Round Trip: ``dict()`` and ``from_dict()`` for JSON persistence.
"""
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
+# from autoconf import setup_notebook; setup_notebook()
import autofit as af
import numpy as np
diff --git a/scripts/cookbooks/multi_level_model.py b/scripts/cookbooks/multi_level_model.py
index e3f1d311..40d94296 100644
--- a/scripts/cookbooks/multi_level_model.py
+++ b/scripts/cookbooks/multi_level_model.py
@@ -1,350 +1,346 @@
-"""
-Cookbook: Multi Level Models
-============================
-
-A multi level model is one where one or more of the input parameters in the model components `__init__`
-constructor are Python classes, as opposed to a float or tuple.
-
-The `af.Model()` object treats these Python classes as model components, enabling the composition of models where
-model components are grouped within other Python classes, in an object oriented fashion.
-
-This enables complex models which are intiutive and extensible to be composed.
-
-This cookbook provides an overview of multi-level model composition.
-
-__Contents__
-
- - Python Class Template: The template of multi level model components written as a Python class.
- - Model Composition: How to compose a multi-level model using the `af.Model()` object.
- - Instances: Creating an instance of a multi-level model via input parameters.
- - Why Use Multi-Level Models?: A description of the benefits of using multi-level models compared to a `Collection`.
- - Model Customization: Customizing a multi-level model (e.g. fixing parameters or linking them to one another).
- - Alternative API: Alternative API for multi-level models which may be more concise and readable for certain models.
- - Json Output (Model): Output a multi-level model in human readable text via a .json file and loading it back again.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import json
-import os
-from os import path
-from typing import List
-
-import autofit as af
-
-"""
-__Python Class Template__
-
-A multi-level model uses standard model components, which are written as a Python class with the usual format
-where the inputs of the `__init__` constructor are the model parameters.
-"""
-
-
-class Gaussian:
- def __init__(
- self,
- normalization=1.0, # <- **PyAutoFit** recognises these constructor arguments
- sigma=5.0, # <- are the Gaussian``s model parameters.
- ):
- self.normalization = normalization
- self.sigma = sigma
-
-
-"""
-The unique aspect of a multi-level model is that a Python class can then be defined where the inputs
-of its `__init__` constructor are instances of these model components.
-
-In the example below, the Python class which will be used to demonstrate a multi-level has an input `gaussian_list`,
-which takes as input a list of instances of the `Gaussian` class above.
-
-This class will represent many individual `Gaussian`'s, which share the same `centre` but have their own unique
-`normalization` and `sigma` values.
-"""
-
-
-class MultiLevelGaussians:
- def __init__(
- self,
- higher_level_centre: float = 50.0, # The centre of all Gaussians in the multi level component.
- gaussian_list: List[Gaussian] = None, # Contains a list of Gaussians
- ):
- self.higher_level_centre = higher_level_centre
-
- self.gaussian_list = gaussian_list
-
-
-"""
-__Model Composition__
-
-A multi-level model is instantiated via the af.Model() command, which is passed:
-
- - `MultiLevelGaussians`: To tell it that the model component will be a `MultiLevelGaussians` object.
- - `gaussian_list`: One or more `Gaussian`'s, each of which are created as an `af.Model()` object with free parameters.
-"""
-model = af.Model(
- MultiLevelGaussians, gaussian_list=[af.Model(Gaussian), af.Model(Gaussian)]
-)
-
-"""
-The multi-level model consists of two `Gaussian`'s, where their centres are shared as a parameter in the higher level
-model component.
-
-Total number of parameters is N=5 (x2 `normalizations`, `x2 `sigma`'s and x1 `higher_level_centre`).
-"""
-print(f"Model Total Free Parameters = {model.total_free_parameters}")
-
-"""
-The structure of the multi-level model, including the hierarchy of Python classes, is shown in the `model.info`.
-"""
-print(model.info)
-
-"""
-__Instances__
-
-Instances of a multi-level model can be created, where an input `vector` of parameters is mapped to create an instance
-of the Python class of the model.
-
-We first need to know the order of parameters in the model, so we know how to define the input `vector`. This
-information is contained in the models `paths` attribute.
-"""
-print(model.paths)
-
-"""
-We now create an instance via a multi-level model.
-
-Its attributes are structured differently to models composed via the `Collection` object..
-"""
-instance = model.instance_from_vector(vector=[1.0, 2.0, 3.0, 4.0, 5.0])
-
-print("Model Instance: \n")
-print(instance)
-
-print("Instance Parameters \n")
-print("Normalization (Gaussian 0) = ", instance.gaussian_list[0].normalization)
-print("Sigma (Gaussian 0) = ", instance.gaussian_list[0].sigma)
-print("Normalization (Gaussian 0) = ", instance.gaussian_list[1].normalization)
-print("Sigma (Gaussian 0) = ", instance.gaussian_list[1].sigma)
-print("Higher Level Centre= ", instance.higher_level_centre)
-
-"""
-__Why Use Multi Level Models?__
-
-An identical model in terms of functionality could of been created via the `Collection` object as follows:
-"""
-
-
-class GaussianCentre:
- def __init__(
- self,
- centre=30.0, # <- **PyAutoFit** recognises these constructor arguments
- normalization=1.0, # <- are the Gaussian``s model parameters.
- sigma=5.0,
- ):
- self.centre = centre
- self.normalization = normalization
- self.sigma = sigma
-
-
-model = af.Collection(gaussian_0=GaussianCentre, gaussian_1=GaussianCentre)
-
-model.gaussian_0.centre = model.gaussian_1.centre
-
-"""
-This raises the question of when to use a `Collection` and when to use multi-level models?
-
-The answer depends on the structure of the models you are composing and fitting.
-
-Many problems have models which have a natural multi-level structure.
-
-For example, imagine a dataset had 3 separate groups of 1D `Gaussian`'s, where each group had multiple Gaussians with
-a shared centre.
-
-This model is concise and easy to define using the multi-level API:
-"""
-group_0 = af.Model(MultiLevelGaussians, gaussian_list=3 * [Gaussian])
-
-group_1 = af.Model(MultiLevelGaussians, gaussian_list=3 * [Gaussian])
-
-group_2 = af.Model(MultiLevelGaussians, gaussian_list=3 * [Gaussian])
-
-model = af.Collection(group_0=group_0, group_1=group_1, group_2=group_2)
-
-print(model.info)
-
-"""
-Composing the same model without the multi-level model is less concise, less readable and prone to error:
-"""
-group_0 = af.Collection(
- gaussian_0=GaussianCentre, gaussian_1=GaussianCentre, gaussian_2=GaussianCentre
-)
-
-group_0.gaussian_0.centre = group_0.gaussian_1.centre
-group_0.gaussian_0.centre = group_0.gaussian_2.centre
-group_0.gaussian_1.centre = group_0.gaussian_2.centre
-
-group_1 = af.Collection(
- gaussian_0=GaussianCentre, gaussian_1=GaussianCentre, gaussian_2=GaussianCentre
-)
-
-group_1.gaussian_0.centre = group_1.gaussian_1.centre
-group_1.gaussian_0.centre = group_1.gaussian_2.centre
-group_1.gaussian_1.centre = group_1.gaussian_2.centre
-
-group_2 = af.Collection(
- gaussian_0=GaussianCentre, gaussian_1=GaussianCentre, gaussian_2=GaussianCentre
-)
-
-group_2.gaussian_0.centre = group_2.gaussian_1.centre
-group_2.gaussian_0.centre = group_2.gaussian_2.centre
-group_2.gaussian_1.centre = group_2.gaussian_2.centre
-
-model = af.Collection(group_0=group_0, group_1=group_1, group_2=group_2)
-
-"""
-In many situations, multi-levels models are more extensible than the `Collection` API.
-
-For example, imagine we wanted to add even more 1D profiles to a group with a shared `centre`. This can easily be
-achieved using the multi-level API:
-
- multi = af.Model(
- MultiLevelGaussians, gaussian_list=[Gaussian, Gaussian, Exponential, YourProfileHere]
- )
-
-Composing the same model using just a `Model` and `Collection` is again possible, but would be even more cumbersome,
-less readable and is not extensible.
-
-__Model Customization__
-
-To customize the higher level parameters of a multi-level the usual model API is used:
-"""
-multi = af.Model(MultiLevelGaussians, gaussian_list=[Gaussian, Gaussian])
-
-multi.higher_level_centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-
-"""
-To customize a multi-level model instantiated via lists, each model component is accessed via its index:
-"""
-multi = af.Model(MultiLevelGaussians, gaussian_list=[Gaussian, Gaussian])
-
-group_level = af.Model(MultiLevelGaussians, gaussian_list=[Gaussian, Gaussian])
-
-group_level.gaussian_list[0].normalization = group_level.gaussian_list[1].normalization
-
-"""
-Any combination of the API’s shown above can be used for customizing this model:
-"""
-gaussian_0 = af.Model(Gaussian)
-gaussian_1 = af.Model(Gaussian)
-
-gaussian_0.normalization = gaussian_1.normalization
-
-group_level = af.Model(
- MultiLevelGaussians, gaussian_list=[gaussian_0, gaussian_1, af.Model(Gaussian)]
-)
-
-group_level.higher_level_centre = 1.0
-group_level.gaussian_list[2].normalization = group_level.gaussian_list[1].normalization
-
-"""
-The `info` shows how the customization of the model has been performed:
-"""
-print(group_level.info)
-
-
-"""
-__Alternative API__
-
-A multi-level model can be instantiated where each model sub-component is setup using a name (as opposed to a list).
-
-This means no list input parameter is required in the Python class of the model component, but we do need to include
-the `**kwargs` input.
-"""
-
-
-class MultiLevelGaussians:
- def __init__(self, higher_level_centre=1.0, **kwargs):
- self.higher_level_centre = higher_level_centre
-
-
-model = af.Model(
- MultiLevelGaussians, gaussian_0=af.Model(Gaussian), gaussian_1=af.Model(Gaussian)
-)
-
-print(model)
-
-instance = model.instance_from_vector(vector=[1.0, 2.0, 3.0, 4.0, 5.0])
-
-print("Instance Parameters \n")
-print("Normalization (Gaussian 0) = ", instance.gaussian_0.normalization)
-print("Sigma (Gaussian 0) = ", instance.gaussian_0.sigma)
-print("Normalization (Gaussian 0) = ", instance.gaussian_1.normalization)
-print("Sigma (Gaussian 0) = ", instance.gaussian_1.sigma)
-print("Higher Level Centre= ", instance.higher_level_centre)
-
-"""
-The use of Python dictionaries illustrated in previous cookbooks can also be used with multi-level models.
-"""
-
-model_dict = {"gaussian_0": Gaussian, "gaussian_1": Gaussian}
-
-model = af.Model(MultiLevelGaussians, **model_dict)
-
-print(f"Multi-level Model Prior Count = {model.prior_count}")
-
-instance = model.instance_from_vector(vector=[1.0, 2.0, 3.0, 4.0, 5.0])
-
-print("Instance Parameters \n")
-print("Normalization (Gaussian 0) = ", instance.gaussian_0.normalization)
-print("Sigma (Gaussian 0) = ", instance.gaussian_0.sigma)
-print("Normalization (Gaussian 0) = ", instance.gaussian_1.normalization)
-print("Sigma (Gaussian 0) = ", instance.gaussian_1.sigma)
-print("Higher Level Centre= ", instance.higher_level_centre)
-
-"""
-__JSon Outputs__
-
-A model has a `dict` attribute, which expresses all information about the model as a Python dictionary.
-
-By printing this dictionary we can therefore get a concise summary of the model.
-"""
-model = af.Model(Gaussian)
-
-print(model.dict())
-
-"""
-The dictionary representation printed above can be saved to hard disk as a `.json` file.
-
-This means we can save any multi-level model to hard-disk in a human readable format.
-
-Checkout the file `autofit_workspace/*/cookbooks/jsons/group_level_model.json` to see the model written as a .json.
-"""
-model_path = path.join("scripts", "cookbooks", "jsons")
-
-os.makedirs(model_path, exist_ok=True)
-
-model_file = path.join(model_path, "multi_level_model.json")
-
-with open(model_file, "w+") as f:
- json.dump(model.dict(), f, indent=4)
-
-"""
-We can load the model from its `.json` file, meaning that one can easily save a model to hard disk and load it
-elsewhere.
-"""
-model = af.Model.from_json(file=model_file)
-
-print(model.info)
-
-"""
-__Wrap Up__
-
-This cookbook shows how to multi-level models consisting of multiple components using the `af.Model()`
-and `af.Collection()` objects.
-
-You should think carefully about whether your model fitting problem can use multi-level models, as they can make
-your model definition more concise and extensible.
-"""
+"""
+Cookbook: Multi Level Models
+============================
+
+A multi level model is one where one or more of the input parameters in the model components `__init__`
+constructor are Python classes, as opposed to a float or tuple.
+
+The `af.Model()` object treats these Python classes as model components, enabling the composition of models where
+model components are grouped within other Python classes, in an object oriented fashion.
+
+This enables complex models which are intiutive and extensible to be composed.
+
+This cookbook provides an overview of multi-level model composition.
+
+__Contents__
+
+ - Python Class Template: The template of multi level model components written as a Python class.
+ - Model Composition: How to compose a multi-level model using the `af.Model()` object.
+ - Instances: Creating an instance of a multi-level model via input parameters.
+ - Why Use Multi-Level Models?: A description of the benefits of using multi-level models compared to a `Collection`.
+ - Model Customization: Customizing a multi-level model (e.g. fixing parameters or linking them to one another).
+ - Alternative API: Alternative API for multi-level models which may be more concise and readable for certain models.
+ - Json Output (Model): Output a multi-level model in human readable text via a .json file and loading it back again.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import json
+import os
+from os import path
+from typing import List
+
+import autofit as af
+
+"""
+__Python Class Template__
+
+A multi-level model uses standard model components, which are written as a Python class with the usual format
+where the inputs of the `__init__` constructor are the model parameters.
+"""
+
+
+class Gaussian:
+ def __init__(
+ self,
+ normalization=1.0, # <- **PyAutoFit** recognises these constructor arguments
+ sigma=5.0, # <- are the Gaussian``s model parameters.
+ ):
+ self.normalization = normalization
+ self.sigma = sigma
+
+
+"""
+The unique aspect of a multi-level model is that a Python class can then be defined where the inputs
+of its `__init__` constructor are instances of these model components.
+
+In the example below, the Python class which will be used to demonstrate a multi-level has an input `gaussian_list`,
+which takes as input a list of instances of the `Gaussian` class above.
+
+This class will represent many individual `Gaussian`'s, which share the same `centre` but have their own unique
+`normalization` and `sigma` values.
+"""
+
+
+class MultiLevelGaussians:
+ def __init__(
+ self,
+ higher_level_centre: float = 50.0, # The centre of all Gaussians in the multi level component.
+ gaussian_list: List[Gaussian] = None, # Contains a list of Gaussians
+ ):
+ self.higher_level_centre = higher_level_centre
+
+ self.gaussian_list = gaussian_list
+
+
+"""
+__Model Composition__
+
+A multi-level model is instantiated via the af.Model() command, which is passed:
+
+ - `MultiLevelGaussians`: To tell it that the model component will be a `MultiLevelGaussians` object.
+ - `gaussian_list`: One or more `Gaussian`'s, each of which are created as an `af.Model()` object with free parameters.
+"""
+model = af.Model(
+ MultiLevelGaussians, gaussian_list=[af.Model(Gaussian), af.Model(Gaussian)]
+)
+
+"""
+The multi-level model consists of two `Gaussian`'s, where their centres are shared as a parameter in the higher level
+model component.
+
+Total number of parameters is N=5 (x2 `normalizations`, `x2 `sigma`'s and x1 `higher_level_centre`).
+"""
+print(f"Model Total Free Parameters = {model.total_free_parameters}")
+
+"""
+The structure of the multi-level model, including the hierarchy of Python classes, is shown in the `model.info`.
+"""
+print(model.info)
+
+"""
+__Instances__
+
+Instances of a multi-level model can be created, where an input `vector` of parameters is mapped to create an instance
+of the Python class of the model.
+
+We first need to know the order of parameters in the model, so we know how to define the input `vector`. This
+information is contained in the models `paths` attribute.
+"""
+print(model.paths)
+
+"""
+We now create an instance via a multi-level model.
+
+Its attributes are structured differently to models composed via the `Collection` object..
+"""
+instance = model.instance_from_vector(vector=[1.0, 2.0, 3.0, 4.0, 5.0])
+
+print("Model Instance: \n")
+print(instance)
+
+print("Instance Parameters \n")
+print("Normalization (Gaussian 0) = ", instance.gaussian_list[0].normalization)
+print("Sigma (Gaussian 0) = ", instance.gaussian_list[0].sigma)
+print("Normalization (Gaussian 0) = ", instance.gaussian_list[1].normalization)
+print("Sigma (Gaussian 0) = ", instance.gaussian_list[1].sigma)
+print("Higher Level Centre= ", instance.higher_level_centre)
+
+"""
+__Why Use Multi Level Models?__
+
+An identical model in terms of functionality could of been created via the `Collection` object as follows:
+"""
+
+
+class GaussianCentre:
+ def __init__(
+ self,
+ centre=30.0, # <- **PyAutoFit** recognises these constructor arguments
+ normalization=1.0, # <- are the Gaussian``s model parameters.
+ sigma=5.0,
+ ):
+ self.centre = centre
+ self.normalization = normalization
+ self.sigma = sigma
+
+
+model = af.Collection(gaussian_0=GaussianCentre, gaussian_1=GaussianCentre)
+
+model.gaussian_0.centre = model.gaussian_1.centre
+
+"""
+This raises the question of when to use a `Collection` and when to use multi-level models?
+
+The answer depends on the structure of the models you are composing and fitting.
+
+Many problems have models which have a natural multi-level structure.
+
+For example, imagine a dataset had 3 separate groups of 1D `Gaussian`'s, where each group had multiple Gaussians with
+a shared centre.
+
+This model is concise and easy to define using the multi-level API:
+"""
+group_0 = af.Model(MultiLevelGaussians, gaussian_list=3 * [Gaussian])
+
+group_1 = af.Model(MultiLevelGaussians, gaussian_list=3 * [Gaussian])
+
+group_2 = af.Model(MultiLevelGaussians, gaussian_list=3 * [Gaussian])
+
+model = af.Collection(group_0=group_0, group_1=group_1, group_2=group_2)
+
+print(model.info)
+
+"""
+Composing the same model without the multi-level model is less concise, less readable and prone to error:
+"""
+group_0 = af.Collection(
+ gaussian_0=GaussianCentre, gaussian_1=GaussianCentre, gaussian_2=GaussianCentre
+)
+
+group_0.gaussian_0.centre = group_0.gaussian_1.centre
+group_0.gaussian_0.centre = group_0.gaussian_2.centre
+group_0.gaussian_1.centre = group_0.gaussian_2.centre
+
+group_1 = af.Collection(
+ gaussian_0=GaussianCentre, gaussian_1=GaussianCentre, gaussian_2=GaussianCentre
+)
+
+group_1.gaussian_0.centre = group_1.gaussian_1.centre
+group_1.gaussian_0.centre = group_1.gaussian_2.centre
+group_1.gaussian_1.centre = group_1.gaussian_2.centre
+
+group_2 = af.Collection(
+ gaussian_0=GaussianCentre, gaussian_1=GaussianCentre, gaussian_2=GaussianCentre
+)
+
+group_2.gaussian_0.centre = group_2.gaussian_1.centre
+group_2.gaussian_0.centre = group_2.gaussian_2.centre
+group_2.gaussian_1.centre = group_2.gaussian_2.centre
+
+model = af.Collection(group_0=group_0, group_1=group_1, group_2=group_2)
+
+"""
+In many situations, multi-levels models are more extensible than the `Collection` API.
+
+For example, imagine we wanted to add even more 1D profiles to a group with a shared `centre`. This can easily be
+achieved using the multi-level API:
+
+ multi = af.Model(
+ MultiLevelGaussians, gaussian_list=[Gaussian, Gaussian, Exponential, YourProfileHere]
+ )
+
+Composing the same model using just a `Model` and `Collection` is again possible, but would be even more cumbersome,
+less readable and is not extensible.
+
+__Model Customization__
+
+To customize the higher level parameters of a multi-level the usual model API is used:
+"""
+multi = af.Model(MultiLevelGaussians, gaussian_list=[Gaussian, Gaussian])
+
+multi.higher_level_centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+
+"""
+To customize a multi-level model instantiated via lists, each model component is accessed via its index:
+"""
+multi = af.Model(MultiLevelGaussians, gaussian_list=[Gaussian, Gaussian])
+
+group_level = af.Model(MultiLevelGaussians, gaussian_list=[Gaussian, Gaussian])
+
+group_level.gaussian_list[0].normalization = group_level.gaussian_list[1].normalization
+
+"""
+Any combination of the API’s shown above can be used for customizing this model:
+"""
+gaussian_0 = af.Model(Gaussian)
+gaussian_1 = af.Model(Gaussian)
+
+gaussian_0.normalization = gaussian_1.normalization
+
+group_level = af.Model(
+ MultiLevelGaussians, gaussian_list=[gaussian_0, gaussian_1, af.Model(Gaussian)]
+)
+
+group_level.higher_level_centre = 1.0
+group_level.gaussian_list[2].normalization = group_level.gaussian_list[1].normalization
+
+"""
+The `info` shows how the customization of the model has been performed:
+"""
+print(group_level.info)
+
+
+"""
+__Alternative API__
+
+A multi-level model can be instantiated where each model sub-component is setup using a name (as opposed to a list).
+
+This means no list input parameter is required in the Python class of the model component, but we do need to include
+the `**kwargs` input.
+"""
+
+
+class MultiLevelGaussians:
+ def __init__(self, higher_level_centre=1.0, **kwargs):
+ self.higher_level_centre = higher_level_centre
+
+
+model = af.Model(
+ MultiLevelGaussians, gaussian_0=af.Model(Gaussian), gaussian_1=af.Model(Gaussian)
+)
+
+print(model)
+
+instance = model.instance_from_vector(vector=[1.0, 2.0, 3.0, 4.0, 5.0])
+
+print("Instance Parameters \n")
+print("Normalization (Gaussian 0) = ", instance.gaussian_0.normalization)
+print("Sigma (Gaussian 0) = ", instance.gaussian_0.sigma)
+print("Normalization (Gaussian 0) = ", instance.gaussian_1.normalization)
+print("Sigma (Gaussian 0) = ", instance.gaussian_1.sigma)
+print("Higher Level Centre= ", instance.higher_level_centre)
+
+"""
+The use of Python dictionaries illustrated in previous cookbooks can also be used with multi-level models.
+"""
+
+model_dict = {"gaussian_0": Gaussian, "gaussian_1": Gaussian}
+
+model = af.Model(MultiLevelGaussians, **model_dict)
+
+print(f"Multi-level Model Prior Count = {model.prior_count}")
+
+instance = model.instance_from_vector(vector=[1.0, 2.0, 3.0, 4.0, 5.0])
+
+print("Instance Parameters \n")
+print("Normalization (Gaussian 0) = ", instance.gaussian_0.normalization)
+print("Sigma (Gaussian 0) = ", instance.gaussian_0.sigma)
+print("Normalization (Gaussian 0) = ", instance.gaussian_1.normalization)
+print("Sigma (Gaussian 0) = ", instance.gaussian_1.sigma)
+print("Higher Level Centre= ", instance.higher_level_centre)
+
+"""
+__JSon Outputs__
+
+A model has a `dict` attribute, which expresses all information about the model as a Python dictionary.
+
+By printing this dictionary we can therefore get a concise summary of the model.
+"""
+model = af.Model(Gaussian)
+
+print(model.dict())
+
+"""
+The dictionary representation printed above can be saved to hard disk as a `.json` file.
+
+This means we can save any multi-level model to hard-disk in a human readable format.
+
+Checkout the file `autofit_workspace/*/cookbooks/jsons/group_level_model.json` to see the model written as a .json.
+"""
+model_path = path.join("scripts", "cookbooks", "jsons")
+
+os.makedirs(model_path, exist_ok=True)
+
+model_file = path.join(model_path, "multi_level_model.json")
+
+with open(model_file, "w+") as f:
+ json.dump(model.dict(), f, indent=4)
+
+"""
+We can load the model from its `.json` file, meaning that one can easily save a model to hard disk and load it
+elsewhere.
+"""
+model = af.Model.from_json(file=model_file)
+
+print(model.info)
+
+"""
+__Wrap Up__
+
+This cookbook shows how to multi-level models consisting of multiple components using the `af.Model()`
+and `af.Collection()` objects.
+
+You should think carefully about whether your model fitting problem can use multi-level models, as they can make
+your model definition more concise and extensible.
+"""
diff --git a/scripts/cookbooks/multiple_datasets.py b/scripts/cookbooks/multiple_datasets.py
index 00650883..634b4479 100644
--- a/scripts/cookbooks/multiple_datasets.py
+++ b/scripts/cookbooks/multiple_datasets.py
@@ -1,482 +1,478 @@
-"""
-Cookbook: Multiple Datasets
-===========================
-
-This cookbook illustrates how to fit multiple datasets simultaneously, where each dataset is fitted by a different
-`Analysis` class.
-
-The `Analysis` classes are combined to give an overall log likelihood function that is the sum of the
-individual log likelihood functions, which a single model is fitted to via non-linear search.
-
-If one has multiple observations of the same signal, it is often desirable to fit them simultaneously. This ensures
-that better constraints are placed on the model, as the full amount of information in the datasets is used.
-
-In some scenarios, the signal may vary across the datasets in a way that requires that the model is updated
-accordingly. **PyAutoFit** provides tools to customize the model composition such that specific parameters of the model
-vary across the datasets.
-
-This cookbook illustrates using observations of 3 1D Gaussians, which have the same `centre` (which is the same
-for the model fitted to each dataset) but different `normalization and `sigma` values (which vary for the model
-fitted to each dataset).
-
-It is common for each individual dataset to only constrain specific aspects of a model. The high level of model
-customization provided by **PyAutoFit** ensures that composing a model that is appropriate for fitting large and diverse
-datasets is straight forward. This is because different `Analysis` classes can be written for each dataset and combined.
-
-__Contents__
-
- - Model Fit: Setup a model-fit to 3 datasets to illustrate multi-dataset fitting.
- - Analysis List: Create a list of `Analysis` objects, one for each dataset, which are fitted simultaneously.
- - Analysis Factor: Wrap each `Analysis` object in an `AnalysisFactor`, which pairs it with the model and prepares it for model fitting.
- - Factor Graph: Combine all `AnalysisFactor` objects into a `FactorGraphModel`, which represents a global model fit to multiple datasets.
- - Result List: Use the output of fits to multiple datasets which are a list of `Result` objects.
- - Variable Model Across Datasets: Fit a model where certain parameters vary across the datasets whereas others
- stay fixed.
- - Relational Model: Fit models where certain parameters vary across the dataset as a user
- defined relation (e.g. `y = mx + c`).
- - Different Analysis Objects: Fit multiple datasets where each dataset is fitted by a different `Analysis` class,
- meaning that datasets with different formats can be fitted simultaneously.
- - Hierarchical / Graphical Models: Use hierarchical / graphical models to fit multiple datasets simultaneously,
- which fit for global trends in the model across the datasets.
- - Interpolation: Fit multiple datasets with a model one-by-one and interpolation over a smoothly varying parameter
- (e.g. time) to infer the model between datasets.
- - Wrap Up: A summary of multi-dataset fitting in PyAutoFit.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import matplotlib.pyplot as plt
-import numpy as np
-from os import path
-
-import autofit as af
-
-"""
-__Model Fit__
-
-Load 3 1D Gaussian datasets from .json files in the directory `autofit_workspace/dataset/`.
-
-All three datasets contain an identical signal, therefore fitting the same model to all three datasets simultaneously
-is appropriate.
-
-Each dataset has a different noise realization, therefore fitting them simultaneously will offer improved constraints
-over individual fits.
-
-__Example Source Code (`af.ex`)__
-
-The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
-
- - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
- `visualize` functions.
-
- - `Gaussian`: a model component representing a 1D Gaussian profile.
-
-These are functionally identical to the `Analysis` and `Gaussian` objects you have seen elsewhere in the workspace.
-"""
-dataset_size = 3
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(path.join("dataset", "example_1d", "gaussian_x1_identical_0")):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data_list = []
-noise_map_list = []
-
-for dataset_index in range(dataset_size):
- dataset_path = path.join(
- "dataset", "example_1d", f"gaussian_x1_identical_{dataset_index}"
- )
-
- data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
- data_list.append(data)
-
- noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
- )
- noise_map_list.append(noise_map)
-
-"""
-Plot all 3 datasets, including their error bars.
-"""
-for data, noise_map in zip(data_list, noise_map_list):
- xvalues = range(data.shape[0])
-
- plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- color="k",
- ecolor="k",
- linestyle="",
- elinewidth=1,
- capsize=2,
- )
- plt.show()
- plt.close()
-
-"""
-Create our model corresponding to a single 1D Gaussian that is fitted to all 3 datasets simultaneously.
-"""
-model = af.Model(af.ex.Gaussian)
-
-model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.normalization = af.LogUniformPrior(lower_limit=1e-2, upper_limit=1e2)
-model.sigma = af.TruncatedGaussianPrior(
- mean=10.0, sigma=5.0, lower_limit=0.0, upper_limit=np.inf
-)
-
-"""
-__Analysis List__
-
-Set up three instances of the `Analysis` class which fit 1D Gaussian.
-"""
-analysis_list = []
-
-for data, noise_map in zip(data_list, noise_map_list):
- analysis = af.ex.Analysis(data=data, noise_map=noise_map)
- analysis_list.append(analysis)
-
-"""
-__Analysis Factor__
-
-Each analysis object is wrapped in an `AnalysisFactor`, which pairs it with the model and prepares it for use in a
-factor graph. This step allows us to flexibly define how each dataset relates to the model.
-
-The term "Factor" comes from factor graphs, a type of probabilistic graphical model. In this context, each factor
-represents the connection between one dataset and the shared model.
-"""
-analysis_factor_list = []
-
-for analysis in analysis_list:
-
- analysis_factor = af.AnalysisFactor(prior_model=model, analysis=analysis)
-
- analysis_factor_list.append(analysis_factor)
-
-"""
-__Factor Graph__
-
-All `AnalysisFactor` objects are combined into a `FactorGraphModel`, which represents a global model fit to
-multiple datasets using a graphical model structure.
-
-The key outcomes of this setup are:
-
- - The individual log likelihoods from each `Analysis` object are summed to form the total log likelihood
- evaluated during the model-fitting process.
-
- - Results from all datasets are output to a unified directory, with subdirectories for visualizations
- from each analysis object, as defined by their `visualize` methods.
-
-This is a basic use of **PyAutoFit**'s graphical modeling capabilities, which support advanced hierarchical
-and probabilistic modeling for large, multi-dataset analyses.
-"""
-factor_graph = af.FactorGraphModel(*analysis_factor_list)
-
-"""
-To inspect the model, we print `factor_graph.global_prior_model.info`.
-"""
-print(factor_graph.global_prior_model.info)
-
-"""
-To fit multiple datasets, we pass the `FactorGraphModel` to a non-linear search.
-
-Unlike single-dataset fitting, we now pass the `factor_graph.global_prior_model` as the model and
-the `factor_graph` itself as the analysis object.
-
-This structure enables simultaneous fitting of multiple datasets in a consistent and scalable way.
-"""
-search = af.DynestyStatic(
- path_prefix="features", sample="rwalk", name="multiple_datasets_simple"
-)
-
-result_list = search.fit(model=factor_graph.global_prior_model, analysis=factor_graph)
-
-"""
-__Result List__
-
-The result object returned by the fit is a list of the `Result` objects, which is described in the result cookbook.
-
-Each `Result` in the list corresponds to each `Analysis` object in the `analysis_list` we passed to the fit.
-
-The same model was fitted across all analyses, thus every `Result` in the `result_list` contains the same information
-on the samples and the same `max_log_likelihood_instance`.
-"""
-print(result_list[0].max_log_likelihood_instance.centre)
-print(result_list[0].max_log_likelihood_instance.normalization)
-print(result_list[0].max_log_likelihood_instance.sigma)
-
-print(result_list[1].max_log_likelihood_instance.centre)
-print(result_list[1].max_log_likelihood_instance.normalization)
-print(result_list[1].max_log_likelihood_instance.sigma)
-
-"""
-We can plot the model-fit to each dataset by iterating over the results:
-"""
-for data, result in zip(data_list, result_list):
- instance = result.max_log_likelihood_instance
-
- model_data = instance.model_data_from(xvalues=np.arange(data.shape[0]))
-
- plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
- )
- plt.plot(xvalues, model_data, color="r")
- plt.title("Dynesty model fit to 1D Gaussian dataset.")
- plt.xlabel("x values of profile")
- plt.ylabel("Profile normalization")
- plt.show()
- plt.close()
-
-"""
-__Variable Model Across Datasets__
-
-The same model was fitted to every dataset simultaneously because all 3 datasets contained an identical signal with
-only the noise varying across the datasets.
-
-If the signal varied across the datasets, we would instead want to fit a different model to each dataset. The model
-composition can be updated by changing the model passed to each `AnalysisFactor`.
-
-We will use an example of 3 1D Gaussians which have the same `centre` but the `normalization` and `sigma` vary across
-datasets:
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1_variable")
-
-dataset_name_list = ["sigma_0", "sigma_1", "sigma_2"]
-
-data_list = []
-noise_map_list = []
-
-for dataset_name in dataset_name_list:
- dataset_time_path = path.join(dataset_path, dataset_name)
-
- data = af.util.numpy_array_from_json(
- file_path=path.join(dataset_time_path, "data.json")
- )
- noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_time_path, "noise_map.json")
- )
-
- data_list.append(data)
- noise_map_list.append(noise_map)
-
-"""
-Plotting these datasets shows that the `normalization` and` `sigma` of each Gaussian vary.
-"""
-for data, noise_map in zip(data_list, noise_map_list):
- xvalues = range(data.shape[0])
-
- af.ex.plot_profile_1d(xvalues=xvalues, profile_1d=data)
-
-"""
-The `centre` of all three 1D Gaussians are the same in each dataset, but their `normalization` and `sigma` values
-are decreasing.
-
-We will therefore fit a model to all three datasets simultaneously, whose `centre` is the same for all 3 datasets but
-the `normalization` and `sigma` vary.
-
-To do that, we use a summed list of `Analysis` objects, where each `Analysis` object contains a different dataset.
-"""
-analysis_list = []
-
-for data, noise_map in zip(data_list, noise_map_list):
- analysis = af.ex.Analysis(data=data, noise_map=noise_map)
- analysis_list.append(analysis)
-
-"""
-We now update the model passed to each `AnalysisFactor `object to compose a model where:
-
- - The `centre` values of the Gaussian fitted to every dataset in every `Analysis` object are identical.
-
- - The`normalization` and `sigma` value of the every Gaussian fitted to every dataset in every `Analysis` object
- are different.
-
-The model has 7 free parameters in total, x1 shared `centre`, x3 unique `normalization`'s and x3 unique `sigma`'s.
-
-We do this by overwriting the `normalization` and `sigma` variables of the model passed to each `AnalysisFactor` object
-with new priors, that make them free parameters of the model.
-"""
-analysis_factor_list = []
-
-for analysis in analysis_list:
-
- model_analysis = model.copy()
-
- model_analysis.normalization = af.LogUniformPrior(lower_limit=1e-2, upper_limit=1e2)
- model_analysis.sigma = af.TruncatedGaussianPrior(
- mean=10.0, sigma=5.0, lower_limit=0.0, upper_limit=np.inf
- )
-
- analysis_factor = af.AnalysisFactor(prior_model=model_analysis, analysis=analysis)
-
- analysis_factor_list.append(analysis_factor)
-
-"""
-To inspect this model, with extra parameters for each dataset created, we print `factor_graph.global_prior_model.info`.
-"""
-factor_graph = af.FactorGraphModel(*analysis_factor_list)
-
-print(factor_graph.global_prior_model.info)
-
-"""
-Fit this model to the data using dynesty.
-"""
-search = af.DynestyStatic(
- path_prefix="features", sample="rwalk", name="multiple_datasets_free_sigma"
-)
-
-result_list = search.fit(model=factor_graph.global_prior_model, analysis=factor_graph)
-
-"""
-The `normalization` and `sigma` values of the maximum log likelihood models fitted to each dataset are different,
-which is shown by printing the `sigma` values of the maximum log likelihood instances of each result.
-
-The `centre` values of the maximum log likelihood models fitted to each dataset are the same.
-"""
-for result in result_list:
- instance = result.max_log_likelihood_instance
-
- print("Max Log Likelihood Model:")
- print("Centre = ", instance.centre)
- print("Normalization = ", instance.normalization)
- print("Sigma = ", instance.sigma)
- print()
-
-
-"""
-__Relational Model__
-
-In the model above, two extra free parameters (`normalization and `sigma`) were added for every dataset.
-
-For just 3 datasets the model stays low dimensional and this is not a problem. However, for 30+ datasets the model
-will become complex and difficult to fit.
-
-In these circumstances, one can instead compose a model where the parameters vary smoothly across the datasets
-via a user defined relation.
-
-Below, we compose a model where the `sigma` value fitted to each dataset is computed according to:
-
- `y = m * x + c` : `sigma` = sigma_m * x + sigma_c`
-
-Where x is an integer number specifying the index of the dataset (e.g. 1, 2 and 3).
-
-By defining a relation of this form, `sigma_m` and `sigma_c` are the only free parameters of the model which vary
-across the datasets.
-
-Of more datasets are added the number of model parameters therefore does not increase.
-"""
-model = af.Collection(gaussian=af.Model(af.ex.Gaussian))
-
-sigma_m = af.UniformPrior(lower_limit=-10.0, upper_limit=10.0)
-sigma_c = af.UniformPrior(lower_limit=-10.0, upper_limit=10.0)
-
-x_list = [1.0, 2.0, 3.0]
-
-analysis_factor_list = []
-
-for x, analysis in zip(x_list, analysis_list):
- sigma_relation = (sigma_m * x) + sigma_c
-
- model_analysis = model.copy()
- model_analysis.gaussian.sigma = sigma_relation
-
- analysis_factor = af.AnalysisFactor(prior_model=model_analysis, analysis=analysis)
-
- analysis_factor_list.append(analysis_factor)
-
-"""
-The factor graph is created and its info can be printed after the relational model has been defined.
-"""
-factor_graph = af.FactorGraphModel(*analysis_factor_list)
-
-print(factor_graph.global_prior_model.info)
-
-"""
-We can fit the model as per usual.
-"""
-search = af.DynestyStatic(
- path_prefix="features", sample="rwalk", name="multiple_datasets_relation"
-)
-
-result_list = search.fit(model=factor_graph.global_prior_model, analysis=factor_graph)
-
-"""
-The `centre` and `sigma` values of the maximum log likelihood models fitted to each dataset are different,
-which is shown by printing the `sigma` values of the maximum log likelihood instances of each result.
-
-They now follow the relation we defined above.
-
-The `centre` normalization of the maximum log likelihood models fitted to each dataset are the same.
-"""
-for result in result_list:
- instance = result.max_log_likelihood_instance
-
- print("Max Log Likelihood Model:")
- print("Centre = ", instance.gaussian.centre)
- print("Normalization = ", instance.gaussian.normalization)
- print("Sigma = ", instance.gaussian.sigma)
- print()
-
-"""
-__Different Analysis Objects__
-
-For simplicity, this example used a single `Analysis` class which fitted 1D Gaussian's to 1D data.
-
-For many problems one may have multiple datasets which are quite different in their format and structure. In this
-situation, one can simply define unique `Analysis` objects for each type of dataset, which will contain a
-unique `log_likelihood_function` and methods for visualization.
-
-__Hierarchical / Graphical Models__
-
-The analysis factor API illustrated here can then be used to fit this large variety of datasets, noting that the
-the model can also be customized as necessary for fitting models to multiple datasets that are different in their
-format and structure.
-
-This allows us to fit large heterogeneous datasets simultaneously, but also forms the basis of the graphical
-modeling API which can be used to fit complex models, such as hierarchical models, to extract more information
-from large datasets.
-
-**PyAutoFit** has a dedicated feature set for fitting hierarchical and graphical models and interested readers should
-checkout the hierarchical and graphical modeling
-chapter of **HowToFit** (https://pyautofit.readthedocs.io/en/latest/howtofit/chapter_graphical_models.html)
-
-__Interpolation__
-
-One may have many datasets which vary according to a smooth function, for example a dataset taken over time where
-the signal varies smoothly as a function of time.
-
-This could be fitted using the tools above, all at once. However, in many use cases this is not possible due to the
-model complexity, number of datasets or computational time.
-
-An alternative approach is to fit each dataset individually, and then interpolate the results over the smoothly
-varying parameter (e.g. time) to estimate the model parameters at any point.
-
-**PyAutoFit** has interpolation tools to do exactly this, which are described in the `features/interpolation.ipynb`
-example.
-
-__Wrap Up__
-
-We have shown how **PyAutoFit** can fit large datasets simultaneously, using custom models that vary specific
-parameters across the dataset.
-"""
+"""
+Cookbook: Multiple Datasets
+===========================
+
+This cookbook illustrates how to fit multiple datasets simultaneously, where each dataset is fitted by a different
+`Analysis` class.
+
+The `Analysis` classes are combined to give an overall log likelihood function that is the sum of the
+individual log likelihood functions, which a single model is fitted to via non-linear search.
+
+If one has multiple observations of the same signal, it is often desirable to fit them simultaneously. This ensures
+that better constraints are placed on the model, as the full amount of information in the datasets is used.
+
+In some scenarios, the signal may vary across the datasets in a way that requires that the model is updated
+accordingly. **PyAutoFit** provides tools to customize the model composition such that specific parameters of the model
+vary across the datasets.
+
+This cookbook illustrates using observations of 3 1D Gaussians, which have the same `centre` (which is the same
+for the model fitted to each dataset) but different `normalization and `sigma` values (which vary for the model
+fitted to each dataset).
+
+It is common for each individual dataset to only constrain specific aspects of a model. The high level of model
+customization provided by **PyAutoFit** ensures that composing a model that is appropriate for fitting large and diverse
+datasets is straight forward. This is because different `Analysis` classes can be written for each dataset and combined.
+
+__Contents__
+
+ - Model Fit: Setup a model-fit to 3 datasets to illustrate multi-dataset fitting.
+ - Analysis List: Create a list of `Analysis` objects, one for each dataset, which are fitted simultaneously.
+ - Analysis Factor: Wrap each `Analysis` object in an `AnalysisFactor`, which pairs it with the model and prepares it for model fitting.
+ - Factor Graph: Combine all `AnalysisFactor` objects into a `FactorGraphModel`, which represents a global model fit to multiple datasets.
+ - Result List: Use the output of fits to multiple datasets which are a list of `Result` objects.
+ - Variable Model Across Datasets: Fit a model where certain parameters vary across the datasets whereas others
+ stay fixed.
+ - Relational Model: Fit models where certain parameters vary across the dataset as a user
+ defined relation (e.g. `y = mx + c`).
+ - Different Analysis Objects: Fit multiple datasets where each dataset is fitted by a different `Analysis` class,
+ meaning that datasets with different formats can be fitted simultaneously.
+ - Hierarchical / Graphical Models: Use hierarchical / graphical models to fit multiple datasets simultaneously,
+ which fit for global trends in the model across the datasets.
+ - Interpolation: Fit multiple datasets with a model one-by-one and interpolation over a smoothly varying parameter
+ (e.g. time) to infer the model between datasets.
+ - Wrap Up: A summary of multi-dataset fitting in PyAutoFit.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import matplotlib.pyplot as plt
+import numpy as np
+from os import path
+
+import autofit as af
+
+"""
+__Model Fit__
+
+Load 3 1D Gaussian datasets from .json files in the directory `autofit_workspace/dataset/`.
+
+All three datasets contain an identical signal, therefore fitting the same model to all three datasets simultaneously
+is appropriate.
+
+Each dataset has a different noise realization, therefore fitting them simultaneously will offer improved constraints
+over individual fits.
+
+__Example Source Code (`af.ex`)__
+
+The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
+
+ - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
+ `visualize` functions.
+
+ - `Gaussian`: a model component representing a 1D Gaussian profile.
+
+These are functionally identical to the `Analysis` and `Gaussian` objects you have seen elsewhere in the workspace.
+"""
+dataset_size = 3
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(path.join("dataset", "example_1d", "gaussian_x1_identical_0")):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data_list = []
+noise_map_list = []
+
+for dataset_index in range(dataset_size):
+ dataset_path = path.join(
+ "dataset", "example_1d", f"gaussian_x1_identical_{dataset_index}"
+ )
+
+ data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+ data_list.append(data)
+
+ noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+ )
+ noise_map_list.append(noise_map)
+
+"""
+Plot all 3 datasets, including their error bars.
+"""
+for data, noise_map in zip(data_list, noise_map_list):
+ xvalues = range(data.shape[0])
+
+ plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ color="k",
+ ecolor="k",
+ linestyle="",
+ elinewidth=1,
+ capsize=2,
+ )
+ plt.show()
+ plt.close()
+
+"""
+Create our model corresponding to a single 1D Gaussian that is fitted to all 3 datasets simultaneously.
+"""
+model = af.Model(af.ex.Gaussian)
+
+model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.normalization = af.LogUniformPrior(lower_limit=1e-2, upper_limit=1e2)
+model.sigma = af.TruncatedGaussianPrior(
+ mean=10.0, sigma=5.0, lower_limit=0.0, upper_limit=np.inf
+)
+
+"""
+__Analysis List__
+
+Set up three instances of the `Analysis` class which fit 1D Gaussian.
+"""
+analysis_list = []
+
+for data, noise_map in zip(data_list, noise_map_list):
+ analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+ analysis_list.append(analysis)
+
+"""
+__Analysis Factor__
+
+Each analysis object is wrapped in an `AnalysisFactor`, which pairs it with the model and prepares it for use in a
+factor graph. This step allows us to flexibly define how each dataset relates to the model.
+
+The term "Factor" comes from factor graphs, a type of probabilistic graphical model. In this context, each factor
+represents the connection between one dataset and the shared model.
+"""
+analysis_factor_list = []
+
+for analysis in analysis_list:
+
+ analysis_factor = af.AnalysisFactor(prior_model=model, analysis=analysis)
+
+ analysis_factor_list.append(analysis_factor)
+
+"""
+__Factor Graph__
+
+All `AnalysisFactor` objects are combined into a `FactorGraphModel`, which represents a global model fit to
+multiple datasets using a graphical model structure.
+
+The key outcomes of this setup are:
+
+ - The individual log likelihoods from each `Analysis` object are summed to form the total log likelihood
+ evaluated during the model-fitting process.
+
+ - Results from all datasets are output to a unified directory, with subdirectories for visualizations
+ from each analysis object, as defined by their `visualize` methods.
+
+This is a basic use of **PyAutoFit**'s graphical modeling capabilities, which support advanced hierarchical
+and probabilistic modeling for large, multi-dataset analyses.
+"""
+factor_graph = af.FactorGraphModel(*analysis_factor_list)
+
+"""
+To inspect the model, we print `factor_graph.global_prior_model.info`.
+"""
+print(factor_graph.global_prior_model.info)
+
+"""
+To fit multiple datasets, we pass the `FactorGraphModel` to a non-linear search.
+
+Unlike single-dataset fitting, we now pass the `factor_graph.global_prior_model` as the model and
+the `factor_graph` itself as the analysis object.
+
+This structure enables simultaneous fitting of multiple datasets in a consistent and scalable way.
+"""
+search = af.DynestyStatic(
+ path_prefix="features", sample="rwalk", name="multiple_datasets_simple"
+)
+
+result_list = search.fit(model=factor_graph.global_prior_model, analysis=factor_graph)
+
+"""
+__Result List__
+
+The result object returned by the fit is a list of the `Result` objects, which is described in the result cookbook.
+
+Each `Result` in the list corresponds to each `Analysis` object in the `analysis_list` we passed to the fit.
+
+The same model was fitted across all analyses, thus every `Result` in the `result_list` contains the same information
+on the samples and the same `max_log_likelihood_instance`.
+"""
+print(result_list[0].max_log_likelihood_instance.centre)
+print(result_list[0].max_log_likelihood_instance.normalization)
+print(result_list[0].max_log_likelihood_instance.sigma)
+
+print(result_list[1].max_log_likelihood_instance.centre)
+print(result_list[1].max_log_likelihood_instance.normalization)
+print(result_list[1].max_log_likelihood_instance.sigma)
+
+"""
+We can plot the model-fit to each dataset by iterating over the results:
+"""
+for data, result in zip(data_list, result_list):
+ instance = result.max_log_likelihood_instance
+
+ model_data = instance.model_data_from(xvalues=np.arange(data.shape[0]))
+
+ plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+ )
+ plt.plot(xvalues, model_data, color="r")
+ plt.title("Dynesty model fit to 1D Gaussian dataset.")
+ plt.xlabel("x values of profile")
+ plt.ylabel("Profile normalization")
+ plt.show()
+ plt.close()
+
+"""
+__Variable Model Across Datasets__
+
+The same model was fitted to every dataset simultaneously because all 3 datasets contained an identical signal with
+only the noise varying across the datasets.
+
+If the signal varied across the datasets, we would instead want to fit a different model to each dataset. The model
+composition can be updated by changing the model passed to each `AnalysisFactor`.
+
+We will use an example of 3 1D Gaussians which have the same `centre` but the `normalization` and `sigma` vary across
+datasets:
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1_variable")
+
+dataset_name_list = ["sigma_0", "sigma_1", "sigma_2"]
+
+data_list = []
+noise_map_list = []
+
+for dataset_name in dataset_name_list:
+ dataset_time_path = path.join(dataset_path, dataset_name)
+
+ data = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_time_path, "data.json")
+ )
+ noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_time_path, "noise_map.json")
+ )
+
+ data_list.append(data)
+ noise_map_list.append(noise_map)
+
+"""
+Plotting these datasets shows that the `normalization` and` `sigma` of each Gaussian vary.
+"""
+for data, noise_map in zip(data_list, noise_map_list):
+ xvalues = range(data.shape[0])
+
+ af.ex.plot_profile_1d(xvalues=xvalues, profile_1d=data)
+
+"""
+The `centre` of all three 1D Gaussians are the same in each dataset, but their `normalization` and `sigma` values
+are decreasing.
+
+We will therefore fit a model to all three datasets simultaneously, whose `centre` is the same for all 3 datasets but
+the `normalization` and `sigma` vary.
+
+To do that, we use a summed list of `Analysis` objects, where each `Analysis` object contains a different dataset.
+"""
+analysis_list = []
+
+for data, noise_map in zip(data_list, noise_map_list):
+ analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+ analysis_list.append(analysis)
+
+"""
+We now update the model passed to each `AnalysisFactor `object to compose a model where:
+
+ - The `centre` values of the Gaussian fitted to every dataset in every `Analysis` object are identical.
+
+ - The`normalization` and `sigma` value of the every Gaussian fitted to every dataset in every `Analysis` object
+ are different.
+
+The model has 7 free parameters in total, x1 shared `centre`, x3 unique `normalization`'s and x3 unique `sigma`'s.
+
+We do this by overwriting the `normalization` and `sigma` variables of the model passed to each `AnalysisFactor` object
+with new priors, that make them free parameters of the model.
+"""
+analysis_factor_list = []
+
+for analysis in analysis_list:
+
+ model_analysis = model.copy()
+
+ model_analysis.normalization = af.LogUniformPrior(lower_limit=1e-2, upper_limit=1e2)
+ model_analysis.sigma = af.TruncatedGaussianPrior(
+ mean=10.0, sigma=5.0, lower_limit=0.0, upper_limit=np.inf
+ )
+
+ analysis_factor = af.AnalysisFactor(prior_model=model_analysis, analysis=analysis)
+
+ analysis_factor_list.append(analysis_factor)
+
+"""
+To inspect this model, with extra parameters for each dataset created, we print `factor_graph.global_prior_model.info`.
+"""
+factor_graph = af.FactorGraphModel(*analysis_factor_list)
+
+print(factor_graph.global_prior_model.info)
+
+"""
+Fit this model to the data using dynesty.
+"""
+search = af.DynestyStatic(
+ path_prefix="features", sample="rwalk", name="multiple_datasets_free_sigma"
+)
+
+result_list = search.fit(model=factor_graph.global_prior_model, analysis=factor_graph)
+
+"""
+The `normalization` and `sigma` values of the maximum log likelihood models fitted to each dataset are different,
+which is shown by printing the `sigma` values of the maximum log likelihood instances of each result.
+
+The `centre` values of the maximum log likelihood models fitted to each dataset are the same.
+"""
+for result in result_list:
+ instance = result.max_log_likelihood_instance
+
+ print("Max Log Likelihood Model:")
+ print("Centre = ", instance.centre)
+ print("Normalization = ", instance.normalization)
+ print("Sigma = ", instance.sigma)
+ print()
+
+
+"""
+__Relational Model__
+
+In the model above, two extra free parameters (`normalization and `sigma`) were added for every dataset.
+
+For just 3 datasets the model stays low dimensional and this is not a problem. However, for 30+ datasets the model
+will become complex and difficult to fit.
+
+In these circumstances, one can instead compose a model where the parameters vary smoothly across the datasets
+via a user defined relation.
+
+Below, we compose a model where the `sigma` value fitted to each dataset is computed according to:
+
+ `y = m * x + c` : `sigma` = sigma_m * x + sigma_c`
+
+Where x is an integer number specifying the index of the dataset (e.g. 1, 2 and 3).
+
+By defining a relation of this form, `sigma_m` and `sigma_c` are the only free parameters of the model which vary
+across the datasets.
+
+Of more datasets are added the number of model parameters therefore does not increase.
+"""
+model = af.Collection(gaussian=af.Model(af.ex.Gaussian))
+
+sigma_m = af.UniformPrior(lower_limit=-10.0, upper_limit=10.0)
+sigma_c = af.UniformPrior(lower_limit=-10.0, upper_limit=10.0)
+
+x_list = [1.0, 2.0, 3.0]
+
+analysis_factor_list = []
+
+for x, analysis in zip(x_list, analysis_list):
+ sigma_relation = (sigma_m * x) + sigma_c
+
+ model_analysis = model.copy()
+ model_analysis.gaussian.sigma = sigma_relation
+
+ analysis_factor = af.AnalysisFactor(prior_model=model_analysis, analysis=analysis)
+
+ analysis_factor_list.append(analysis_factor)
+
+"""
+The factor graph is created and its info can be printed after the relational model has been defined.
+"""
+factor_graph = af.FactorGraphModel(*analysis_factor_list)
+
+print(factor_graph.global_prior_model.info)
+
+"""
+We can fit the model as per usual.
+"""
+search = af.DynestyStatic(
+ path_prefix="features", sample="rwalk", name="multiple_datasets_relation"
+)
+
+result_list = search.fit(model=factor_graph.global_prior_model, analysis=factor_graph)
+
+"""
+The `centre` and `sigma` values of the maximum log likelihood models fitted to each dataset are different,
+which is shown by printing the `sigma` values of the maximum log likelihood instances of each result.
+
+They now follow the relation we defined above.
+
+The `centre` normalization of the maximum log likelihood models fitted to each dataset are the same.
+"""
+for result in result_list:
+ instance = result.max_log_likelihood_instance
+
+ print("Max Log Likelihood Model:")
+ print("Centre = ", instance.gaussian.centre)
+ print("Normalization = ", instance.gaussian.normalization)
+ print("Sigma = ", instance.gaussian.sigma)
+ print()
+
+"""
+__Different Analysis Objects__
+
+For simplicity, this example used a single `Analysis` class which fitted 1D Gaussian's to 1D data.
+
+For many problems one may have multiple datasets which are quite different in their format and structure. In this
+situation, one can simply define unique `Analysis` objects for each type of dataset, which will contain a
+unique `log_likelihood_function` and methods for visualization.
+
+__Hierarchical / Graphical Models__
+
+The analysis factor API illustrated here can then be used to fit this large variety of datasets, noting that the
+the model can also be customized as necessary for fitting models to multiple datasets that are different in their
+format and structure.
+
+This allows us to fit large heterogeneous datasets simultaneously, but also forms the basis of the graphical
+modeling API which can be used to fit complex models, such as hierarchical models, to extract more information
+from large datasets.
+
+**PyAutoFit** has a dedicated feature set for fitting hierarchical and graphical models and interested readers should
+checkout the hierarchical and graphical modeling
+chapter of **HowToFit** (https://pyautofit.readthedocs.io/en/latest/howtofit/chapter_graphical_models.html)
+
+__Interpolation__
+
+One may have many datasets which vary according to a smooth function, for example a dataset taken over time where
+the signal varies smoothly as a function of time.
+
+This could be fitted using the tools above, all at once. However, in many use cases this is not possible due to the
+model complexity, number of datasets or computational time.
+
+An alternative approach is to fit each dataset individually, and then interpolate the results over the smoothly
+varying parameter (e.g. time) to estimate the model parameters at any point.
+
+**PyAutoFit** has interpolation tools to do exactly this, which are described in the `features/interpolation.ipynb`
+example.
+
+__Wrap Up__
+
+We have shown how **PyAutoFit** can fit large datasets simultaneously, using custom models that vary specific
+parameters across the dataset.
+"""
diff --git a/scripts/cookbooks/result.py b/scripts/cookbooks/result.py
index de877541..f409bacb 100644
--- a/scripts/cookbooks/result.py
+++ b/scripts/cookbooks/result.py
@@ -1,751 +1,747 @@
-"""
-Cookbook: Result
-================
-
-A non-linear search fits a model to a dataset, returning a `Result` object that contains a lot of information on the
-model-fit.
-
-This cookbook provides a concise reference to the result API.
-
-The cookbook then describes how the results of a search can be output to hard-disk and loaded back into Python,
-either using the `Aggregator` object or by building an sqlite database of results. Result loading supports
-queries, so that only the results of interest are returned.
-
-The samples of the non-linear search, which are used to estimate quantities the maximum likelihood model and
-parameter errors, are described separately in the `samples.py` cookbook.
-
-__Contents__
-
-An overview of the `Result` object's functionality is given in the following sections:
-
- - Simple Fit: Perform a simple model-fit to generate a `Result` object.
- - Info: Print the `info` attribute of the `Result` object to display a summary of the model-fit.
- - Max Log Likelihood Instance: Getting the maximum likelihood model instance.
- - Samples: Getting the samples of the non-linear search from a result.
- - Custom Result: Extending the `Result` object with custom attributes specific to the model-fit.
-
-The cookbook next describes how results can be output to hard-disk and loaded back into Python via the `Aggregator`:
-
- - Output To Hard-Disk: Output results to hard-disk so they can be inspected and used to restart a crashed search.
- - Files: The files that are stored in the `files` folder that is created when results are output to hard-disk.
- - Loading From Hard-disk: Loading results from hard-disk to Python variables via the aggregator.
- - Generators: Why loading results uses Python generators to ensure memory efficiency.
-
-The cookbook next gives examples of how to load all the following results from the database:
-
- - Samples: The samples of the non-linear search loaded via the aggregator.
- - Loading Model: The model fitted by the non-linear search.
- - Loading Search: The search used to perform the model-fit.
- - Loading Samples: The samples of the non-linear search (e.g. all parameter values, log likelihoods, etc.).
- - Loading Samples Info: Additional information on the samples.
- - Loading Samples Summary: A summary of the samples of the non-linear search (e.g. the maximum log likelihood model).
- - Loading Info: The `info` dictionary passed to the search.
-
-The output of results to hard-disk is customizeable and described in the following section:
-
- - Custom Output: Extend `Analysis` classes to output additional information which can be loaded via the aggregator.
-
-Using queries to load specific results is described in the following sections:
-
- - Querying Datasets: Query based on the name of the dataset.
- - Querying Searches: Query based on the name of the search.
- - Querying Models: Query based on the model that is fitted.
- - Querying Results: Query based on the results of the model-fit.
- - Querying with Logic: Use logic to combine queries to load specific results (e.g. AND, OR, etc.).
-
-The final section describes how to use results built in an sqlite database file:
-
- - Database: Building a database file from the output folder.
- - Unique Identifiers: The unique identifier of each model-fit.
- - Building From Output Folder: Build the database from the output folder on hard-disk.
- - Writing Directly To Database: Writing results directly to the database.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import json
-from os import path
-import numpy as np
-from typing import Optional
-
-import autofit as af
-
-"""
-__Simple Fit__
-
-To illustrate the API of the result object, we first fit a 1D `Gaussian` profile with a `Gaussian` model.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-model = af.Model(af.ex.Gaussian)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-search = af.Emcee(
- nwalkers=30,
- nsteps=1000,
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-"""
-__Info__
-
-Printing the `info` attribute shows the overall result of the model-fit in a human readable format.
-"""
-print(result.info)
-
-"""
-__Max Log Likelihood Instance__
-
-The `max_log_likelihood_instance` is the model instance of the maximum log likelihood model, which is the model
-that maximizes the likelihood of the data given the model.
-"""
-instance = result.max_log_likelihood_instance
-
-print("Max Log Likelihood `Gaussian` Instance:")
-print("Centre = ", instance.centre)
-print("Normalization = ", instance.normalization)
-print("Sigma = ", instance.sigma)
-
-"""
-__Samples__
-
-The `Samples` class contains all information on the non-linear search samples, for example the value of every parameter
-sampled using the fit or an instance of the maximum likelihood model.
-"""
-print(result.samples)
-
-"""
-The samples are described in detail separately in the `samples.py` cookbook.
-
-__Custom Result__
-
-The result can be can be customized to include additional information about the model-fit that is specific to your
-model-fitting problem.
-
-For example, for fitting 1D profiles, the `Result` could include the maximum log likelihood model 1D data:
-
-`print(result.max_log_likelihood_model_data_1d)`
-
-In other examples, this quantity has been manually computed after the model-fit has completed.
-
-The custom result API allows us to do this. First, we define a custom `Result` class, which includes the property
-`max_log_likelihood_model_data_1d`.
-"""
-
-
-class ResultExample(af.Result):
- @property
- def max_log_likelihood_model_data_1d(self) -> np.ndarray:
- """
- Returns the maximum log likelihood model's 1D model data.
-
- This is an example of how we can pass the `Analysis` class a custom `Result` object and extend this result
- object with new properties that are specific to the model-fit we are performing.
- """
- xvalues = np.arange(self.analysis.data.shape[0])
-
- return self.instance.model_data_from(xvalues=xvalues)
-
-
-"""
-The custom result has access to the analysis class, meaning that we can use any of its methods or properties to
-compute custom result properties.
-
-To make it so that the `ResultExample` object above is returned by the search we overwrite the `Result` class attribute
-of the `Analysis` and define a `make_result` object describing what we want it to contain:
-"""
-
-
-class Analysis(af.ex.Analysis):
- """
- This overwrite means the `ResultExample` class is returned after the model-fit.
- """
-
- Result = ResultExample
-
- def make_result(
- self,
- samples_summary: af.SamplesSummary,
- paths: af.AbstractPaths,
- samples: Optional[af.SamplesPDF] = None,
- search_internal: Optional[object] = None,
- analysis: Optional[object] = None,
- ) -> Result:
- """
- Returns the `Result` of the non-linear search after it is completed.
-
- The result type is defined as a class variable in the `Analysis` class (see top of code under the python code
- `class Analysis(af.Analysis)`.
-
- The result can be manually overwritten by a user to return a user-defined result object, which can be extended
- with additional methods and attribute specific to the model-fit.
-
- This example class does example this, whereby the analysis result has been overwritten with the `ResultExample`
- class, which contains a property `max_log_likelihood_model_data_1d` that returns the model data of the
- best-fit model. This API means you can customize your result object to include whatever attributes you want
- and therefore make a result object specific to your model-fit and model-fitting problem.
-
- The `Result` object you return can be customized to include:
-
- - The samples summary, which contains the maximum log likelihood instance and median PDF model.
-
- - The paths of the search, which are used for loading the samples and search internal below when a search
- is resumed.
-
- - The samples of the non-linear search (e.g. MCMC chains) also stored in `samples.csv`.
-
- - The non-linear search used for the fit in its internal representation, which is used for resuming a search
- and making bespoke visualization using the search's internal results.
-
- - The analysis used to fit the model (default disabled to save memory, but option may be useful for certain
- projects).
-
- Parameters
- ----------
- samples_summary
- The summary of the samples of the non-linear search, which include the maximum log likelihood instance and
- median PDF model.
- paths
- An object describing the paths for saving data (e.g. hard-disk directories or entries in sqlite database).
- samples
- The samples of the non-linear search, for example the chains of an MCMC run.
- search_internal
- The internal representation of the non-linear search used to perform the model-fit.
- analysis
- The analysis used to fit the model.
-
- Returns
- -------
- Result
- The result of the non-linear search, which is defined as a class variable in the `Analysis` class.
- """
- return self.Result(
- samples_summary=samples_summary,
- paths=paths,
- samples=samples,
- search_internal=search_internal,
- analysis=self,
- )
-
-
-"""
-Using the `Analysis` class above, the `Result` object returned by the search is now a `ResultExample` object.
-"""
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-search = af.Emcee(
- nwalkers=30,
- nsteps=1000,
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-print(result.max_log_likelihood_model_data_1d)
-
-"""
-__Output To Hard-Disk__
-
-By default, a non-linear search does not output its results to hard-disk and its results can only be inspected
-in Python via the `result` object.
-
-However, the results of any non-linear search can be output to hard-disk by passing the `name` and / or `path_prefix`
-attributes, which are used to name files and output the results to a folder on your hard-disk.
-
-This cookbook now runs the three searches with output to hard-disk enabled, so you can see how the results are output
-to hard-disk and to then illustrate how they can be loaded back into Python.
-
-Note that an `info` dictionary is also passed to the search, which includes the date of the model-fit and the exposure
-time of the dataset. This information is stored output to hard-disk and can be loaded to help interpret the results.
-"""
-info = {"date_of_observation": "01-02-18", "exposure_time": 1000.0}
-
-dataset_name_list = ["gaussian_x1_0", "gaussian_x1_1", "gaussian_x1_2"]
-
-model = af.Collection(gaussian=af.ex.Gaussian)
-
-model.gaussian.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.gaussian.normalization = af.LogUniformPrior(lower_limit=1e-2, upper_limit=1e2)
-model.gaussian.sigma = af.TruncatedGaussianPrior(
- mean=10.0, sigma=5.0, lower_limit=0.0, upper_limit=np.inf
-)
-
-for dataset_name in dataset_name_list:
- dataset_path = path.join("dataset", "example_1d", dataset_name)
-
- data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
- noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
- )
-
- analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
- search = af.DynestyStatic(
- name="multi_result_example",
- path_prefix=path.join("cookbooks", "result"),
- unique_tag=dataset_name, # This makes the unique identifier use the dataset name
- nlive=50,
- )
-
- print(
- """
- The non-linear search has begun running.
- This Jupyter notebook cell with progress once search has completed, this could take a few minutes!
- """
- )
-
- result = search.fit(model=model, analysis=analysis, info=info)
-
-print("Search has finished run - you may now continue the notebook.")
-
-"""
-__Files__
-
-By outputting results to hard-disk, a `files` folder is created containing .json / .csv files of the model,
-samples, search, etc, for each fit.
-
-You should check it out now for the completed fits on your hard-disk.
-
-A description of all files is as follows:
-
- - `model`: The `model` defined above and used in the model-fit (`model.json`).
- - `search`: The non-linear search settings (`search.json`).
- - `samples`: The non-linear search samples (`samples.csv`).
- - `samples_info`: Additional information about the samples (`samples_info.json`).
- - `samples_summary`: A summary of key results of the samples (`samples_summary.json`).
- - `info`: The info dictionary passed to the search (`info.json`).
- - `covariance`: The inferred covariance matrix (`covariance.csv`).
- - `data`: The 1D noisy data used that is fitted (`data.json`).
- - `noise_map`: The 1D noise-map fitted (`noise_map.json`).
-
-The `samples` and `samples_summary` results contain a lot of repeated information. The `samples` result contains
-the full non-linear search samples, for example every parameter sample and its log likelihood. The `samples_summary`
-contains a summary of the results, for example the maximum log likelihood model and error estimates on parameters
-at 1 and 3 sigma confidence.
-
-Accessing results via the `samples_summary` is much faster, because as it does not reperform calculations using the full
-list of samples. Therefore, if the result you want is accessible via the `samples_summary` you should use it
-but if not you can revert to the `samples.
-
-__Loading From Hard-disk__
-
-The multi-fits above wrote the results to hard-disk in three distinct folders, one for each dataset.
-
-Their results are loaded using the `Aggregator` object, which finds the results in the output directory and can
-load them into Python objects.
-"""
-from autofit.aggregator.aggregator import Aggregator
-
-agg = Aggregator.from_directory(
- directory=path.join("multi_result_example"),
-)
-
-"""
-__Generators__
-
-Before using the aggregator to inspect results, lets discuss Python generators.
-
-A generator is an object that iterates over a function when it is called. The aggregator creates all of the objects
-that it loads from the database as generators (as opposed to a list, or dictionary, or another Python type).
-
-This is because generators are memory efficient, as they do not store the entries of the database in memory
-simultaneously. This contrasts objects like lists and dictionaries, which store all entries in memory all at once.
-If you fit a large number of datasets, lists and dictionaries will use a lot of memory and could crash your computer!
-
-Once we use a generator in the Python code, it cannot be used again. To perform the same task twice, the
-generator must be remade it. This cookbook therefore rarely stores generators as variables and instead uses the
-aggregator to create each generator at the point of use.
-
-To create a generator of a specific set of results, we use the `values` method. This takes the `name` of the
-object we want to create a generator of, for example inputting `name=samples` will return the results `Samples`
-object (which is illustrated in detail below).
-"""
-for samples in agg.values("samples"):
- print(samples.parameter_lists[0])
-
-"""
-__Samples__
-
-By converting this generator to a list and printing it, it is a list of 3 `SamplesNest` objects, corresponding to
-the 3 model-fits performed above.
-"""
-print("Samples:\n")
-samples_gen = agg.values("samples")
-print(samples_gen)
-print("Total Samples Objects = ", len(agg), "\n")
-
-"""
-__Loading Model__
-
-The model used to perform the model fit for each of the 3 datasets can be loaded via the aggregator and printed.
-"""
-model_gen = agg.values("model")
-
-for model in model_gen:
- print(model.info)
-
-"""
-__Loading Search__
-
-The non-linear search used to perform the model fit can be loaded via the aggregator and printed.
-"""
-search_gen = agg.values("search")
-
-for search in search_gen:
- print(search)
-
-"""
-__Loading Samples__
-
-The `Samples` class contains all information on the non-linear search samples, for example the value of every parameter
-sampled using the fit or an instance of the maximum likelihood model.
-
-The `Samples` class is described fully in the results cookbook.
-"""
-for samples in agg.values("samples"):
- print("The tenth sample`s third parameter")
- print(samples.parameter_lists[9][2], "\n")
-
- instance = samples.max_log_likelihood()
-
- print("Max Log Likelihood `Gaussian` Instance:")
- print("Centre = ", instance.gaussian.centre)
- print("Normalization = ", instance.gaussian.normalization)
- print("Sigma = ", instance.gaussian.sigma, "\n")
-
-"""
-__Loading Samples Info__
-
-The samples info contains additional information on the samples, which depends on the non-linear search used.
-
-For example, for a nested sampling algorithm it contains information on the number of live points, for a MCMC
-algorithm it contains information on the number of steps, etc.
-"""
-for samples_info in agg.values("samples_info"):
- print(samples_info)
-
-"""
-__Loading Samples Summary__
-
-The samples summary contains a subset of results access via the `Samples`, for example the maximum likelihood model
-and parameter error estimates.
-
-Using the samples method above can be slow, as the quantities have to be computed from all non-linear search samples
-(e.g. computing errors requires that all samples are marginalized over). This information is stored directly in the
-samples summary and can therefore be accessed instantly.
-"""
-# for samples_summary in agg.values("samples_summary"):
-#
-# instance = samples_summary.max_log_likelihood()
-#
-# print("Max Log Likelihood `Gaussian` Instance:")
-# print("Centre = ", instance.centre)
-# print("Normalization = ", instance.normalization)
-# print("Sigma = ", instance.sigma, "\n")
-
-"""
-__Loading Info__
-
-The info dictionary passed to the search, discussed earlier in this cookbook, is accessible.
-"""
-for info in agg.values("info"):
- print(info["date_of_observation"])
- print(info["exposure_time"])
-
-"""
-__Custom Output__
-
-The results accessible via the database (e.g. `model`, `samples`) are those contained in the `files` folder.
-
-By extending an `Analysis` class with the methods `save_attributes` and `save_results`,
-custom files can be written to the `files` folder and become accessible via the database.
-
-To save the objects in a human readable and loaded .json format, the `data` and `noise_map`, which are natively stored
-as 1D numpy arrays, are converted to a suitable dictionary output format. This uses the **PyAutoConf** method
-`to_dict`.
-"""
-
-
-class Analysis(af.Analysis):
- def __init__(self, data: np.ndarray, noise_map: np.ndarray):
- """
- Standard Analysis class example used throughout PyAutoFit examples.
- """
- super().__init__()
-
- self.data = data
- self.noise_map = noise_map
-
- def log_likelihood_function(self, instance) -> float:
- """
- Standard log likelihood function used throughout PyAutoFit examples.
- """
-
- xvalues = np.arange(self.data.shape[0])
-
- model_data = instance.model_data_from(xvalues=xvalues)
-
- residual_map = self.data - model_data
- chi_squared_map = (residual_map / self.noise_map) ** 2.0
- chi_squared = sum(chi_squared_map)
- noise_normalization = np.sum(np.log(2 * np.pi * self.noise_map**2.0))
- log_likelihood = -0.5 * (chi_squared + noise_normalization)
-
- return log_likelihood
-
- def save_attributes(self, paths: af.DirectoryPaths):
- """
- Before the non-linear search begins, this routine saves attributes of the `Analysis` object to the `files`
- folder such that they can be loaded after the analysis using PyAutoFit's database and aggregator tools.
-
- For this analysis, it uses the `AnalysisDataset` object's method to output the following:
-
- - The dataset's data as a .json file.
- - The dataset's noise-map as a .json file.
-
- These are accessed using the aggregator via `agg.values("data")` and `agg.values("noise_map")`.
-
- They are saved using the paths function `save_json`, noting that this saves outputs appropriate for the
- sqlite3 database.
-
- Parameters
- ----------
- paths
- The paths object which manages all paths, e.g. where the non-linear search outputs are stored,
- visualization, and the pickled objects used by the aggregator output by this function.
- """
- from autoconf.dictable import to_dict
-
- paths.save_json(name="data", object_dict=to_dict(self.data))
- paths.save_json(name="noise_map", object_dict=to_dict(self.noise_map))
-
- def save_results(self, paths: af.DirectoryPaths, result: af.Result):
- """
- At the end of a model-fit, this routine saves attributes of the `Analysis` object to the `files`
- folder such that they can be loaded after the analysis using PyAutoFit's database and aggregator tools.
-
- For this analysis it outputs the following:
-
- - The maximum log likelihood model data as a .json file.
-
- This is accessed using the aggregator via `agg.values("model_data")`.
-
- Parameters
- ----------
- paths
- The paths object which manages all paths, e.g. where the non-linear search outputs are stored,
- visualization and the pickled objects used by the aggregator output by this function.
- result
- The result of a model fit, including the non-linear search, samples and maximum likelihood model.
- """
- xvalues = np.arange(self.data.shape[0])
-
- instance = result.max_log_likelihood_instance
-
- model_data = instance.model_data_from(xvalues=xvalues)
-
- # The path where model_data.json is saved, e.g. output/dataset_name/unique_id/files/model_data.json
-
- file_path = (path.join(paths._json_path, "model_data.json"),)
-
- with open(file_path, "w+") as f:
- json.dump(model_data, f, indent=4)
-
-
-"""
-__Querying Datasets__
-
-The aggregator can query the database, returning only specific fits of interested.
-
-We can query using the `dataset_name` string we input into the model-fit above, in order to get the results
-of a fit to a specific dataset.
-
-For example, querying using the string `gaussian_x1_1` returns results for only the fit using the
-second `Gaussian` dataset.
-"""
-unique_tag = agg.search.unique_tag
-agg_query = agg.query(unique_tag == "gaussian_x1_1")
-
-"""
-As expected, this list has only 1 `SamplesNest` corresponding to the second dataset.
-"""
-print(agg_query.values("samples"))
-print("Total Samples Objects via dataset_name Query = ", len(agg_query), "\n")
-
-"""
-If we query using an incorrect dataset name we get no results.
-"""
-unique_tag = agg.search.unique_tag
-agg_query = agg.query(unique_tag == "incorrect_name")
-samples_gen = agg_query.values("samples")
-
-"""
-__Querying Searches__
-
-We can query using the `name` of the non-linear search used to fit the model.
-
-In this cookbook, all three fits used the same search, named `database_example`. Query based on search name in this
-example is therefore somewhat pointless.
-
-However, querying based on the search name is useful for model-fits which use a range of searches, for example
-if different non-linear searches are used multiple times.
-
-As expected, the query using search name below contains all 3 results.
-"""
-name = agg.search.name
-agg_query = agg.query(name == "database_example")
-
-print(agg_query.values("samples"))
-print("Total Samples Objects via name Query = ", len(agg_query), "\n")
-
-"""
-__Querying Models__
-
-We can query based on the model fitted.
-
-For example, we can load all results which fitted a `Gaussian` model-component, which in this simple example is all
-3 model-fits.
-
-Querying via the model is useful for loading results after performing many model-fits with many different model
-parameterizations to large (e.g. Bayesian model comparison).
-
-[Note: the code `agg.model.gaussian` corresponds to the fact that in the `Collection` above, we named the model
-component `gaussian`. If this `Collection` had used a different name the code below would change
-correspondingly. Models with multiple model components (e.g., `gaussian` and `exponential`) are therefore also easily
-accessed via the database.]
-"""
-gaussian = agg.model.gaussian
-agg_query = agg.query(gaussian == af.ex.Gaussian)
-print("Total Samples Objects via `Gaussian` model query = ", len(agg_query), "\n")
-
-"""
-__Querying Results__
-
-We can query based on the results of the model-fit.
-
-Below, we query the database to find all fits where the inferred value of `sigma` for the `Gaussian` is less
-than 3.0 (which returns only the first of the three model-fits).
-"""
-gaussian = agg.model.gaussian
-agg_query = agg.query(gaussian.sigma < 3.0)
-print("Total Samples Objects In Query `gaussian.sigma < 3.0` = ", len(agg_query), "\n")
-
-"""
-__Querying with Logic__
-
-Advanced queries can be constructed using logic.
-
-Below, we combine the two queries above to find all results which fitted a `Gaussian` AND (using the & symbol)
-inferred a value of sigma less than 3.0.
-
-The OR logical clause is also supported via the symbol |.
-"""
-gaussian = agg.model.gaussian
-agg_query = agg.query((gaussian == af.ex.Gaussian) & (gaussian.sigma < 3.0))
-print(
- "Total Samples Objects In Query `Gaussian & sigma < 3.0` = ", len(agg_query), "\n"
-)
-
-"""
-__Database__
-
-The default behaviour of model-fitting results output is to be written to hard-disc in folders. These are simple to
-navigate and manually check.
-
-For small model-fitting tasks this is sufficient, however it does not scale well when performing many model fits to
-large datasets, because manual inspection of results becomes time consuming.
-
-All results can therefore be output to an sqlite3 (https://docs.python.org/3/library/sqlite3.html) relational database,
-meaning that results can be loaded into a Jupyter notebook or Python script for inspection, analysis and interpretation.
-This database supports advanced querying, so that specific model-fits (e.g., which fit a certain model or dataset) can
-be loaded.
-
-__Unique Identifiers__
-
-We have discussed how every model-fit is given a unique identifier, which is used to ensure that the results of the
-model-fit are output to a separate folder on hard-disk.
-
-Each unique identifier is also used to define every entry of the database as it is built. Unique identifiers
-therefore play the same vital role for the database of ensuring that every set of results written to it are unique.
-
-__Building From Output Folder__
-
-The fits above wrote the results to hard-disk in folders, not as an .sqlite database file.
-
-We build the database below, where the `database_name` corresponds to the name of your output folder and is also the
-name of the `.sqlite` database file that is created.
-
-If you are fitting a relatively small number of datasets (e.g. 10-100) having all results written to hard-disk (e.g.
-for quick visual inspection) and using the database for sample wide analysis is beneficial.
-
-We can optionally only include completed model-fits but setting `completed_only=True`.
-
-If you inspect the `output` folder, you will see a `database.sqlite` file which contains the results.
-"""
-database_name = "database"
-
-agg = af.Aggregator.from_database(
- filename=f"{database_name}.sqlite", completed_only=False
-)
-
-agg.add_directory(directory=path.join("output", "cookbooks", database_name))
-
-"""
-__Writing Directly To Database__
-
-Results can be written directly to the .sqlite database file, skipping output to hard-disk entirely, by creating
-a session and passing this to the non-linear search.
-
-The code below shows how to do this, but it is commented out to avoid rerunning the non-linear searches.
-
-This is ideal for tasks where model-fits to hundreds or thousands of datasets are performed, as it becomes unfeasible
-to inspect the results of all fits on the hard-disk.
-
-Our recommended workflow is to set up database analysis scripts using ~10 model-fits, and then scaling these up
-to large samples by writing directly to the database.
-"""
-session = af.db.open_database("database.sqlite")
-
-search = af.DynestyStatic(
- name="multi_result_example",
- path_prefix=path.join("cookbooks", "result"),
- unique_tag=dataset_name, # This makes the unique identifier use the dataset name
- session=session, # This can instruct the search to write to the .sqlite database.
- nlive=50,
-)
-
-"""
-If you run the above code and inspect the `output` folder, you will see a `database.sqlite` file which contains
-the results.
-
-The API for loading a database and creating an aggregator to query is as follows:
-
-# agg = af.Aggregator.from_database("database.sqlite")
-
-Once we have the Aggregator, we can use it to query the database and load results as we did before.
-"""
+"""
+Cookbook: Result
+================
+
+A non-linear search fits a model to a dataset, returning a `Result` object that contains a lot of information on the
+model-fit.
+
+This cookbook provides a concise reference to the result API.
+
+The cookbook then describes how the results of a search can be output to hard-disk and loaded back into Python,
+either using the `Aggregator` object or by building an sqlite database of results. Result loading supports
+queries, so that only the results of interest are returned.
+
+The samples of the non-linear search, which are used to estimate quantities the maximum likelihood model and
+parameter errors, are described separately in the `samples.py` cookbook.
+
+__Contents__
+
+An overview of the `Result` object's functionality is given in the following sections:
+
+ - Simple Fit: Perform a simple model-fit to generate a `Result` object.
+ - Info: Print the `info` attribute of the `Result` object to display a summary of the model-fit.
+ - Max Log Likelihood Instance: Getting the maximum likelihood model instance.
+ - Samples: Getting the samples of the non-linear search from a result.
+ - Custom Result: Extending the `Result` object with custom attributes specific to the model-fit.
+
+The cookbook next describes how results can be output to hard-disk and loaded back into Python via the `Aggregator`:
+
+ - Output To Hard-Disk: Output results to hard-disk so they can be inspected and used to restart a crashed search.
+ - Files: The files that are stored in the `files` folder that is created when results are output to hard-disk.
+ - Loading From Hard-disk: Loading results from hard-disk to Python variables via the aggregator.
+ - Generators: Why loading results uses Python generators to ensure memory efficiency.
+
+The cookbook next gives examples of how to load all the following results from the database:
+
+ - Samples: The samples of the non-linear search loaded via the aggregator.
+ - Loading Model: The model fitted by the non-linear search.
+ - Loading Search: The search used to perform the model-fit.
+ - Loading Samples: The samples of the non-linear search (e.g. all parameter values, log likelihoods, etc.).
+ - Loading Samples Info: Additional information on the samples.
+ - Loading Samples Summary: A summary of the samples of the non-linear search (e.g. the maximum log likelihood model).
+ - Loading Info: The `info` dictionary passed to the search.
+
+The output of results to hard-disk is customizeable and described in the following section:
+
+ - Custom Output: Extend `Analysis` classes to output additional information which can be loaded via the aggregator.
+
+Using queries to load specific results is described in the following sections:
+
+ - Querying Datasets: Query based on the name of the dataset.
+ - Querying Searches: Query based on the name of the search.
+ - Querying Models: Query based on the model that is fitted.
+ - Querying Results: Query based on the results of the model-fit.
+ - Querying with Logic: Use logic to combine queries to load specific results (e.g. AND, OR, etc.).
+
+The final section describes how to use results built in an sqlite database file:
+
+ - Database: Building a database file from the output folder.
+ - Unique Identifiers: The unique identifier of each model-fit.
+ - Building From Output Folder: Build the database from the output folder on hard-disk.
+ - Writing Directly To Database: Writing results directly to the database.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import json
+from os import path
+import numpy as np
+from typing import Optional
+
+import autofit as af
+
+"""
+__Simple Fit__
+
+To illustrate the API of the result object, we first fit a 1D `Gaussian` profile with a `Gaussian` model.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+model = af.Model(af.ex.Gaussian)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+search = af.Emcee(
+ nwalkers=30,
+ nsteps=1000,
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+"""
+__Info__
+
+Printing the `info` attribute shows the overall result of the model-fit in a human readable format.
+"""
+print(result.info)
+
+"""
+__Max Log Likelihood Instance__
+
+The `max_log_likelihood_instance` is the model instance of the maximum log likelihood model, which is the model
+that maximizes the likelihood of the data given the model.
+"""
+instance = result.max_log_likelihood_instance
+
+print("Max Log Likelihood `Gaussian` Instance:")
+print("Centre = ", instance.centre)
+print("Normalization = ", instance.normalization)
+print("Sigma = ", instance.sigma)
+
+"""
+__Samples__
+
+The `Samples` class contains all information on the non-linear search samples, for example the value of every parameter
+sampled using the fit or an instance of the maximum likelihood model.
+"""
+print(result.samples)
+
+"""
+The samples are described in detail separately in the `samples.py` cookbook.
+
+__Custom Result__
+
+The result can be can be customized to include additional information about the model-fit that is specific to your
+model-fitting problem.
+
+For example, for fitting 1D profiles, the `Result` could include the maximum log likelihood model 1D data:
+
+`print(result.max_log_likelihood_model_data_1d)`
+
+In other examples, this quantity has been manually computed after the model-fit has completed.
+
+The custom result API allows us to do this. First, we define a custom `Result` class, which includes the property
+`max_log_likelihood_model_data_1d`.
+"""
+
+
+class ResultExample(af.Result):
+ @property
+ def max_log_likelihood_model_data_1d(self) -> np.ndarray:
+ """
+ Returns the maximum log likelihood model's 1D model data.
+
+ This is an example of how we can pass the `Analysis` class a custom `Result` object and extend this result
+ object with new properties that are specific to the model-fit we are performing.
+ """
+ xvalues = np.arange(self.analysis.data.shape[0])
+
+ return self.instance.model_data_from(xvalues=xvalues)
+
+
+"""
+The custom result has access to the analysis class, meaning that we can use any of its methods or properties to
+compute custom result properties.
+
+To make it so that the `ResultExample` object above is returned by the search we overwrite the `Result` class attribute
+of the `Analysis` and define a `make_result` object describing what we want it to contain:
+"""
+
+
+class Analysis(af.ex.Analysis):
+ """
+ This overwrite means the `ResultExample` class is returned after the model-fit.
+ """
+
+ Result = ResultExample
+
+ def make_result(
+ self,
+ samples_summary: af.SamplesSummary,
+ paths: af.AbstractPaths,
+ samples: Optional[af.SamplesPDF] = None,
+ search_internal: Optional[object] = None,
+ analysis: Optional[object] = None,
+ ) -> Result:
+ """
+ Returns the `Result` of the non-linear search after it is completed.
+
+ The result type is defined as a class variable in the `Analysis` class (see top of code under the python code
+ `class Analysis(af.Analysis)`.
+
+ The result can be manually overwritten by a user to return a user-defined result object, which can be extended
+ with additional methods and attribute specific to the model-fit.
+
+ This example class does example this, whereby the analysis result has been overwritten with the `ResultExample`
+ class, which contains a property `max_log_likelihood_model_data_1d` that returns the model data of the
+ best-fit model. This API means you can customize your result object to include whatever attributes you want
+ and therefore make a result object specific to your model-fit and model-fitting problem.
+
+ The `Result` object you return can be customized to include:
+
+ - The samples summary, which contains the maximum log likelihood instance and median PDF model.
+
+ - The paths of the search, which are used for loading the samples and search internal below when a search
+ is resumed.
+
+ - The samples of the non-linear search (e.g. MCMC chains) also stored in `samples.csv`.
+
+ - The non-linear search used for the fit in its internal representation, which is used for resuming a search
+ and making bespoke visualization using the search's internal results.
+
+ - The analysis used to fit the model (default disabled to save memory, but option may be useful for certain
+ projects).
+
+ Parameters
+ ----------
+ samples_summary
+ The summary of the samples of the non-linear search, which include the maximum log likelihood instance and
+ median PDF model.
+ paths
+ An object describing the paths for saving data (e.g. hard-disk directories or entries in sqlite database).
+ samples
+ The samples of the non-linear search, for example the chains of an MCMC run.
+ search_internal
+ The internal representation of the non-linear search used to perform the model-fit.
+ analysis
+ The analysis used to fit the model.
+
+ Returns
+ -------
+ Result
+ The result of the non-linear search, which is defined as a class variable in the `Analysis` class.
+ """
+ return self.Result(
+ samples_summary=samples_summary,
+ paths=paths,
+ samples=samples,
+ search_internal=search_internal,
+ analysis=self,
+ )
+
+
+"""
+Using the `Analysis` class above, the `Result` object returned by the search is now a `ResultExample` object.
+"""
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+search = af.Emcee(
+ nwalkers=30,
+ nsteps=1000,
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+print(result.max_log_likelihood_model_data_1d)
+
+"""
+__Output To Hard-Disk__
+
+By default, a non-linear search does not output its results to hard-disk and its results can only be inspected
+in Python via the `result` object.
+
+However, the results of any non-linear search can be output to hard-disk by passing the `name` and / or `path_prefix`
+attributes, which are used to name files and output the results to a folder on your hard-disk.
+
+This cookbook now runs the three searches with output to hard-disk enabled, so you can see how the results are output
+to hard-disk and to then illustrate how they can be loaded back into Python.
+
+Note that an `info` dictionary is also passed to the search, which includes the date of the model-fit and the exposure
+time of the dataset. This information is stored output to hard-disk and can be loaded to help interpret the results.
+"""
+info = {"date_of_observation": "01-02-18", "exposure_time": 1000.0}
+
+dataset_name_list = ["gaussian_x1_0", "gaussian_x1_1", "gaussian_x1_2"]
+
+model = af.Collection(gaussian=af.ex.Gaussian)
+
+model.gaussian.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.gaussian.normalization = af.LogUniformPrior(lower_limit=1e-2, upper_limit=1e2)
+model.gaussian.sigma = af.TruncatedGaussianPrior(
+ mean=10.0, sigma=5.0, lower_limit=0.0, upper_limit=np.inf
+)
+
+for dataset_name in dataset_name_list:
+ dataset_path = path.join("dataset", "example_1d", dataset_name)
+
+ data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+ noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+ )
+
+ analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+ search = af.DynestyStatic(
+ name="multi_result_example",
+ path_prefix=path.join("cookbooks", "result"),
+ unique_tag=dataset_name, # This makes the unique identifier use the dataset name
+ nlive=50,
+ )
+
+ print(
+ """
+ The non-linear search has begun running.
+ This Jupyter notebook cell with progress once search has completed, this could take a few minutes!
+ """
+ )
+
+ result = search.fit(model=model, analysis=analysis, info=info)
+
+print("Search has finished run - you may now continue the notebook.")
+
+"""
+__Files__
+
+By outputting results to hard-disk, a `files` folder is created containing .json / .csv files of the model,
+samples, search, etc, for each fit.
+
+You should check it out now for the completed fits on your hard-disk.
+
+A description of all files is as follows:
+
+ - `model`: The `model` defined above and used in the model-fit (`model.json`).
+ - `search`: The non-linear search settings (`search.json`).
+ - `samples`: The non-linear search samples (`samples.csv`).
+ - `samples_info`: Additional information about the samples (`samples_info.json`).
+ - `samples_summary`: A summary of key results of the samples (`samples_summary.json`).
+ - `info`: The info dictionary passed to the search (`info.json`).
+ - `covariance`: The inferred covariance matrix (`covariance.csv`).
+ - `data`: The 1D noisy data used that is fitted (`data.json`).
+ - `noise_map`: The 1D noise-map fitted (`noise_map.json`).
+
+The `samples` and `samples_summary` results contain a lot of repeated information. The `samples` result contains
+the full non-linear search samples, for example every parameter sample and its log likelihood. The `samples_summary`
+contains a summary of the results, for example the maximum log likelihood model and error estimates on parameters
+at 1 and 3 sigma confidence.
+
+Accessing results via the `samples_summary` is much faster, because as it does not reperform calculations using the full
+list of samples. Therefore, if the result you want is accessible via the `samples_summary` you should use it
+but if not you can revert to the `samples.
+
+__Loading From Hard-disk__
+
+The multi-fits above wrote the results to hard-disk in three distinct folders, one for each dataset.
+
+Their results are loaded using the `Aggregator` object, which finds the results in the output directory and can
+load them into Python objects.
+"""
+from autofit.aggregator.aggregator import Aggregator
+
+agg = Aggregator.from_directory(
+ directory=path.join("multi_result_example"),
+)
+
+"""
+__Generators__
+
+Before using the aggregator to inspect results, lets discuss Python generators.
+
+A generator is an object that iterates over a function when it is called. The aggregator creates all of the objects
+that it loads from the database as generators (as opposed to a list, or dictionary, or another Python type).
+
+This is because generators are memory efficient, as they do not store the entries of the database in memory
+simultaneously. This contrasts objects like lists and dictionaries, which store all entries in memory all at once.
+If you fit a large number of datasets, lists and dictionaries will use a lot of memory and could crash your computer!
+
+Once we use a generator in the Python code, it cannot be used again. To perform the same task twice, the
+generator must be remade it. This cookbook therefore rarely stores generators as variables and instead uses the
+aggregator to create each generator at the point of use.
+
+To create a generator of a specific set of results, we use the `values` method. This takes the `name` of the
+object we want to create a generator of, for example inputting `name=samples` will return the results `Samples`
+object (which is illustrated in detail below).
+"""
+for samples in agg.values("samples"):
+ print(samples.parameter_lists[0])
+
+"""
+__Samples__
+
+By converting this generator to a list and printing it, it is a list of 3 `SamplesNest` objects, corresponding to
+the 3 model-fits performed above.
+"""
+print("Samples:\n")
+samples_gen = agg.values("samples")
+print(samples_gen)
+print("Total Samples Objects = ", len(agg), "\n")
+
+"""
+__Loading Model__
+
+The model used to perform the model fit for each of the 3 datasets can be loaded via the aggregator and printed.
+"""
+model_gen = agg.values("model")
+
+for model in model_gen:
+ print(model.info)
+
+"""
+__Loading Search__
+
+The non-linear search used to perform the model fit can be loaded via the aggregator and printed.
+"""
+search_gen = agg.values("search")
+
+for search in search_gen:
+ print(search)
+
+"""
+__Loading Samples__
+
+The `Samples` class contains all information on the non-linear search samples, for example the value of every parameter
+sampled using the fit or an instance of the maximum likelihood model.
+
+The `Samples` class is described fully in the results cookbook.
+"""
+for samples in agg.values("samples"):
+ print("The tenth sample`s third parameter")
+ print(samples.parameter_lists[9][2], "\n")
+
+ instance = samples.max_log_likelihood()
+
+ print("Max Log Likelihood `Gaussian` Instance:")
+ print("Centre = ", instance.gaussian.centre)
+ print("Normalization = ", instance.gaussian.normalization)
+ print("Sigma = ", instance.gaussian.sigma, "\n")
+
+"""
+__Loading Samples Info__
+
+The samples info contains additional information on the samples, which depends on the non-linear search used.
+
+For example, for a nested sampling algorithm it contains information on the number of live points, for a MCMC
+algorithm it contains information on the number of steps, etc.
+"""
+for samples_info in agg.values("samples_info"):
+ print(samples_info)
+
+"""
+__Loading Samples Summary__
+
+The samples summary contains a subset of results access via the `Samples`, for example the maximum likelihood model
+and parameter error estimates.
+
+Using the samples method above can be slow, as the quantities have to be computed from all non-linear search samples
+(e.g. computing errors requires that all samples are marginalized over). This information is stored directly in the
+samples summary and can therefore be accessed instantly.
+"""
+# for samples_summary in agg.values("samples_summary"):
+#
+# instance = samples_summary.max_log_likelihood()
+#
+# print("Max Log Likelihood `Gaussian` Instance:")
+# print("Centre = ", instance.centre)
+# print("Normalization = ", instance.normalization)
+# print("Sigma = ", instance.sigma, "\n")
+
+"""
+__Loading Info__
+
+The info dictionary passed to the search, discussed earlier in this cookbook, is accessible.
+"""
+for info in agg.values("info"):
+ print(info["date_of_observation"])
+ print(info["exposure_time"])
+
+"""
+__Custom Output__
+
+The results accessible via the database (e.g. `model`, `samples`) are those contained in the `files` folder.
+
+By extending an `Analysis` class with the methods `save_attributes` and `save_results`,
+custom files can be written to the `files` folder and become accessible via the database.
+
+To save the objects in a human readable and loaded .json format, the `data` and `noise_map`, which are natively stored
+as 1D numpy arrays, are converted to a suitable dictionary output format. This uses the **PyAutoConf** method
+`to_dict`.
+"""
+
+
+class Analysis(af.Analysis):
+ def __init__(self, data: np.ndarray, noise_map: np.ndarray):
+ """
+ Standard Analysis class example used throughout PyAutoFit examples.
+ """
+ super().__init__()
+
+ self.data = data
+ self.noise_map = noise_map
+
+ def log_likelihood_function(self, instance) -> float:
+ """
+ Standard log likelihood function used throughout PyAutoFit examples.
+ """
+
+ xvalues = np.arange(self.data.shape[0])
+
+ model_data = instance.model_data_from(xvalues=xvalues)
+
+ residual_map = self.data - model_data
+ chi_squared_map = (residual_map / self.noise_map) ** 2.0
+ chi_squared = sum(chi_squared_map)
+ noise_normalization = np.sum(np.log(2 * np.pi * self.noise_map**2.0))
+ log_likelihood = -0.5 * (chi_squared + noise_normalization)
+
+ return log_likelihood
+
+ def save_attributes(self, paths: af.DirectoryPaths):
+ """
+ Before the non-linear search begins, this routine saves attributes of the `Analysis` object to the `files`
+ folder such that they can be loaded after the analysis using PyAutoFit's database and aggregator tools.
+
+ For this analysis, it uses the `AnalysisDataset` object's method to output the following:
+
+ - The dataset's data as a .json file.
+ - The dataset's noise-map as a .json file.
+
+ These are accessed using the aggregator via `agg.values("data")` and `agg.values("noise_map")`.
+
+ They are saved using the paths function `save_json`, noting that this saves outputs appropriate for the
+ sqlite3 database.
+
+ Parameters
+ ----------
+ paths
+ The paths object which manages all paths, e.g. where the non-linear search outputs are stored,
+ visualization, and the pickled objects used by the aggregator output by this function.
+ """
+ from autoconf.dictable import to_dict
+
+ paths.save_json(name="data", object_dict=to_dict(self.data))
+ paths.save_json(name="noise_map", object_dict=to_dict(self.noise_map))
+
+ def save_results(self, paths: af.DirectoryPaths, result: af.Result):
+ """
+ At the end of a model-fit, this routine saves attributes of the `Analysis` object to the `files`
+ folder such that they can be loaded after the analysis using PyAutoFit's database and aggregator tools.
+
+ For this analysis it outputs the following:
+
+ - The maximum log likelihood model data as a .json file.
+
+ This is accessed using the aggregator via `agg.values("model_data")`.
+
+ Parameters
+ ----------
+ paths
+ The paths object which manages all paths, e.g. where the non-linear search outputs are stored,
+ visualization and the pickled objects used by the aggregator output by this function.
+ result
+ The result of a model fit, including the non-linear search, samples and maximum likelihood model.
+ """
+ xvalues = np.arange(self.data.shape[0])
+
+ instance = result.max_log_likelihood_instance
+
+ model_data = instance.model_data_from(xvalues=xvalues)
+
+ # The path where model_data.json is saved, e.g. output/dataset_name/unique_id/files/model_data.json
+
+ file_path = (path.join(paths._json_path, "model_data.json"),)
+
+ with open(file_path, "w+") as f:
+ json.dump(model_data, f, indent=4)
+
+
+"""
+__Querying Datasets__
+
+The aggregator can query the database, returning only specific fits of interested.
+
+We can query using the `dataset_name` string we input into the model-fit above, in order to get the results
+of a fit to a specific dataset.
+
+For example, querying using the string `gaussian_x1_1` returns results for only the fit using the
+second `Gaussian` dataset.
+"""
+unique_tag = agg.search.unique_tag
+agg_query = agg.query(unique_tag == "gaussian_x1_1")
+
+"""
+As expected, this list has only 1 `SamplesNest` corresponding to the second dataset.
+"""
+print(agg_query.values("samples"))
+print("Total Samples Objects via dataset_name Query = ", len(agg_query), "\n")
+
+"""
+If we query using an incorrect dataset name we get no results.
+"""
+unique_tag = agg.search.unique_tag
+agg_query = agg.query(unique_tag == "incorrect_name")
+samples_gen = agg_query.values("samples")
+
+"""
+__Querying Searches__
+
+We can query using the `name` of the non-linear search used to fit the model.
+
+In this cookbook, all three fits used the same search, named `database_example`. Query based on search name in this
+example is therefore somewhat pointless.
+
+However, querying based on the search name is useful for model-fits which use a range of searches, for example
+if different non-linear searches are used multiple times.
+
+As expected, the query using search name below contains all 3 results.
+"""
+name = agg.search.name
+agg_query = agg.query(name == "database_example")
+
+print(agg_query.values("samples"))
+print("Total Samples Objects via name Query = ", len(agg_query), "\n")
+
+"""
+__Querying Models__
+
+We can query based on the model fitted.
+
+For example, we can load all results which fitted a `Gaussian` model-component, which in this simple example is all
+3 model-fits.
+
+Querying via the model is useful for loading results after performing many model-fits with many different model
+parameterizations to large (e.g. Bayesian model comparison).
+
+[Note: the code `agg.model.gaussian` corresponds to the fact that in the `Collection` above, we named the model
+component `gaussian`. If this `Collection` had used a different name the code below would change
+correspondingly. Models with multiple model components (e.g., `gaussian` and `exponential`) are therefore also easily
+accessed via the database.]
+"""
+gaussian = agg.model.gaussian
+agg_query = agg.query(gaussian == af.ex.Gaussian)
+print("Total Samples Objects via `Gaussian` model query = ", len(agg_query), "\n")
+
+"""
+__Querying Results__
+
+We can query based on the results of the model-fit.
+
+Below, we query the database to find all fits where the inferred value of `sigma` for the `Gaussian` is less
+than 3.0 (which returns only the first of the three model-fits).
+"""
+gaussian = agg.model.gaussian
+agg_query = agg.query(gaussian.sigma < 3.0)
+print("Total Samples Objects In Query `gaussian.sigma < 3.0` = ", len(agg_query), "\n")
+
+"""
+__Querying with Logic__
+
+Advanced queries can be constructed using logic.
+
+Below, we combine the two queries above to find all results which fitted a `Gaussian` AND (using the & symbol)
+inferred a value of sigma less than 3.0.
+
+The OR logical clause is also supported via the symbol |.
+"""
+gaussian = agg.model.gaussian
+agg_query = agg.query((gaussian == af.ex.Gaussian) & (gaussian.sigma < 3.0))
+print(
+ "Total Samples Objects In Query `Gaussian & sigma < 3.0` = ", len(agg_query), "\n"
+)
+
+"""
+__Database__
+
+The default behaviour of model-fitting results output is to be written to hard-disc in folders. These are simple to
+navigate and manually check.
+
+For small model-fitting tasks this is sufficient, however it does not scale well when performing many model fits to
+large datasets, because manual inspection of results becomes time consuming.
+
+All results can therefore be output to an sqlite3 (https://docs.python.org/3/library/sqlite3.html) relational database,
+meaning that results can be loaded into a Jupyter notebook or Python script for inspection, analysis and interpretation.
+This database supports advanced querying, so that specific model-fits (e.g., which fit a certain model or dataset) can
+be loaded.
+
+__Unique Identifiers__
+
+We have discussed how every model-fit is given a unique identifier, which is used to ensure that the results of the
+model-fit are output to a separate folder on hard-disk.
+
+Each unique identifier is also used to define every entry of the database as it is built. Unique identifiers
+therefore play the same vital role for the database of ensuring that every set of results written to it are unique.
+
+__Building From Output Folder__
+
+The fits above wrote the results to hard-disk in folders, not as an .sqlite database file.
+
+We build the database below, where the `database_name` corresponds to the name of your output folder and is also the
+name of the `.sqlite` database file that is created.
+
+If you are fitting a relatively small number of datasets (e.g. 10-100) having all results written to hard-disk (e.g.
+for quick visual inspection) and using the database for sample wide analysis is beneficial.
+
+We can optionally only include completed model-fits but setting `completed_only=True`.
+
+If you inspect the `output` folder, you will see a `database.sqlite` file which contains the results.
+"""
+database_name = "database"
+
+agg = af.Aggregator.from_database(
+ filename=f"{database_name}.sqlite", completed_only=False
+)
+
+agg.add_directory(directory=path.join("output", "cookbooks", database_name))
+
+"""
+__Writing Directly To Database__
+
+Results can be written directly to the .sqlite database file, skipping output to hard-disk entirely, by creating
+a session and passing this to the non-linear search.
+
+The code below shows how to do this, but it is commented out to avoid rerunning the non-linear searches.
+
+This is ideal for tasks where model-fits to hundreds or thousands of datasets are performed, as it becomes unfeasible
+to inspect the results of all fits on the hard-disk.
+
+Our recommended workflow is to set up database analysis scripts using ~10 model-fits, and then scaling these up
+to large samples by writing directly to the database.
+"""
+session = af.db.open_database("database.sqlite")
+
+search = af.DynestyStatic(
+ name="multi_result_example",
+ path_prefix=path.join("cookbooks", "result"),
+ unique_tag=dataset_name, # This makes the unique identifier use the dataset name
+ session=session, # This can instruct the search to write to the .sqlite database.
+ nlive=50,
+)
+
+"""
+If you run the above code and inspect the `output` folder, you will see a `database.sqlite` file which contains
+the results.
+
+The API for loading a database and creating an aggregator to query is as follows:
+
+# agg = af.Aggregator.from_database("database.sqlite")
+
+Once we have the Aggregator, we can use it to query the database and load results as we did before.
+"""
diff --git a/scripts/cookbooks/samples.py b/scripts/cookbooks/samples.py
index 96b7f8da..937b2d9b 100644
--- a/scripts/cookbooks/samples.py
+++ b/scripts/cookbooks/samples.py
@@ -1,589 +1,585 @@
-"""
-Cookbook: Samples
-=================
-
-A non-linear search samples parameter space to find the global maximum log likelihood solution.
-
-The `Samples` object contains the history of the non-linear search, including the model parameters and log likelihood
-of every accepted sample.
-
-This cookbook provides an overview of using the samples.
-
-__Contents__
-
- - Model Fit: Perform a simple model-fit to create a ``Samples`` object.
- - Samples: The `Samples` object`, containing all non-linear samples (e.g. parameters, log likelihoods, etc.).
- - Parameters: Accessing the parameters of the model from the samples.
- - Figures of Merit: The log likelihood, log prior, log posterior and weight_list of every accepted sample.
- - Instances: Returning instances of the model corresponding to a particular sample (e.g. the maximum log likelihood).
- - Posterior / PDF: The median PDF model instance and PDF vectors of all model parameters via 1D marginalization.
- - Errors: The errors on every parameter estimated from the PDF, computed via marginalized 1D PDFs at an input sigma.
- - Samples Summary: A summary of the samples of the non-linear search (e.g. the maximum log likelihood model) which can
- be faster to load than the full set of samples.
- - Sample Instance: The model instance of any accepted sample.
- - Search Plots: Plots of the non-linear search, for example a corner plot or 1D PDF of every parameter.
- - Maximum Likelihood: The maximum log likelihood model value.
- - Bayesian Evidence: The log evidence estimated via a nested sampling algorithm.
- - Collection: Results created from models defined via a `Collection` object.
- - Lists: Extracting results as Python lists instead of instances.
- - Latex: Producing latex tables of results (e.g. for a paper).
-
-The following sections outline how to use advanced features of the results, which you may skip on a first read:
-
- - Derived Quantities: Computing quantities and errors for quantities and parameters not included directly in the model.
- - Derived Errors Manual (Advanced): Manually computing errors on derived quantities from the PDF of samples.
- - Samples Filtering (Advanced): Filter the `Samples` object to only contain samples fulfilling certain criteria.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import autofit as af
-import autofit.plot as aplt
-
-from os import path
-import matplotlib.pyplot as plt
-import numpy as np
-
-"""
-__Model Fit__
-
-To illustrate results, we need to perform a model-fit in order to create a `Result` object.
-
-We do this below using the standard API and noisy 1D signal example, which you should be familiar with from other
-example scripts.
-
-Note that the `Gaussian` and `Analysis` classes come via the `af.ex` module, which contains example model components
-that are identical to those found throughout the examples.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-model = af.Model(af.ex.Gaussian)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-search = af.Emcee(
- name="cookbook_result",
- nwalkers=30,
- nsteps=1000,
- number_of_cores=1,
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-"""
-__Samples__
-
-The result contains a `Samples` object, which contains all samples of the non-linear search.
-
-Each sample corresponds to a set of model parameters that were evaluated and accepted by the non linear search,
-in this example `emcee.`
-
-This includes their log likelihoods, which are used for computing additional information about the model-fit,
-for example the error on every parameter.
-
-Our model-fit used the MCMC algorithm Emcee, so the `Samples` object returned is a `SamplesMCMC` object.
-"""
-samples = result.samples
-
-print("MCMC Samples: \n")
-print(samples)
-
-"""
-__Parameters__
-
-The parameters are stored as a list of lists, where:
-
- - The outer list is the size of the total number of samples.
- - The inner list is the size of the number of free parameters in the fit.
-"""
-samples = result.samples
-
-print("Sample 5's second parameter value (Gaussian -> normalization):")
-print(samples.parameter_lists[4][1])
-print("Sample 10`s third parameter value (Gaussian -> sigma)")
-print(samples.parameter_lists[9][2], "\n")
-
-"""
-__Figures of Merit__
-
-The `Samples` class contains the log likelihood, log prior, log posterior and weight_list of every accepted sample, where:
-
-- The `log_likelihood` is the value evaluated in the `log_likelihood_function`.
-
-- The `log_prior` encodes information on how parameter priors map log likelihood values to log posterior values.
-
-- The `log_posterior` is `log_likelihood + log_prior`.
-
-- The `weight` gives information on how samples are combined to estimate the posterior, which depends on type of search
- used (for `Emcee` they are all 1's meaning they are weighted equally).
-
-Lets inspect these values for the tenth sample.
-"""
-print("log(likelihood), log(prior), log(posterior) and weight of the tenth sample.")
-print(samples.log_likelihood_list[9])
-print(samples.log_prior_list[9])
-print(samples.log_posterior_list[9])
-print(samples.weight_list[9])
-
-"""
-__Instances__
-
-Many results can be returned as an instance of the model, using the Python class structure of the model composition.
-
-For example, we can return the model parameters corresponding to the maximum log likelihood sample.
-
-The attributes of the `instance` (`centre`, `normalization` and `sigma`) have these names due to how we composed
-the `Gaussian` class via the `Model` above. They would be named structured and named differently if we hd
-used a `Collection` and different names.
-"""
-instance = samples.max_log_likelihood()
-
-print("Max Log Likelihood `Gaussian` Instance:")
-print("Centre = ", instance.centre)
-print("Normalization = ", instance.normalization)
-print("Sigma = ", instance.sigma, "\n")
-
-"""
-This makes it straight forward to plot the median PDF model:
-"""
-model_data = instance.model_data_from(xvalues=np.arange(data.shape[0]))
-
-plt.plot(range(data.shape[0]), data)
-plt.plot(range(data.shape[0]), model_data)
-plt.title("Illustrative model fit to 1D `Gaussian` profile data.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-__Posterior / PDF__
-
-The result contains the full posterior information of our non-linear search, which can be used for parameter
-estimation.
-
-PDF stands for "Probability Density Function" and it quantifies probability of each model parameter having values
-that are sampled. It therefore enables error estimation via a process called marginalization.
-
-The median pdf vector is available, which estimates every parameter via 1D marginalization of their PDFs.
-"""
-instance = samples.median_pdf()
-
-print("Median PDF `Gaussian` Instance:")
-print("Centre = ", instance.centre)
-print("Normalization = ", instance.normalization)
-print("Sigma = ", instance.sigma, "\n")
-
-"""
-__Errors__
-
-Methods for computing error estimates on all parameters are provided.
-
-This again uses 1D marginalization, now at an input sigma confidence limit.
-
-By inputting `sigma=3.0` margnialization find the values spanning 99.7% of 1D PDF. Changing this to `sigma=1.0`
-would give the errors at the 68.3% confidence limit.
-"""
-instance_upper_sigma = samples.errors_at_upper_sigma(sigma=3.0)
-instance_lower_sigma = samples.errors_at_lower_sigma(sigma=3.0)
-
-print("Upper Error values (at 3.0 sigma confidence):")
-print("Centre = ", instance_upper_sigma.centre)
-print("Normalization = ", instance_upper_sigma.normalization)
-print("Sigma = ", instance_upper_sigma.sigma, "\n")
-
-print("lower Error values (at 3.0 sigma confidence):")
-print("Centre = ", instance_lower_sigma.centre)
-print("Normalization = ", instance_lower_sigma.normalization)
-print("Sigma = ", instance_lower_sigma.sigma, "\n")
-
-"""
-They can also be returned at the values of the parameters at their error values.
-"""
-instance_upper_values = samples.values_at_upper_sigma(sigma=3.0)
-instance_lower_values = samples.values_at_lower_sigma(sigma=3.0)
-
-print("Upper Parameter values w/ error (at 3.0 sigma confidence):")
-print("Centre = ", instance_upper_values.centre)
-print("Normalization = ", instance_upper_values.normalization)
-print("Sigma = ", instance_upper_values.sigma, "\n")
-
-print("lower Parameter values w/ errors (at 3.0 sigma confidence):")
-print("Centre = ", instance_lower_values.centre)
-print("Normalization = ", instance_lower_values.normalization)
-print("Sigma = ", instance_lower_values.sigma, "\n")
-
-"""
-__Samples Summary__
-
-The samples summary contains a subset of results access via the `Samples`, for example the maximum likelihood model
-and parameter error estimates.
-
-Using the samples method above can be slow, as the quantities have to be computed from all non-linear search samples
-(e.g. computing errors requires that all samples are marginalized over). This information is stored directly in the
-samples summary and can therefore be accessed instantly.
-"""
-print(samples.summary().max_log_likelihood_sample)
-
-"""
-__Sample Instance__
-
-A non-linear search retains every model that is accepted during the model-fit.
-
-We can create an instance of any model -- below we create an instance of the last accepted model.
-"""
-instance = samples.from_sample_index(sample_index=-1)
-
-print("Gaussian Instance of last sample")
-print("Centre = ", instance.centre)
-print("Normalization = ", instance.normalization)
-print("Sigma = ", instance.sigma, "\n")
-
-"""
-__Search Plots__
-
-The Probability Density Functions (PDF's) of the results can be plotted using the non-linear search in-built
-visualization tools.
-
-This fit used `Emcee` therefore we use `corner.py` for visualization via the `aplt.corner_cornerpy` function.
-
-The `autofit_workspace/*/plots` folder illustrates other packages that can be used to make these plots using
-the standard output results formats (e.g. `GetDist.py`).
-"""
-aplt.corner_cornerpy(samples=result.samples)
-
-"""
-__Maximum Likelihood__
-
-The maximum log likelihood value of the model-fit can be estimated by simple taking the maximum of all log
-likelihoods of the samples.
-
-If different models are fitted to the same dataset, this value can be compared to determine which model provides
-the best fit (e.g. which model has the highest maximum likelihood)?
-"""
-print("Maximum Log Likelihood: \n")
-print(max(samples.log_likelihood_list))
-
-"""
-__Bayesian Evidence__
-
-If a nested sampling non-linear search is used, the evidence of the model is also available which enables Bayesian
-model comparison to be performed (given we are using Emcee, which is not a nested sampling algorithm, the log evidence
-is None).
-
-A full discussion of Bayesian model comparison is given in `autofit_workspace/*/features/bayes_model_comparison.py`.
-"""
-log_evidence = samples.log_evidence
-print(f"Log Evidence: {log_evidence}")
-
-"""
-__Collection__
-
-The examples correspond to a model where `af.Model(Gaussian)` was used to compose the model.
-
-Below, we illustrate how the results API slightly changes if we compose our model using a `Collection`:
-"""
-model = af.Collection(gaussian=af.ex.Gaussian, exponential=af.ex.Exponential)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-search = af.Emcee(
- nwalkers=50,
- nsteps=1000,
- number_of_cores=1,
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-"""
-The `result.info` shows the result for the model with both a `Gaussian` and `Exponential` profile.
-"""
-print(result.info)
-
-"""
-Result instances again use the Python classes used to compose the model.
-
-However, because our fit uses a `Collection` the `instance` has attributes named according to the names given to the
-`Collection`, which above were `gaussian` and `exponential`.
-
-For complex models, with a large number of model components and parameters, this offers a readable API to interpret
-the results.
-"""
-samples = result.samples
-
-instance = samples.max_log_likelihood()
-
-print("Max Log Likelihood `Gaussian` Instance:")
-print("Centre = ", instance.gaussian.centre)
-print("Normalization = ", instance.gaussian.normalization)
-print("Sigma = ", instance.gaussian.sigma, "\n")
-
-print("Max Log Likelihood Exponential Instance:")
-print("Centre = ", instance.exponential.centre)
-print("Normalization = ", instance.exponential.normalization)
-print("Sigma = ", instance.exponential.rate, "\n")
-
-"""
-__Lists__
-
-All results can alternatively be returned as a 1D list of values, by passing `as_instance=False`:
-"""
-max_lh_list = samples.max_log_likelihood(as_instance=False)
-print("Max Log Likelihood Model Parameters: \n")
-print(max_lh_list, "\n\n")
-
-"""
-The list above does not tell us which values correspond to which parameters.
-
-The following quantities are available in the `Model`, where the order of their entries correspond to the parameters
-in the `ml_vector` above:
-
- - `paths`: a list of tuples which give the path of every parameter in the `Model`.
- - `parameter_names`: a list of shorthand parameter names derived from the `paths`.
- - `parameter_labels`: a list of parameter labels used when visualizing non-linear search results (see below).
-
-For simple models like the one fitted in this tutorial, the quantities below are somewhat redundant. For the
-more complex models they are important for tracking the parameters of the model.
-"""
-model = samples.model
-
-print(model.paths)
-print(model.parameter_names)
-print(model.parameter_labels)
-print(model.model_component_and_parameter_names)
-print("\n")
-
-"""
-All the methods above are available as lists.
-"""
-instance = samples.median_pdf(as_instance=False)
-values_at_upper_sigma = samples.values_at_upper_sigma(sigma=3.0, as_instance=False)
-values_at_lower_sigma = samples.values_at_lower_sigma(sigma=3.0, as_instance=False)
-errors_at_upper_sigma = samples.errors_at_upper_sigma(sigma=3.0, as_instance=False)
-errors_at_lower_sigma = samples.errors_at_lower_sigma(sigma=3.0, as_instance=False)
-
-"""
-__Latex__
-
-If you are writing modeling results up in a paper, you can use inbuilt latex tools to create latex table code which
-you can copy to your .tex document.
-
-By combining this with the filtering tools below, specific parameters can be included or removed from the latex.
-
-Remember that the superscripts of a parameter are loaded from the config file `notation/label.yaml`, providing high
-levels of customization for how the parameter names appear in the latex table. This is especially useful if your model
-uses the same model components with the same parameter, which therefore need to be distinguished via superscripts.
-"""
-latex = af.text.Samples.latex(
- samples=result.samples,
- median_pdf_model=True,
- sigma=3.0,
- name_to_label=True,
- include_name=True,
- include_quickmath=True,
- prefix="Example Prefix ",
- suffix=" \\[-2pt]",
-)
-
-print(latex)
-
-"""
-__Derived Quantities__
-
-The parameters `centre`, `normalization` and `sigma` are the model parameters of the `Gaussian`. They are sampled
-directly by the non-linear search and we can therefore use the `Samples` object to easily determine their values and
-errors.
-
-Derived quantities (also called latent variables) are those which are not sampled directly by the non-linear search,
-but one may still wish to know their values and errors after the fit is complete. For example, what if we want the
-error on the full width half maximum (FWHM) of the Gaussian?
-
-This is achieved by adding them to the `compute_latent_variables` method of the `Analysis` class, which is called
-after the non-linear search has completed. The analysis cookbook illustrates how to do this.
-
-The example analysis used above includes a `compute_latent_variables` method that computes the FWHM of the Gaussian
-profile.
-
-This leads to a number of noteworthy outputs:
-
- - A `latent.results` file is output to the results folder, which includes the value and error of all derived quantities
- based on the non-linear search samples (in this example only the `fwhm`).
-
- - A `latent/samples.csv` is output which lists every accepted sample's value of every derived quantity, which is again
- analogous to the `samples.csv` file (in this example only the `fwhm`).
-
- - A `latent/samples_summary.json` is output which acts analogously to `samples_summary.json` but for the derived
- quantities of the model (in this example only the `fwhm`).
-
-Derived quantities are also accessible via the `Samples` object, following a similar API to the model parameters:
-"""
-latent = analysis.compute_latent_samples(result.samples)
-
-instance = latent.max_log_likelihood()
-
-print(f"Max Likelihood FWHM: {instance.gaussian.fwhm}")
-
-instance = latent.median_pdf()
-
-print(f"Median PDF FWHM {instance.gaussian.fwhm}")
-
-"""
-__Derived Errors Manual (Advanced)__
-
-The derived quantities decorator above provides a simple interface for computing the errors of a derived quantity and
-ensuring all results are easily inspected in the output results folder.
-
-However, you may wish to compute the errors of a derived quantity manually. For example, if it is a quantity that
-you did not decorate before performing the fit, or if it is computationally expensive to compute and you only want
-to compute it specific circumstances.
-
-We create the PDF of the derived quantity, the FWHM, manually, which we marginalize over using the same function
-we use to marginalize model parameters. We compute the FWHM of every accepted model sampled by the non-linear search
-and use this determine the PDF of the FWHM.
-
-When combining the FWHM's we weight each value by its `weight`. For Emcee, an MCMC algorithm, the weight of every
-sample is 1, but weights may take different values for other non-linear searches.
-
-In order to pass these samples to the function `marginalize`, which marginalizes over the PDF of the FWHM to compute
-its error, we also pass the weight list of the samples.
-"""
-fwhm_list = []
-
-for sample in samples.sample_list:
- instance = sample.instance_for_model(model=samples.model)
-
- sigma = instance.gaussian.sigma
-
- fwhm = 2 * np.sqrt(2 * np.log(2)) * sigma
-
- fwhm_list.append(fwhm)
-
-median_fwhm, lower_fwhm, upper_fwhm = af.marginalize(
- parameter_list=fwhm_list, sigma=3.0, weight_list=samples.weight_list
-)
-
-print(f"FWHM = {median_fwhm} ({upper_fwhm} {lower_fwhm}")
-
-"""
-The calculation above could be computationally expensive, if there are many samples and the derived quantity is
-slow to compute.
-
-An alternative approach, which will provide comparable accuracy provided enough draws are used, is to sample
-points randomy from the PDF of the model and use these to compute the derived quantity.
-
-Draws are from the PDF of the model, so the weights of the samples are accounted for and we therefore do not
-pass them to the `marginalize` function (it essentially treats all samples as having equal weight).
-
-TRY AND EXCEPT INCLUDED TO FIX BUG, NEED TO SOLVE IN FUTURE.
-"""
-try:
- random_draws = 50
-
- fwhm_list = []
-
- for i in range(random_draws):
- instance = samples.draw_randomly_via_pdf()
-
- sigma = instance.gaussian.sigma
-
- fwhm = 2 * np.sqrt(2 * np.log(2)) * sigma
-
- fwhm_list.append(fwhm)
-
- median_fwhm, lower_fwhm, upper_fwhm = af.marginalize(
- parameter_list=fwhm_list, sigma=3.0, weight_list=samples.weight_list
- )
-
- print(f"fwhm = {median_fwhm} ({upper_fwhm} {lower_fwhm}")
-
-except ValueError:
- pass
-
-"""
-__Samples Filtering (Advanced)__
-
-Our samples object has the results for all three parameters in our model. However, we might only be interested in the
-results of a specific parameter.
-
-The basic form of filtering specifies parameters via their path, which was printed above via the model and is printed
-again below.
-"""
-samples = result.samples
-
-print("Parameter paths in the model which are used for filtering:")
-print(samples.model.paths)
-
-print("All parameters of the very first sample")
-print(samples.parameter_lists[0])
-
-samples = samples.with_paths([("gaussian", "centre")])
-
-print("All parameters of the very first sample (containing only the Gaussian centre.")
-print(samples.parameter_lists[0])
-
-print("Maximum Log Likelihood Model Instances (containing only the Gaussian centre):\n")
-print(samples.max_log_likelihood(as_instance=False))
-
-"""
-We specified each path as a list of tuples of strings.
-
-This is how the source code internally stores the path to different components of the model, but it is not
-consistent with the API used to compose a model.
-
-We can alternatively use the following API:
-"""
-samples = result.samples
-
-samples = samples.with_paths(["gaussian.centre"])
-
-print("All parameters of the very first sample (containing only the Gaussian centre).")
-print(samples.parameter_lists[0])
-
-"""
-We filtered the `Samples` above by asking for all parameters which included the path ("gaussian", "centre").
-
-We can alternatively filter the `Samples` object by removing all parameters with a certain path. Below, we remove
-the Gaussian's `centre` to be left with 2 parameters; the `normalization` and `sigma`.
-"""
-samples = result.samples
-
-print("Parameter paths in the model which are used for filtering:")
-print(samples.model.paths)
-
-print("All parameters of the very first sample")
-print(samples.parameter_lists[0])
-
-samples = samples.without_paths(["gaussian.centre"])
-
-print(
- "All parameters of the very first sample (containing only the Gaussian normalization and sigma)."
-)
-print(samples.parameter_lists[0])
-
-"""
-Finish.
-"""
+"""
+Cookbook: Samples
+=================
+
+A non-linear search samples parameter space to find the global maximum log likelihood solution.
+
+The `Samples` object contains the history of the non-linear search, including the model parameters and log likelihood
+of every accepted sample.
+
+This cookbook provides an overview of using the samples.
+
+__Contents__
+
+ - Model Fit: Perform a simple model-fit to create a ``Samples`` object.
+ - Samples: The `Samples` object`, containing all non-linear samples (e.g. parameters, log likelihoods, etc.).
+ - Parameters: Accessing the parameters of the model from the samples.
+ - Figures of Merit: The log likelihood, log prior, log posterior and weight_list of every accepted sample.
+ - Instances: Returning instances of the model corresponding to a particular sample (e.g. the maximum log likelihood).
+ - Posterior / PDF: The median PDF model instance and PDF vectors of all model parameters via 1D marginalization.
+ - Errors: The errors on every parameter estimated from the PDF, computed via marginalized 1D PDFs at an input sigma.
+ - Samples Summary: A summary of the samples of the non-linear search (e.g. the maximum log likelihood model) which can
+ be faster to load than the full set of samples.
+ - Sample Instance: The model instance of any accepted sample.
+ - Search Plots: Plots of the non-linear search, for example a corner plot or 1D PDF of every parameter.
+ - Maximum Likelihood: The maximum log likelihood model value.
+ - Bayesian Evidence: The log evidence estimated via a nested sampling algorithm.
+ - Collection: Results created from models defined via a `Collection` object.
+ - Lists: Extracting results as Python lists instead of instances.
+ - Latex: Producing latex tables of results (e.g. for a paper).
+
+The following sections outline how to use advanced features of the results, which you may skip on a first read:
+
+ - Derived Quantities: Computing quantities and errors for quantities and parameters not included directly in the model.
+ - Derived Errors Manual (Advanced): Manually computing errors on derived quantities from the PDF of samples.
+ - Samples Filtering (Advanced): Filter the `Samples` object to only contain samples fulfilling certain criteria.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import autofit as af
+import autofit.plot as aplt
+
+from os import path
+import matplotlib.pyplot as plt
+import numpy as np
+
+"""
+__Model Fit__
+
+To illustrate results, we need to perform a model-fit in order to create a `Result` object.
+
+We do this below using the standard API and noisy 1D signal example, which you should be familiar with from other
+example scripts.
+
+Note that the `Gaussian` and `Analysis` classes come via the `af.ex` module, which contains example model components
+that are identical to those found throughout the examples.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+model = af.Model(af.ex.Gaussian)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+search = af.Emcee(
+ name="cookbook_result",
+ nwalkers=30,
+ nsteps=1000,
+ number_of_cores=1,
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+"""
+__Samples__
+
+The result contains a `Samples` object, which contains all samples of the non-linear search.
+
+Each sample corresponds to a set of model parameters that were evaluated and accepted by the non linear search,
+in this example `emcee.`
+
+This includes their log likelihoods, which are used for computing additional information about the model-fit,
+for example the error on every parameter.
+
+Our model-fit used the MCMC algorithm Emcee, so the `Samples` object returned is a `SamplesMCMC` object.
+"""
+samples = result.samples
+
+print("MCMC Samples: \n")
+print(samples)
+
+"""
+__Parameters__
+
+The parameters are stored as a list of lists, where:
+
+ - The outer list is the size of the total number of samples.
+ - The inner list is the size of the number of free parameters in the fit.
+"""
+samples = result.samples
+
+print("Sample 5's second parameter value (Gaussian -> normalization):")
+print(samples.parameter_lists[4][1])
+print("Sample 10`s third parameter value (Gaussian -> sigma)")
+print(samples.parameter_lists[9][2], "\n")
+
+"""
+__Figures of Merit__
+
+The `Samples` class contains the log likelihood, log prior, log posterior and weight_list of every accepted sample, where:
+
+- The `log_likelihood` is the value evaluated in the `log_likelihood_function`.
+
+- The `log_prior` encodes information on how parameter priors map log likelihood values to log posterior values.
+
+- The `log_posterior` is `log_likelihood + log_prior`.
+
+- The `weight` gives information on how samples are combined to estimate the posterior, which depends on type of search
+ used (for `Emcee` they are all 1's meaning they are weighted equally).
+
+Lets inspect these values for the tenth sample.
+"""
+print("log(likelihood), log(prior), log(posterior) and weight of the tenth sample.")
+print(samples.log_likelihood_list[9])
+print(samples.log_prior_list[9])
+print(samples.log_posterior_list[9])
+print(samples.weight_list[9])
+
+"""
+__Instances__
+
+Many results can be returned as an instance of the model, using the Python class structure of the model composition.
+
+For example, we can return the model parameters corresponding to the maximum log likelihood sample.
+
+The attributes of the `instance` (`centre`, `normalization` and `sigma`) have these names due to how we composed
+the `Gaussian` class via the `Model` above. They would be named structured and named differently if we hd
+used a `Collection` and different names.
+"""
+instance = samples.max_log_likelihood()
+
+print("Max Log Likelihood `Gaussian` Instance:")
+print("Centre = ", instance.centre)
+print("Normalization = ", instance.normalization)
+print("Sigma = ", instance.sigma, "\n")
+
+"""
+This makes it straight forward to plot the median PDF model:
+"""
+model_data = instance.model_data_from(xvalues=np.arange(data.shape[0]))
+
+plt.plot(range(data.shape[0]), data)
+plt.plot(range(data.shape[0]), model_data)
+plt.title("Illustrative model fit to 1D `Gaussian` profile data.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+__Posterior / PDF__
+
+The result contains the full posterior information of our non-linear search, which can be used for parameter
+estimation.
+
+PDF stands for "Probability Density Function" and it quantifies probability of each model parameter having values
+that are sampled. It therefore enables error estimation via a process called marginalization.
+
+The median pdf vector is available, which estimates every parameter via 1D marginalization of their PDFs.
+"""
+instance = samples.median_pdf()
+
+print("Median PDF `Gaussian` Instance:")
+print("Centre = ", instance.centre)
+print("Normalization = ", instance.normalization)
+print("Sigma = ", instance.sigma, "\n")
+
+"""
+__Errors__
+
+Methods for computing error estimates on all parameters are provided.
+
+This again uses 1D marginalization, now at an input sigma confidence limit.
+
+By inputting `sigma=3.0` margnialization find the values spanning 99.7% of 1D PDF. Changing this to `sigma=1.0`
+would give the errors at the 68.3% confidence limit.
+"""
+instance_upper_sigma = samples.errors_at_upper_sigma(sigma=3.0)
+instance_lower_sigma = samples.errors_at_lower_sigma(sigma=3.0)
+
+print("Upper Error values (at 3.0 sigma confidence):")
+print("Centre = ", instance_upper_sigma.centre)
+print("Normalization = ", instance_upper_sigma.normalization)
+print("Sigma = ", instance_upper_sigma.sigma, "\n")
+
+print("lower Error values (at 3.0 sigma confidence):")
+print("Centre = ", instance_lower_sigma.centre)
+print("Normalization = ", instance_lower_sigma.normalization)
+print("Sigma = ", instance_lower_sigma.sigma, "\n")
+
+"""
+They can also be returned at the values of the parameters at their error values.
+"""
+instance_upper_values = samples.values_at_upper_sigma(sigma=3.0)
+instance_lower_values = samples.values_at_lower_sigma(sigma=3.0)
+
+print("Upper Parameter values w/ error (at 3.0 sigma confidence):")
+print("Centre = ", instance_upper_values.centre)
+print("Normalization = ", instance_upper_values.normalization)
+print("Sigma = ", instance_upper_values.sigma, "\n")
+
+print("lower Parameter values w/ errors (at 3.0 sigma confidence):")
+print("Centre = ", instance_lower_values.centre)
+print("Normalization = ", instance_lower_values.normalization)
+print("Sigma = ", instance_lower_values.sigma, "\n")
+
+"""
+__Samples Summary__
+
+The samples summary contains a subset of results access via the `Samples`, for example the maximum likelihood model
+and parameter error estimates.
+
+Using the samples method above can be slow, as the quantities have to be computed from all non-linear search samples
+(e.g. computing errors requires that all samples are marginalized over). This information is stored directly in the
+samples summary and can therefore be accessed instantly.
+"""
+print(samples.summary().max_log_likelihood_sample)
+
+"""
+__Sample Instance__
+
+A non-linear search retains every model that is accepted during the model-fit.
+
+We can create an instance of any model -- below we create an instance of the last accepted model.
+"""
+instance = samples.from_sample_index(sample_index=-1)
+
+print("Gaussian Instance of last sample")
+print("Centre = ", instance.centre)
+print("Normalization = ", instance.normalization)
+print("Sigma = ", instance.sigma, "\n")
+
+"""
+__Search Plots__
+
+The Probability Density Functions (PDF's) of the results can be plotted using the non-linear search in-built
+visualization tools.
+
+This fit used `Emcee` therefore we use `corner.py` for visualization via the `aplt.corner_cornerpy` function.
+
+The `autofit_workspace/*/plots` folder illustrates other packages that can be used to make these plots using
+the standard output results formats (e.g. `GetDist.py`).
+"""
+aplt.corner_cornerpy(samples=result.samples)
+
+"""
+__Maximum Likelihood__
+
+The maximum log likelihood value of the model-fit can be estimated by simple taking the maximum of all log
+likelihoods of the samples.
+
+If different models are fitted to the same dataset, this value can be compared to determine which model provides
+the best fit (e.g. which model has the highest maximum likelihood)?
+"""
+print("Maximum Log Likelihood: \n")
+print(max(samples.log_likelihood_list))
+
+"""
+__Bayesian Evidence__
+
+If a nested sampling non-linear search is used, the evidence of the model is also available which enables Bayesian
+model comparison to be performed (given we are using Emcee, which is not a nested sampling algorithm, the log evidence
+is None).
+
+A full discussion of Bayesian model comparison is given in `autofit_workspace/*/features/bayes_model_comparison.py`.
+"""
+log_evidence = samples.log_evidence
+print(f"Log Evidence: {log_evidence}")
+
+"""
+__Collection__
+
+The examples correspond to a model where `af.Model(Gaussian)` was used to compose the model.
+
+Below, we illustrate how the results API slightly changes if we compose our model using a `Collection`:
+"""
+model = af.Collection(gaussian=af.ex.Gaussian, exponential=af.ex.Exponential)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+search = af.Emcee(
+ nwalkers=50,
+ nsteps=1000,
+ number_of_cores=1,
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+"""
+The `result.info` shows the result for the model with both a `Gaussian` and `Exponential` profile.
+"""
+print(result.info)
+
+"""
+Result instances again use the Python classes used to compose the model.
+
+However, because our fit uses a `Collection` the `instance` has attributes named according to the names given to the
+`Collection`, which above were `gaussian` and `exponential`.
+
+For complex models, with a large number of model components and parameters, this offers a readable API to interpret
+the results.
+"""
+samples = result.samples
+
+instance = samples.max_log_likelihood()
+
+print("Max Log Likelihood `Gaussian` Instance:")
+print("Centre = ", instance.gaussian.centre)
+print("Normalization = ", instance.gaussian.normalization)
+print("Sigma = ", instance.gaussian.sigma, "\n")
+
+print("Max Log Likelihood Exponential Instance:")
+print("Centre = ", instance.exponential.centre)
+print("Normalization = ", instance.exponential.normalization)
+print("Sigma = ", instance.exponential.rate, "\n")
+
+"""
+__Lists__
+
+All results can alternatively be returned as a 1D list of values, by passing `as_instance=False`:
+"""
+max_lh_list = samples.max_log_likelihood(as_instance=False)
+print("Max Log Likelihood Model Parameters: \n")
+print(max_lh_list, "\n\n")
+
+"""
+The list above does not tell us which values correspond to which parameters.
+
+The following quantities are available in the `Model`, where the order of their entries correspond to the parameters
+in the `ml_vector` above:
+
+ - `paths`: a list of tuples which give the path of every parameter in the `Model`.
+ - `parameter_names`: a list of shorthand parameter names derived from the `paths`.
+ - `parameter_labels`: a list of parameter labels used when visualizing non-linear search results (see below).
+
+For simple models like the one fitted in this tutorial, the quantities below are somewhat redundant. For the
+more complex models they are important for tracking the parameters of the model.
+"""
+model = samples.model
+
+print(model.paths)
+print(model.parameter_names)
+print(model.parameter_labels)
+print(model.model_component_and_parameter_names)
+print("\n")
+
+"""
+All the methods above are available as lists.
+"""
+instance = samples.median_pdf(as_instance=False)
+values_at_upper_sigma = samples.values_at_upper_sigma(sigma=3.0, as_instance=False)
+values_at_lower_sigma = samples.values_at_lower_sigma(sigma=3.0, as_instance=False)
+errors_at_upper_sigma = samples.errors_at_upper_sigma(sigma=3.0, as_instance=False)
+errors_at_lower_sigma = samples.errors_at_lower_sigma(sigma=3.0, as_instance=False)
+
+"""
+__Latex__
+
+If you are writing modeling results up in a paper, you can use inbuilt latex tools to create latex table code which
+you can copy to your .tex document.
+
+By combining this with the filtering tools below, specific parameters can be included or removed from the latex.
+
+Remember that the superscripts of a parameter are loaded from the config file `notation/label.yaml`, providing high
+levels of customization for how the parameter names appear in the latex table. This is especially useful if your model
+uses the same model components with the same parameter, which therefore need to be distinguished via superscripts.
+"""
+latex = af.text.Samples.latex(
+ samples=result.samples,
+ median_pdf_model=True,
+ sigma=3.0,
+ name_to_label=True,
+ include_name=True,
+ include_quickmath=True,
+ prefix="Example Prefix ",
+ suffix=" \\[-2pt]",
+)
+
+print(latex)
+
+"""
+__Derived Quantities__
+
+The parameters `centre`, `normalization` and `sigma` are the model parameters of the `Gaussian`. They are sampled
+directly by the non-linear search and we can therefore use the `Samples` object to easily determine their values and
+errors.
+
+Derived quantities (also called latent variables) are those which are not sampled directly by the non-linear search,
+but one may still wish to know their values and errors after the fit is complete. For example, what if we want the
+error on the full width half maximum (FWHM) of the Gaussian?
+
+This is achieved by adding them to the `compute_latent_variables` method of the `Analysis` class, which is called
+after the non-linear search has completed. The analysis cookbook illustrates how to do this.
+
+The example analysis used above includes a `compute_latent_variables` method that computes the FWHM of the Gaussian
+profile.
+
+This leads to a number of noteworthy outputs:
+
+ - A `latent.results` file is output to the results folder, which includes the value and error of all derived quantities
+ based on the non-linear search samples (in this example only the `fwhm`).
+
+ - A `latent/samples.csv` is output which lists every accepted sample's value of every derived quantity, which is again
+ analogous to the `samples.csv` file (in this example only the `fwhm`).
+
+ - A `latent/samples_summary.json` is output which acts analogously to `samples_summary.json` but for the derived
+ quantities of the model (in this example only the `fwhm`).
+
+Derived quantities are also accessible via the `Samples` object, following a similar API to the model parameters:
+"""
+latent = analysis.compute_latent_samples(result.samples)
+
+instance = latent.max_log_likelihood()
+
+print(f"Max Likelihood FWHM: {instance.gaussian.fwhm}")
+
+instance = latent.median_pdf()
+
+print(f"Median PDF FWHM {instance.gaussian.fwhm}")
+
+"""
+__Derived Errors Manual (Advanced)__
+
+The derived quantities decorator above provides a simple interface for computing the errors of a derived quantity and
+ensuring all results are easily inspected in the output results folder.
+
+However, you may wish to compute the errors of a derived quantity manually. For example, if it is a quantity that
+you did not decorate before performing the fit, or if it is computationally expensive to compute and you only want
+to compute it specific circumstances.
+
+We create the PDF of the derived quantity, the FWHM, manually, which we marginalize over using the same function
+we use to marginalize model parameters. We compute the FWHM of every accepted model sampled by the non-linear search
+and use this determine the PDF of the FWHM.
+
+When combining the FWHM's we weight each value by its `weight`. For Emcee, an MCMC algorithm, the weight of every
+sample is 1, but weights may take different values for other non-linear searches.
+
+In order to pass these samples to the function `marginalize`, which marginalizes over the PDF of the FWHM to compute
+its error, we also pass the weight list of the samples.
+"""
+fwhm_list = []
+
+for sample in samples.sample_list:
+ instance = sample.instance_for_model(model=samples.model)
+
+ sigma = instance.gaussian.sigma
+
+ fwhm = 2 * np.sqrt(2 * np.log(2)) * sigma
+
+ fwhm_list.append(fwhm)
+
+median_fwhm, lower_fwhm, upper_fwhm = af.marginalize(
+ parameter_list=fwhm_list, sigma=3.0, weight_list=samples.weight_list
+)
+
+print(f"FWHM = {median_fwhm} ({upper_fwhm} {lower_fwhm}")
+
+"""
+The calculation above could be computationally expensive, if there are many samples and the derived quantity is
+slow to compute.
+
+An alternative approach, which will provide comparable accuracy provided enough draws are used, is to sample
+points randomy from the PDF of the model and use these to compute the derived quantity.
+
+Draws are from the PDF of the model, so the weights of the samples are accounted for and we therefore do not
+pass them to the `marginalize` function (it essentially treats all samples as having equal weight).
+
+TRY AND EXCEPT INCLUDED TO FIX BUG, NEED TO SOLVE IN FUTURE.
+"""
+try:
+ random_draws = 50
+
+ fwhm_list = []
+
+ for i in range(random_draws):
+ instance = samples.draw_randomly_via_pdf()
+
+ sigma = instance.gaussian.sigma
+
+ fwhm = 2 * np.sqrt(2 * np.log(2)) * sigma
+
+ fwhm_list.append(fwhm)
+
+ median_fwhm, lower_fwhm, upper_fwhm = af.marginalize(
+ parameter_list=fwhm_list, sigma=3.0, weight_list=samples.weight_list
+ )
+
+ print(f"fwhm = {median_fwhm} ({upper_fwhm} {lower_fwhm}")
+
+except ValueError:
+ pass
+
+"""
+__Samples Filtering (Advanced)__
+
+Our samples object has the results for all three parameters in our model. However, we might only be interested in the
+results of a specific parameter.
+
+The basic form of filtering specifies parameters via their path, which was printed above via the model and is printed
+again below.
+"""
+samples = result.samples
+
+print("Parameter paths in the model which are used for filtering:")
+print(samples.model.paths)
+
+print("All parameters of the very first sample")
+print(samples.parameter_lists[0])
+
+samples = samples.with_paths([("gaussian", "centre")])
+
+print("All parameters of the very first sample (containing only the Gaussian centre.")
+print(samples.parameter_lists[0])
+
+print("Maximum Log Likelihood Model Instances (containing only the Gaussian centre):\n")
+print(samples.max_log_likelihood(as_instance=False))
+
+"""
+We specified each path as a list of tuples of strings.
+
+This is how the source code internally stores the path to different components of the model, but it is not
+consistent with the API used to compose a model.
+
+We can alternatively use the following API:
+"""
+samples = result.samples
+
+samples = samples.with_paths(["gaussian.centre"])
+
+print("All parameters of the very first sample (containing only the Gaussian centre).")
+print(samples.parameter_lists[0])
+
+"""
+We filtered the `Samples` above by asking for all parameters which included the path ("gaussian", "centre").
+
+We can alternatively filter the `Samples` object by removing all parameters with a certain path. Below, we remove
+the Gaussian's `centre` to be left with 2 parameters; the `normalization` and `sigma`.
+"""
+samples = result.samples
+
+print("Parameter paths in the model which are used for filtering:")
+print(samples.model.paths)
+
+print("All parameters of the very first sample")
+print(samples.parameter_lists[0])
+
+samples = samples.without_paths(["gaussian.centre"])
+
+print(
+ "All parameters of the very first sample (containing only the Gaussian normalization and sigma)."
+)
+print(samples.parameter_lists[0])
+
+"""
+Finish.
+"""
diff --git a/scripts/cookbooks/search.py b/scripts/cookbooks/search.py
index ec051153..bd6686a8 100644
--- a/scripts/cookbooks/search.py
+++ b/scripts/cookbooks/search.py
@@ -1,498 +1,494 @@
-"""
-Cookbook: Searches
-==================
-
-This cookbook provides an overview of the non-linear searches available in **PyAutoFit**, and how to use them.
-
-__Contents__
-
-It first covers standard options available for all non-linear searches:
-
- - Example Fit: A simple example of a non-linear search to remind us how it works.
- - Output To Hard-Disk: Output results to hard-disk so they can be inspected and used to restart a crashed search.
- - Output Customization: Customize the output of a non-linear search to hard-disk.
- - Unique Identifier: Ensure results are output in unique folders, so they do not overwrite each other.
- - Iterations Per Update: Control how often non-linear searches output results to hard-disk.
- - Parallelization: Use parallel processing to speed up the sampling of parameter space.
- - Plots: Perform non-linear search specific visualization using their in-built visualization tools.
- - Start Point: Manually specify the start point of a non-linear search, or sample a specific region of parameter space.
-
-It then provides example code for using every search:
-
- - Emcee (MCMC): The Emcee ensemble sampler MCMC.
- - Zeus (MCMC): The Zeus ensemble sampler MCMC.
- - DynestyDynamic (Nested Sampling): The Dynesty dynamic nested sampler.
- - DynestyStatic (Nested Sampling): The Dynesty static nested sampler.
- - UltraNest (Nested Sampling): The UltraNest nested sampler.
- - PySwarmsGlobal (Particle Swarm Optimization): The global PySwarms particle swarm optimization
- - PySwarmsLocal (Particle Swarm Optimization): The local PySwarms particle swarm optimization.
- - LBFGS: The L-BFGS scipy optimization.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import numpy as np
-from os import path
-
-import autofit as af
-import autofit.plot as aplt
-
-"""
-__Example Fit__
-
-An example of how to use a `search` to fit a model to data is given in other example scripts, but is shown below
-for completeness.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-model = af.Model(af.ex.Gaussian)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-"""
-It is this line, where the command `af.Emcee()` can be swapped out for the examples provided throughout this
-cookbook to use different non-linear searches.
-"""
-search = af.Emcee()
-
-result = search.fit(model=model, analysis=analysis)
-
-"""
-__Output To Hard-Disk__
-
-By default, a non-linear search does not output its results to hard-disk and its results can only be inspected
-in Python via the `result` object.
-
-However, the results of any non-linear search can be output to hard-disk by passing the `name` and / or `path_prefix`
-attributes, which are used to name files and output the results to a folder on your hard-disk.
-
-The benefits of doing this include:
-
-- Inspecting results via folders on your computer can be more efficient than using a Jupyter Notebook.
-- Results are output on-the-fly, making it possible to check that a fit i progressing as expected mid way through.
-- Additional information about a fit (e.g. visualization) is output.
-- Unfinished runs can be resumed from where they left off if they are terminated.
-- On high performance super computers which use a batch system, results must be output in this way.
-
-If you run a search with outputting to hard-disk enabled, checkout the `files` folder which includes files such as
-`samples.csv` (every accepted samples with their parameters and log likelihoods), `model.json` (a .json
-file describing the model) and `search.json` (a .json file describing the search).
-
-These outputs are fully described in the scientific workflow example.
-"""
-search = af.Emcee(path_prefix=path.join("folder_0", "folder_1"), name="example_mcmc")
-
-"""
-__Output Customization__
-
-For large model fitting problems outputs may use up a lot of hard-disk space, therefore full customization of the
-outputs is supported.
-
-This is controlled by the `output.yaml` config file found in the `config` folder of the workspace. This file contains
-a full description of all customization options.
-
-A few examples of the options available include:
-
-- Control over every file which is output to the `files` folder (e.g. `model.json`, `samples.csv`, etc.).
-
-- For the `samples.csv` file, all samples with a weight below a certain value can be automatically removed.
-
-- Customization of the `samples_summary.json` file, which summarises the results of the model-fit (e.g. the maximum
- log likelihood model, the median PDF model and 3 sigma error). These results are computed using the full set of
- samples, ensuring samples removal via a weight cut does not impact the results.
-
-In many use cases, the `samples.csv` takes up the significant majority of the hard-disk space, which for large-scale
-model-fitting problems can exceed gigabytes and be prohibitive to the analysis.
-
-Careful customization of the `output.yaml` file enables a workflow where the `samples.csv` file is never output,
-but all important information is output in the `samples_summary.json` file using the full samples to compute all
-results to high numerical accuracy.
-
-__Unique Identifier__
-
-Results are output to a folder which is a collection of random characters, which is the 'unique_identifier' of
-the model-fit. This identifier is generated based on the model fitted and search used, such that an identical
-combination of model and search generates the same identifier.
-
-This ensures that rerunning an identical fit will use the existing results to resume the model-fit. In contrast, if
-you change the model or search, a new unique identifier will be generated, ensuring that the model-fit results are
-output into a separate folder.
-
-A `unique_tag` can be input into a search, which customizes the unique identifier based on the string you provide.
-For example, if you are performing many fits to different datasets, using an identical model and search, you may
-wish to provide a unique tag for each dataset such that the model-fit results are output into a different folder.
-"""
-search = af.Emcee(unique_tag="example_tag")
-
-"""
-__Iterations Per Update__
-
-If results are output to hard-disk, this occurs every `iterations_per_full_update` number of iterations.
-
-For certain problems, you may want this value to be low, to inspect the results of the model-fit on a regular basis.
-This is especially true if the time it takes for your non-linear search to perform an iteration by evaluating the
-log likelihood is long (e.g. > 1s) and your model-fit often goes to incorrect solutions that you want to monitor.
-
-For other problems, you may want to increase this value, to avoid spending lots of time outputting the results to
-hard-disk. This is especially true if the time it takes for your non-linear search to perform an iteration by
-evaluating the log likelihood is fast (e.g. < 0.1s) and you are confident your model-fit will find the global
-maximum solution given enough iterations.
-
-**If the iteration per update is too low, the model-fit may be significantly slowed down by the time it takes to
-output results and visualization frequently to hard-disk. If your fit is consistent displaying a log saying that it
-is outputting results, try increasing this value to ensure the model-fit runs efficiently.**
-"""
-search = af.Emcee(iterations_per_full_update=1000)
-
-"""
-__Parallelization__
-
-Many searches support parallelization using the Python ``multiprocessing`` module.
-
-This distributes the non-linear search analysis over multiple CPU's, speeding up the run-time roughly by the number
-of CPUs used.
-
-To enable parallelization, input a `number_of_cores` greater than 1. You should aim not to exceed the number of
-physical cores in your computer, as using more cores than exist may actually slow down the non-linear search.
-"""
-search = af.Emcee(number_of_cores=4)
-
-"""
-__Plots__
-
-Every non-linear search supported by **PyAutoFit** has a dedicated `plotter` class that allows the results of the
-model-fit to be plotted and inspected.
-
-This uses that search's in-built visualization libraries, which are fully described in the `plot` package of the
-workspace.
-
-For example, `Emcee` results can be plotted using the `aplt.corner_cornerpy` function as follows.
-
-Checkout the `plot` package for a complete description of the plots that can be made for a given search.
-"""
-samples = result.samples
-
-aplt.corner_cornerpy(
- samples=samples,
- bins=20,
- range=None,
- color="k",
- hist_bin_factor=1,
- smooth=None,
- smooth1d=None,
- label_kwargs=None,
- titles=None,
- show_titles=False,
- title_fmt=".2f",
- title_kwargs=None,
- truths=None,
- truth_color="#4682b4",
- scale_hist=False,
- quantiles=None,
- verbose=False,
- fig=None,
- max_n_ticks=5,
- top_ticks=False,
- use_math_text=False,
- reverse=False,
- labelpad=0.0,
- hist_kwargs=None,
- group="posterior",
- var_names=None,
- filter_vars=None,
- coords=None,
- divergences=False,
- divergences_kwargs=None,
- labeller=None,
-)
-
-"""
-The Python library `GetDist `_ can also be used to create plots of the
-results.
-
-This is described in the `plot` package of the workspace.
-
-__Start Point__
-
-For maximum likelihood estimator (MLE) and Markov Chain Monte Carlo (MCMC) non-linear searches, parameter space
-sampling is built around having a "location" in parameter space.
-
-This could simply be the parameters of the current maximum likelihood model in an MLE fit, or the locations of many
-walkers in parameter space (e.g. MCMC).
-
-For many model-fitting problems, we may have an expectation of where correct solutions lie in parameter space and
-therefore want our non-linear search to start near that location of parameter space. Alternatively, we may want to
-sample a specific region of parameter space, to determine what solutions look like there.
-
-The start-point API allows us to do this, by manually specifying the start-point of an MLE fit or the start-point of
-the walkers in an MCMC fit. Because nested sampling draws from priors, it cannot use the start-point API.
-
-We now define the start point of certain parameters in the model as follows.
-"""
-initializer = af.InitializerParamBounds(
- {
- model.centre: (49.0, 51.0),
- model.normalization: (4.0, 6.0),
- model.sigma: (1.0, 2.0),
- }
-)
-
-"""
-Similar behaviour can be achieved by customizing the priors of a model-fit. We could place `TruncatedGaussianPrior`'s
-centred on the regions of parameter space we want to sample, or we could place tight `UniformPrior`'s on regions
-of parameter space we believe the correct answer lies.
-
-The downside of using priors is that our priors have a direct influence on the parameters we infer and the size
-of the inferred parameter errors. By using priors to control the location of our model-fit, we therefore risk
-inferring a non-representative model.
-
-For users more familiar with statistical inference, adjusting ones priors in the way described above leads to
-changes in the posterior, which therefore impacts the model inferred.
-
-__Emcee (MCMC)__
-
-The Emcee sampler is a Markov Chain Monte Carlo (MCMC) Ensemble sampler. It is a Python implementation of the
-`Goodman & Weare `_ affine-invariant ensemble MCMC sampler.
-
-Information about Emcee can be found at the following links:
-
- - https://github.com/dfm/emcee
- - https://emcee.readthedocs.io/en/stable/
-
-The following workspace example shows examples of fitting data with Emcee and plotting the results.
-
-- `autofit_workspace/notebooks/searches/mcmc/Emcee.ipynb`
-- `autofit_workspace/notebooks/plot/MCMCPlotter.ipynb`
-
-The following code shows how to use Emcee with all available options.
-"""
-search = af.Emcee(
- nwalkers=30,
- nsteps=1000,
- initializer=af.InitializerBall(lower_limit=0.49, upper_limit=0.51),
- auto_correlations_settings=af.AutoCorrelationsSettings(
- check_for_convergence=True,
- check_size=100,
- required_length=50,
- change_threshold=0.01,
- ),
-)
-
-"""
-__Zeus (MCMC)__
-
-The Zeus sampler is a Markov Chain Monte Carlo (MCMC) Ensemble sampler.
-
-Information about Zeus can be found at the following links:
-
- - https://github.com/minaskar/zeus
- - https://zeus-mcmc.readthedocs.io/en/latest/
-"""
-search = af.Zeus(
- nwalkers=30,
- nsteps=1001,
- initializer=af.InitializerBall(lower_limit=0.49, upper_limit=0.51),
- auto_correlations_settings=af.AutoCorrelationsSettings(
- check_for_convergence=True,
- check_size=100,
- required_length=50,
- change_threshold=0.01,
- ),
- tune=False,
- tolerance=0.05,
- patience=5,
- maxsteps=10000,
- mu=1.0,
- maxiter=10000,
- vectorize=False,
- check_walkers=True,
- shuffle_ensemble=True,
- light_mode=False,
-)
-
-"""
-__DynestyDynamic (Nested Sampling)__
-
-The DynestyDynamic sampler is a Dynamic Nested Sampling algorithm. It is a Python implementation of the
-`Speagle `_ algorithm.
-
-Information about Dynesty can be found at the following links:
-
- - https://github.com/joshspeagle/dynesty
- - https://dynesty.readthedocs.io/en/latest/
-"""
-search = af.DynestyDynamic(
- nlive=50,
- bound="multi",
- sample="auto",
- bootstrap=None,
- enlarge=None,
- update_interval=None,
- walks=25,
- facc=0.5,
- slices=5,
- fmove=0.9,
- max_move=100,
-)
-
-"""
-__DynestyStatic (Nested Sampling)__
-
-The DynestyStatic sampler is a Static Nested Sampling algorithm. It is a Python implementation of the
-`Speagle `_ algorithm.
-
-Information about Dynesty can be found at the following links:
-
- - https://github.com/joshspeagle/dynesty
- - https://dynesty.readthedocs.io/en/latest/
-"""
-search = af.DynestyStatic(
- nlive=50,
- bound="multi",
- sample="auto",
- bootstrap=None,
- enlarge=None,
- update_interval=None,
- walks=25,
- facc=0.5,
- slices=5,
- fmove=0.9,
- max_move=100,
-)
-
-"""
-__UltraNest (Nested Sampling)__
-
-The UltraNest sampler is a Nested Sampling algorithm. It is a Python implementation of the
-`Buchner `_ algorithm.
-
-UltraNest is an optional requirement and must be installed manually via the command `pip install ultranest`.
-It is optional as it has certain dependencies which are generally straight forward to install (e.g. Cython).
-
-Information about UltraNest can be found at the following links:
-
- - https://github.com/JohannesBuchner/UltraNest
- - https://johannesbuchner.github.io/UltraNest/readme.html
-"""
-search = af.UltraNest(
- resume=True,
- run_num=None,
- num_test_samples=2,
- draw_multiple=True,
- num_bootstraps=30,
- vectorized=False,
- ndraw_min=128,
- ndraw_max=65536,
- storage_backend="hdf5",
- warmstart_max_tau=-1,
- update_interval_volume_fraction=0.8,
- update_interval_ncall=None,
- log_interval=None,
- show_status=True,
- viz_callback="auto",
- dlogz=0.5,
- dKL=0.5,
- frac_remain=0.01,
- Lepsilon=0.001,
- min_ess=400,
- max_iters=None,
- max_ncalls=None,
- max_num_improvement_loops=-1,
- min_num_live_points=50,
- cluster_num_live_points=40,
- insertion_test_window=10,
- insertion_test_zscore_threshold=2,
- stepsampler_cls="RegionMHSampler",
- nsteps=11,
-)
-
-"""
-__PySwarmsGlobal__
-
-The PySwarmsGlobal sampler is a Global Optimization algorithm. It is a Python implementation of the
-`Bratley `_ algorithm.
-
-Information about PySwarms can be found at the following links:
-
- - https://github.com/ljvmiranda921/pyswarms
- - https://pyswarms.readthedocs.io/en/latest/index.html
- - https://pyswarms.readthedocs.io/en/latest/api/pyswarms.single.html#module-pyswarms.single.global_best
-"""
-search = af.PySwarmsGlobal(
- n_particles=50,
- iters=1000,
- cognitive=0.5,
- social=0.3,
- inertia=0.9,
- ftol=-np.inf,
-)
-
-"""
-__PySwarmsLocal__
-
-The PySwarmsLocal sampler is a Local Optimization algorithm. It is a Python implementation of the
-`Bratley `_ algorithm.
-
-Information about PySwarms can be found at the following links:
-
- - https://github.com/ljvmiranda921/pyswarms
- - https://pyswarms.readthedocs.io/en/latest/index.html
- - https://pyswarms.readthedocs.io/en/latest/api/pyswarms.single.html#module-pyswarms.single.global_best
-"""
-search = af.PySwarmsLocal(
- n_particles=50,
- iters=1000,
- cognitive=0.5,
- social=0.3,
- inertia=0.9,
- number_of_k_neighbors=3,
- minkowski_p_norm=2,
- ftol=-np.inf,
-)
-
-"""
-__LBFGS__
-
-The LBFGS sampler is a Local Optimization algorithm. It is a Python implementation of the scipy.optimize.lbfgs
-algorithm.
-
-Information about the L-BFGS method can be found at the following links:
-
- - https://docs.scipy.org/doc/scipy/reference/optimize.minimize-lbfgsb.html
-"""
-search = af.LBFGS(
- tol=None,
- disp=None,
- maxcor=10,
- ftol=2.220446049250313e-09,
- gtol=1e-05,
- eps=1e-08,
- maxfun=15000,
- maxiter=15000,
- iprint=-1,
- maxls=20,
-)
+"""
+Cookbook: Searches
+==================
+
+This cookbook provides an overview of the non-linear searches available in **PyAutoFit**, and how to use them.
+
+__Contents__
+
+It first covers standard options available for all non-linear searches:
+
+ - Example Fit: A simple example of a non-linear search to remind us how it works.
+ - Output To Hard-Disk: Output results to hard-disk so they can be inspected and used to restart a crashed search.
+ - Output Customization: Customize the output of a non-linear search to hard-disk.
+ - Unique Identifier: Ensure results are output in unique folders, so they do not overwrite each other.
+ - Iterations Per Update: Control how often non-linear searches output results to hard-disk.
+ - Parallelization: Use parallel processing to speed up the sampling of parameter space.
+ - Plots: Perform non-linear search specific visualization using their in-built visualization tools.
+ - Start Point: Manually specify the start point of a non-linear search, or sample a specific region of parameter space.
+
+It then provides example code for using every search:
+
+ - Emcee (MCMC): The Emcee ensemble sampler MCMC.
+ - Zeus (MCMC): The Zeus ensemble sampler MCMC.
+ - DynestyDynamic (Nested Sampling): The Dynesty dynamic nested sampler.
+ - DynestyStatic (Nested Sampling): The Dynesty static nested sampler.
+ - UltraNest (Nested Sampling): The UltraNest nested sampler.
+ - PySwarmsGlobal (Particle Swarm Optimization): The global PySwarms particle swarm optimization
+ - PySwarmsLocal (Particle Swarm Optimization): The local PySwarms particle swarm optimization.
+ - LBFGS: The L-BFGS scipy optimization.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import numpy as np
+from os import path
+
+import autofit as af
+import autofit.plot as aplt
+
+"""
+__Example Fit__
+
+An example of how to use a `search` to fit a model to data is given in other example scripts, but is shown below
+for completeness.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+model = af.Model(af.ex.Gaussian)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+"""
+It is this line, where the command `af.Emcee()` can be swapped out for the examples provided throughout this
+cookbook to use different non-linear searches.
+"""
+search = af.Emcee()
+
+result = search.fit(model=model, analysis=analysis)
+
+"""
+__Output To Hard-Disk__
+
+By default, a non-linear search does not output its results to hard-disk and its results can only be inspected
+in Python via the `result` object.
+
+However, the results of any non-linear search can be output to hard-disk by passing the `name` and / or `path_prefix`
+attributes, which are used to name files and output the results to a folder on your hard-disk.
+
+The benefits of doing this include:
+
+- Inspecting results via folders on your computer can be more efficient than using a Jupyter Notebook.
+- Results are output on-the-fly, making it possible to check that a fit i progressing as expected mid way through.
+- Additional information about a fit (e.g. visualization) is output.
+- Unfinished runs can be resumed from where they left off if they are terminated.
+- On high performance super computers which use a batch system, results must be output in this way.
+
+If you run a search with outputting to hard-disk enabled, checkout the `files` folder which includes files such as
+`samples.csv` (every accepted samples with their parameters and log likelihoods), `model.json` (a .json
+file describing the model) and `search.json` (a .json file describing the search).
+
+These outputs are fully described in the scientific workflow example.
+"""
+search = af.Emcee(path_prefix=path.join("folder_0", "folder_1"), name="example_mcmc")
+
+"""
+__Output Customization__
+
+For large model fitting problems outputs may use up a lot of hard-disk space, therefore full customization of the
+outputs is supported.
+
+This is controlled by the `output.yaml` config file found in the `config` folder of the workspace. This file contains
+a full description of all customization options.
+
+A few examples of the options available include:
+
+- Control over every file which is output to the `files` folder (e.g. `model.json`, `samples.csv`, etc.).
+
+- For the `samples.csv` file, all samples with a weight below a certain value can be automatically removed.
+
+- Customization of the `samples_summary.json` file, which summarises the results of the model-fit (e.g. the maximum
+ log likelihood model, the median PDF model and 3 sigma error). These results are computed using the full set of
+ samples, ensuring samples removal via a weight cut does not impact the results.
+
+In many use cases, the `samples.csv` takes up the significant majority of the hard-disk space, which for large-scale
+model-fitting problems can exceed gigabytes and be prohibitive to the analysis.
+
+Careful customization of the `output.yaml` file enables a workflow where the `samples.csv` file is never output,
+but all important information is output in the `samples_summary.json` file using the full samples to compute all
+results to high numerical accuracy.
+
+__Unique Identifier__
+
+Results are output to a folder which is a collection of random characters, which is the 'unique_identifier' of
+the model-fit. This identifier is generated based on the model fitted and search used, such that an identical
+combination of model and search generates the same identifier.
+
+This ensures that rerunning an identical fit will use the existing results to resume the model-fit. In contrast, if
+you change the model or search, a new unique identifier will be generated, ensuring that the model-fit results are
+output into a separate folder.
+
+A `unique_tag` can be input into a search, which customizes the unique identifier based on the string you provide.
+For example, if you are performing many fits to different datasets, using an identical model and search, you may
+wish to provide a unique tag for each dataset such that the model-fit results are output into a different folder.
+"""
+search = af.Emcee(unique_tag="example_tag")
+
+"""
+__Iterations Per Update__
+
+If results are output to hard-disk, this occurs every `iterations_per_full_update` number of iterations.
+
+For certain problems, you may want this value to be low, to inspect the results of the model-fit on a regular basis.
+This is especially true if the time it takes for your non-linear search to perform an iteration by evaluating the
+log likelihood is long (e.g. > 1s) and your model-fit often goes to incorrect solutions that you want to monitor.
+
+For other problems, you may want to increase this value, to avoid spending lots of time outputting the results to
+hard-disk. This is especially true if the time it takes for your non-linear search to perform an iteration by
+evaluating the log likelihood is fast (e.g. < 0.1s) and you are confident your model-fit will find the global
+maximum solution given enough iterations.
+
+**If the iteration per update is too low, the model-fit may be significantly slowed down by the time it takes to
+output results and visualization frequently to hard-disk. If your fit is consistent displaying a log saying that it
+is outputting results, try increasing this value to ensure the model-fit runs efficiently.**
+"""
+search = af.Emcee(iterations_per_full_update=1000)
+
+"""
+__Parallelization__
+
+Many searches support parallelization using the Python ``multiprocessing`` module.
+
+This distributes the non-linear search analysis over multiple CPU's, speeding up the run-time roughly by the number
+of CPUs used.
+
+To enable parallelization, input a `number_of_cores` greater than 1. You should aim not to exceed the number of
+physical cores in your computer, as using more cores than exist may actually slow down the non-linear search.
+"""
+search = af.Emcee(number_of_cores=4)
+
+"""
+__Plots__
+
+Every non-linear search supported by **PyAutoFit** has a dedicated `plotter` class that allows the results of the
+model-fit to be plotted and inspected.
+
+This uses that search's in-built visualization libraries, which are fully described in the `plot` package of the
+workspace.
+
+For example, `Emcee` results can be plotted using the `aplt.corner_cornerpy` function as follows.
+
+Checkout the `plot` package for a complete description of the plots that can be made for a given search.
+"""
+samples = result.samples
+
+aplt.corner_cornerpy(
+ samples=samples,
+ bins=20,
+ range=None,
+ color="k",
+ hist_bin_factor=1,
+ smooth=None,
+ smooth1d=None,
+ label_kwargs=None,
+ titles=None,
+ show_titles=False,
+ title_fmt=".2f",
+ title_kwargs=None,
+ truths=None,
+ truth_color="#4682b4",
+ scale_hist=False,
+ quantiles=None,
+ verbose=False,
+ fig=None,
+ max_n_ticks=5,
+ top_ticks=False,
+ use_math_text=False,
+ reverse=False,
+ labelpad=0.0,
+ hist_kwargs=None,
+ group="posterior",
+ var_names=None,
+ filter_vars=None,
+ coords=None,
+ divergences=False,
+ divergences_kwargs=None,
+ labeller=None,
+)
+
+"""
+The Python library `GetDist `_ can also be used to create plots of the
+results.
+
+This is described in the `plot` package of the workspace.
+
+__Start Point__
+
+For maximum likelihood estimator (MLE) and Markov Chain Monte Carlo (MCMC) non-linear searches, parameter space
+sampling is built around having a "location" in parameter space.
+
+This could simply be the parameters of the current maximum likelihood model in an MLE fit, or the locations of many
+walkers in parameter space (e.g. MCMC).
+
+For many model-fitting problems, we may have an expectation of where correct solutions lie in parameter space and
+therefore want our non-linear search to start near that location of parameter space. Alternatively, we may want to
+sample a specific region of parameter space, to determine what solutions look like there.
+
+The start-point API allows us to do this, by manually specifying the start-point of an MLE fit or the start-point of
+the walkers in an MCMC fit. Because nested sampling draws from priors, it cannot use the start-point API.
+
+We now define the start point of certain parameters in the model as follows.
+"""
+initializer = af.InitializerParamBounds(
+ {
+ model.centre: (49.0, 51.0),
+ model.normalization: (4.0, 6.0),
+ model.sigma: (1.0, 2.0),
+ }
+)
+
+"""
+Similar behaviour can be achieved by customizing the priors of a model-fit. We could place `TruncatedGaussianPrior`'s
+centred on the regions of parameter space we want to sample, or we could place tight `UniformPrior`'s on regions
+of parameter space we believe the correct answer lies.
+
+The downside of using priors is that our priors have a direct influence on the parameters we infer and the size
+of the inferred parameter errors. By using priors to control the location of our model-fit, we therefore risk
+inferring a non-representative model.
+
+For users more familiar with statistical inference, adjusting ones priors in the way described above leads to
+changes in the posterior, which therefore impacts the model inferred.
+
+__Emcee (MCMC)__
+
+The Emcee sampler is a Markov Chain Monte Carlo (MCMC) Ensemble sampler. It is a Python implementation of the
+`Goodman & Weare `_ affine-invariant ensemble MCMC sampler.
+
+Information about Emcee can be found at the following links:
+
+ - https://github.com/dfm/emcee
+ - https://emcee.readthedocs.io/en/stable/
+
+The following workspace example shows examples of fitting data with Emcee and plotting the results.
+
+- `autofit_workspace/notebooks/searches/mcmc/Emcee.ipynb`
+- `autofit_workspace/notebooks/plot/MCMCPlotter.ipynb`
+
+The following code shows how to use Emcee with all available options.
+"""
+search = af.Emcee(
+ nwalkers=30,
+ nsteps=1000,
+ initializer=af.InitializerBall(lower_limit=0.49, upper_limit=0.51),
+ auto_correlations_settings=af.AutoCorrelationsSettings(
+ check_for_convergence=True,
+ check_size=100,
+ required_length=50,
+ change_threshold=0.01,
+ ),
+)
+
+"""
+__Zeus (MCMC)__
+
+The Zeus sampler is a Markov Chain Monte Carlo (MCMC) Ensemble sampler.
+
+Information about Zeus can be found at the following links:
+
+ - https://github.com/minaskar/zeus
+ - https://zeus-mcmc.readthedocs.io/en/latest/
+"""
+search = af.Zeus(
+ nwalkers=30,
+ nsteps=1001,
+ initializer=af.InitializerBall(lower_limit=0.49, upper_limit=0.51),
+ auto_correlations_settings=af.AutoCorrelationsSettings(
+ check_for_convergence=True,
+ check_size=100,
+ required_length=50,
+ change_threshold=0.01,
+ ),
+ tune=False,
+ tolerance=0.05,
+ patience=5,
+ maxsteps=10000,
+ mu=1.0,
+ maxiter=10000,
+ vectorize=False,
+ check_walkers=True,
+ shuffle_ensemble=True,
+ light_mode=False,
+)
+
+"""
+__DynestyDynamic (Nested Sampling)__
+
+The DynestyDynamic sampler is a Dynamic Nested Sampling algorithm. It is a Python implementation of the
+`Speagle `_ algorithm.
+
+Information about Dynesty can be found at the following links:
+
+ - https://github.com/joshspeagle/dynesty
+ - https://dynesty.readthedocs.io/en/latest/
+"""
+search = af.DynestyDynamic(
+ nlive=50,
+ bound="multi",
+ sample="auto",
+ bootstrap=None,
+ enlarge=None,
+ update_interval=None,
+ walks=25,
+ facc=0.5,
+ slices=5,
+ fmove=0.9,
+ max_move=100,
+)
+
+"""
+__DynestyStatic (Nested Sampling)__
+
+The DynestyStatic sampler is a Static Nested Sampling algorithm. It is a Python implementation of the
+`Speagle `_ algorithm.
+
+Information about Dynesty can be found at the following links:
+
+ - https://github.com/joshspeagle/dynesty
+ - https://dynesty.readthedocs.io/en/latest/
+"""
+search = af.DynestyStatic(
+ nlive=50,
+ bound="multi",
+ sample="auto",
+ bootstrap=None,
+ enlarge=None,
+ update_interval=None,
+ walks=25,
+ facc=0.5,
+ slices=5,
+ fmove=0.9,
+ max_move=100,
+)
+
+"""
+__UltraNest (Nested Sampling)__
+
+The UltraNest sampler is a Nested Sampling algorithm. It is a Python implementation of the
+`Buchner `_ algorithm.
+
+UltraNest is an optional requirement and must be installed manually via the command `pip install ultranest`.
+It is optional as it has certain dependencies which are generally straight forward to install (e.g. Cython).
+
+Information about UltraNest can be found at the following links:
+
+ - https://github.com/JohannesBuchner/UltraNest
+ - https://johannesbuchner.github.io/UltraNest/readme.html
+"""
+search = af.UltraNest(
+ resume=True,
+ run_num=None,
+ num_test_samples=2,
+ draw_multiple=True,
+ num_bootstraps=30,
+ vectorized=False,
+ ndraw_min=128,
+ ndraw_max=65536,
+ storage_backend="hdf5",
+ warmstart_max_tau=-1,
+ update_interval_volume_fraction=0.8,
+ update_interval_ncall=None,
+ log_interval=None,
+ show_status=True,
+ viz_callback="auto",
+ dlogz=0.5,
+ dKL=0.5,
+ frac_remain=0.01,
+ Lepsilon=0.001,
+ min_ess=400,
+ max_iters=None,
+ max_ncalls=None,
+ max_num_improvement_loops=-1,
+ min_num_live_points=50,
+ cluster_num_live_points=40,
+ insertion_test_window=10,
+ insertion_test_zscore_threshold=2,
+ stepsampler_cls="RegionMHSampler",
+ nsteps=11,
+)
+
+"""
+__PySwarmsGlobal__
+
+The PySwarmsGlobal sampler is a Global Optimization algorithm. It is a Python implementation of the
+`Bratley `_ algorithm.
+
+Information about PySwarms can be found at the following links:
+
+ - https://github.com/ljvmiranda921/pyswarms
+ - https://pyswarms.readthedocs.io/en/latest/index.html
+ - https://pyswarms.readthedocs.io/en/latest/api/pyswarms.single.html#module-pyswarms.single.global_best
+"""
+search = af.PySwarmsGlobal(
+ n_particles=50,
+ iters=1000,
+ cognitive=0.5,
+ social=0.3,
+ inertia=0.9,
+ ftol=-np.inf,
+)
+
+"""
+__PySwarmsLocal__
+
+The PySwarmsLocal sampler is a Local Optimization algorithm. It is a Python implementation of the
+`Bratley `_ algorithm.
+
+Information about PySwarms can be found at the following links:
+
+ - https://github.com/ljvmiranda921/pyswarms
+ - https://pyswarms.readthedocs.io/en/latest/index.html
+ - https://pyswarms.readthedocs.io/en/latest/api/pyswarms.single.html#module-pyswarms.single.global_best
+"""
+search = af.PySwarmsLocal(
+ n_particles=50,
+ iters=1000,
+ cognitive=0.5,
+ social=0.3,
+ inertia=0.9,
+ number_of_k_neighbors=3,
+ minkowski_p_norm=2,
+ ftol=-np.inf,
+)
+
+"""
+__LBFGS__
+
+The LBFGS sampler is a Local Optimization algorithm. It is a Python implementation of the scipy.optimize.lbfgs
+algorithm.
+
+Information about the L-BFGS method can be found at the following links:
+
+ - https://docs.scipy.org/doc/scipy/reference/optimize.minimize-lbfgsb.html
+"""
+search = af.LBFGS(
+ tol=None,
+ disp=None,
+ maxcor=10,
+ ftol=2.220446049250313e-09,
+ gtol=1e-05,
+ eps=1e-08,
+ maxfun=15000,
+ maxiter=15000,
+ iprint=-1,
+ maxls=20,
+)
diff --git a/scripts/features/graphical_models.py b/scripts/features/graphical_models.py
index 4a990e3d..8c68d9c3 100644
--- a/scripts/features/graphical_models.py
+++ b/scripts/features/graphical_models.py
@@ -1,275 +1,271 @@
-"""
-Feature: Graphical Models
-=========================
-
-The examples so far have focused on fitting one model to one dataset, for example fitting 1D profiles composed of
-Gaussians to noisy 1D data. When multiple datasets were available each is fitted individually and their results
-interpreted one-by-one.
-
-However, for many problems we may have a large dataset and not be interested in how well the model fits each dataset
-individually. Instead, we may wish to fit this model (or many similar models) to the full dataset and determine
-the 'global' trends of the model across the datasets.
-
-This can be done using graphical models, which compose and fit a model that has 'local' parameters specific to each
-individual dataset but also higher-level model components that fit 'global' parameters of the model across the whole
-dataset. This framework can be easily extended to fit datasets with different properties, complex models with different
-topologies and has the functionality to allow it to be generalized to models with thousands of parameters.
-
-In this example, we demonstrate the API for performing graphical modeling in **PyAutoFit** using the example of
-simultaneously fitting 3 noisy 1D Gaussians. However, graphical models are an extensive feature and at the end of
-this example we will discuss other options available in **PyAutoFit** for composing a fitting a graphical model.
-
-The **HowToFit** tutorials contain a chapter dedicated to composing and fitting graphical models.
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Example Source Code (`af.ex`)**: The example objects used in this script.
-- **Dataset**: Load 3 noisy 1D Gaussian datasets for simultaneous fitting.
-- **Analysis**: Create Analysis objects for each dataset with a log likelihood function.
-- **Model**: Compose a graphical model with a shared prior across multiple model components.
-- **Analysis Factors**: Pair each Model with its corresponding Analysis class at factor graph nodes.
-- **Factor Graph**: Combine the Analysis Factors into a factor graph representing the graphical model.
-- **Search**: Create a non-linear search and fit the factor graph.
-- **Hierarchical Models**: Discuss how shared parameters can be drawn from a common parent distribution.
-- **Expectation Propagation**: Introduce the EP framework for scaling graphical models to high dimensionality.
-
-__Example Source Code (`af.ex`)__
-
-The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
-
- - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
- `visualize` functions.
-
- - `Gaussian`: a model component representing a 1D Gaussian profile.
-
-These are functionally identical to the `Analysis` and `Gaussian` objects you have seen elsewhere in the workspace.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-from os import path
-import matplotlib.pyplot as plt
-
-import autofit as af
-
-"""
-__Dataset__
-
-We are going to build a graphical model that fits three datasets.
-
-We begin by loading noisy 1D data containing 3 Gaussian's.
-"""
-total_datasets = 3
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(
- path.join("dataset", "example_1d", "gaussian_x1__low_snr", "dataset_0")
-):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-dataset_name_list = []
-data_list = []
-noise_map_list = []
-
-for dataset_index in range(total_datasets):
- dataset_name = f"dataset_{dataset_index}"
-
- dataset_path = path.join(
- "dataset", "example_1d", "gaussian_x1__low_snr", dataset_name
- )
-
- data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
- noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
- )
-
- data_list.append(data)
- noise_map_list.append(noise_map)
-
-"""
-Now lets plot the data, including their error bars. One should note that we are fitting much lower signal-to-noise
-datasets than usual.
-
-Note that all three of these `Gaussian`'s were simulated using the same `centre` value. To demonstrate graphical
-modeling we will therefore fit a model where the `centre` a shared parameter across the fit to the 3 `Gaussian`s,
-therefore making it a global parameter.
-"""
-for dataset_index in range(total_datasets):
- xvalues = range(data_list[dataset_index].shape[0])
-
- plt.errorbar(
- x=xvalues,
- y=data_list[dataset_index],
- yerr=noise_map_list[dataset_index],
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
- )
- plt.title("1D Gaussian Data #1.")
- plt.xlabel("x values of profile")
- plt.ylabel("Profile normalization")
- plt.show()
- plt.close()
-
-
-"""
-__Analysis__
-
-They are much lower signal-to-noise than the Gaussian's in other examples.
-
-Graphical models extract a lot more information from lower quantity datasets, something we demonstrate explicitly
-in the **HowToFit** lectures on graphical models.
-
-For each dataset we now create a corresponding `Analysis` class. By associating each dataset with an `Analysis`
-class we are therefore associating it with a unique `log_likelihood_function`.
-
-If our dataset had many different formats which each required their own unique `log_likelihood_function`, it would
-be straight forward to write customized `Analysis` classes for each dataset.
-"""
-analysis_list = []
-
-for data, noise_map in zip(data_list, noise_map_list):
- analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
- analysis_list.append(analysis)
-
-"""
-__Model__
-
-We now compose the graphical model we will fit using the `Model` objects described in the `overview` examples
-and chapter 1 of **HowToFit**.
-
-We begin by setting up a shared prior for `centre`, which is set up this up as a single `TruncatedGaussianPrior` that is
-passed to a unique `Model` for each `Gaussian`. This means all three `Gaussian`'s will be fitted wih the same
-value of `centre`.
-"""
-centre_shared_prior = af.GaussianPrior(mean=50.0, sigma=30.0)
-
-"""
-We now set up three `Model`'s, each of which contain a `Gaussian` that is used to fit each of the
-datasets we loaded above.
-
-All three of these `Model`'s use the `centre_shared_prior`. This means all three model-components use
-the same value of `centre` for every model composed and fitted by the `NonLinearSearch`, reducing the dimensionality
-of parameter space from N=9 (e.g. 3 parameters per Gaussian) to N=7.
-"""
-model_list = []
-
-for model_index in range(len(data_list)):
- gaussian = af.Model(af.ex.Gaussian)
-
- gaussian.centre = centre_shared_prior # This prior is used by all 3 Gaussians!
- gaussian.normalization = af.LogUniformPrior(lower_limit=1e-6, upper_limit=1e6)
- gaussian.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=25.0)
-
- model_list.append(gaussian)
-
-"""
-__Analysis Factors__
-
-Above, we composed a model consisting of three `Gaussian`'s with a shared `centre` prior. We also loaded three
-datasets which we intend to fit with each of these `Gaussians`, setting up each in an `Analysis` class that defines
-how the model is used to fit the data.
-
-We now simply need to pair each model-component to each `Analysis` class, so that:
-
-- `prior_model_0` fits `data_0` via `analysis_0`.
-- `prior_model_1` fits `data_1` via `analysis_1`.
-- `prior_model_2` fits `data_2` via `analysis_2`.
-
-The point where a `Model` and `Analysis` class meet is called a `AnalysisFactor`.
-
-This term is used to denote that we are composing a graphical model, which is commonly termed a 'factor graph'. A
-factor defines a node on this graph where we have some data, a model, and we fit the two together. The 'links' between
-these different nodes then define the global model we are fitting.
-"""
-analysis_factor_list = []
-
-for model, analysis in zip(model_list, analysis_list):
- analysis_factor = af.AnalysisFactor(prior_model=model, analysis=analysis)
-
- analysis_factor_list.append(analysis_factor)
-
-"""
-__Factor Graph__
-
-We combine our `AnalysisFactors` into one, to compose the factor graph.
-
-So, what is a factor graph?
-
-A factor graph defines the graphical model we have composed. For example, it defines the different model components
-that make up our model (e.g. the three `Gaussian` classes) and how their parameters are linked or shared (e.g. that
-each `Gaussian` has its own unique `normalization` and `sigma`, but a shared `centre` parameter.
-
-This is what our factor graph looks like:
-
-The factor graph above is made up of two components:
-
-- Nodes: these are points on the graph where we have a unique set of data and a model that is made up of a subset of
-our overall graphical model. This is effectively the `AnalysisFactor` objects we created above.
-
-- Links: these define the model components and parameters that are shared across different nodes and thus retain the
-same values when fitting different datasets.
-"""
-factor_graph = af.FactorGraphModel(*analysis_factor_list)
-
-"""
-__Search__
-
-We can now create a non-linear search and used it to the fit the factor graph, using its `global_prior_model` property.
-"""
-search = af.DynestyStatic(
- path_prefix="features", name="graphical_model", sample="rwalk"
-)
-
-result = search.fit(model=factor_graph.global_prior_model, analysis=factor_graph)
-
-"""
-This will fit the N=7 dimension parameter space where every Gaussian has a shared centre!
-
-This is all expanded upon in the **HowToFit** chapter on graphical models, where we will give a more detailed
-description of why this approach to model-fitting extracts a lot more information than fitting each dataset
-one-by-one.
-
-__Hierarchical Models__
-
-A specific type of graphical model is a hierarchical model, where the shared parameter(s) of a graph are assumed
-to be drawn from a common parent distribution.
-
-Fitting the datasets simultaneously enables better estimate of this global hierarchical distribution.
-
-__Expectation Propagation__
-
-For large datasets, a graphical model may have hundreds, thousands, or *hundreds of thousands* of parameters. The
-high dimensionality of such a parameter space can make it inefficient or impossible to fit the model.
-
-Fitting high dimensionality graphical models in **PyAutoFit** can use an Expectation Propagation (EP) framework to
-make scaling up feasible. This framework fits every dataset individually and pass messages throughout the graph to
-inform every fit the expected
-values of each parameter.
-
-The following paper describes the EP framework in formal Bayesian notation:
-
-https://arxiv.org/pdf/1412.4869.pdf
-
-Hierarchical models can also be scaled up to large datasets via Expectation Propagation.
-"""
+"""
+Feature: Graphical Models
+=========================
+
+The examples so far have focused on fitting one model to one dataset, for example fitting 1D profiles composed of
+Gaussians to noisy 1D data. When multiple datasets were available each is fitted individually and their results
+interpreted one-by-one.
+
+However, for many problems we may have a large dataset and not be interested in how well the model fits each dataset
+individually. Instead, we may wish to fit this model (or many similar models) to the full dataset and determine
+the 'global' trends of the model across the datasets.
+
+This can be done using graphical models, which compose and fit a model that has 'local' parameters specific to each
+individual dataset but also higher-level model components that fit 'global' parameters of the model across the whole
+dataset. This framework can be easily extended to fit datasets with different properties, complex models with different
+topologies and has the functionality to allow it to be generalized to models with thousands of parameters.
+
+In this example, we demonstrate the API for performing graphical modeling in **PyAutoFit** using the example of
+simultaneously fitting 3 noisy 1D Gaussians. However, graphical models are an extensive feature and at the end of
+this example we will discuss other options available in **PyAutoFit** for composing a fitting a graphical model.
+
+The **HowToFit** tutorials contain a chapter dedicated to composing and fitting graphical models.
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Example Source Code (`af.ex`)**: The example objects used in this script.
+- **Dataset**: Load 3 noisy 1D Gaussian datasets for simultaneous fitting.
+- **Analysis**: Create Analysis objects for each dataset with a log likelihood function.
+- **Model**: Compose a graphical model with a shared prior across multiple model components.
+- **Analysis Factors**: Pair each Model with its corresponding Analysis class at factor graph nodes.
+- **Factor Graph**: Combine the Analysis Factors into a factor graph representing the graphical model.
+- **Search**: Create a non-linear search and fit the factor graph.
+- **Hierarchical Models**: Discuss how shared parameters can be drawn from a common parent distribution.
+- **Expectation Propagation**: Introduce the EP framework for scaling graphical models to high dimensionality.
+
+__Example Source Code (`af.ex`)__
+
+The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
+
+ - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
+ `visualize` functions.
+
+ - `Gaussian`: a model component representing a 1D Gaussian profile.
+
+These are functionally identical to the `Analysis` and `Gaussian` objects you have seen elsewhere in the workspace.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+from os import path
+import matplotlib.pyplot as plt
+
+import autofit as af
+
+"""
+__Dataset__
+
+We are going to build a graphical model that fits three datasets.
+
+We begin by loading noisy 1D data containing 3 Gaussian's.
+"""
+total_datasets = 3
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(
+ path.join("dataset", "example_1d", "gaussian_x1__low_snr", "dataset_0")
+):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+dataset_name_list = []
+data_list = []
+noise_map_list = []
+
+for dataset_index in range(total_datasets):
+ dataset_name = f"dataset_{dataset_index}"
+
+ dataset_path = path.join(
+ "dataset", "example_1d", "gaussian_x1__low_snr", dataset_name
+ )
+
+ data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+ noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+ )
+
+ data_list.append(data)
+ noise_map_list.append(noise_map)
+
+"""
+Now lets plot the data, including their error bars. One should note that we are fitting much lower signal-to-noise
+datasets than usual.
+
+Note that all three of these `Gaussian`'s were simulated using the same `centre` value. To demonstrate graphical
+modeling we will therefore fit a model where the `centre` a shared parameter across the fit to the 3 `Gaussian`s,
+therefore making it a global parameter.
+"""
+for dataset_index in range(total_datasets):
+ xvalues = range(data_list[dataset_index].shape[0])
+
+ plt.errorbar(
+ x=xvalues,
+ y=data_list[dataset_index],
+ yerr=noise_map_list[dataset_index],
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+ )
+ plt.title("1D Gaussian Data #1.")
+ plt.xlabel("x values of profile")
+ plt.ylabel("Profile normalization")
+ plt.show()
+ plt.close()
+
+
+"""
+__Analysis__
+
+They are much lower signal-to-noise than the Gaussian's in other examples.
+
+Graphical models extract a lot more information from lower quantity datasets, something we demonstrate explicitly
+in the **HowToFit** lectures on graphical models.
+
+For each dataset we now create a corresponding `Analysis` class. By associating each dataset with an `Analysis`
+class we are therefore associating it with a unique `log_likelihood_function`.
+
+If our dataset had many different formats which each required their own unique `log_likelihood_function`, it would
+be straight forward to write customized `Analysis` classes for each dataset.
+"""
+analysis_list = []
+
+for data, noise_map in zip(data_list, noise_map_list):
+ analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+ analysis_list.append(analysis)
+
+"""
+__Model__
+
+We now compose the graphical model we will fit using the `Model` objects described in the `overview` examples
+and chapter 1 of **HowToFit**.
+
+We begin by setting up a shared prior for `centre`, which is set up this up as a single `TruncatedGaussianPrior` that is
+passed to a unique `Model` for each `Gaussian`. This means all three `Gaussian`'s will be fitted wih the same
+value of `centre`.
+"""
+centre_shared_prior = af.GaussianPrior(mean=50.0, sigma=30.0)
+
+"""
+We now set up three `Model`'s, each of which contain a `Gaussian` that is used to fit each of the
+datasets we loaded above.
+
+All three of these `Model`'s use the `centre_shared_prior`. This means all three model-components use
+the same value of `centre` for every model composed and fitted by the `NonLinearSearch`, reducing the dimensionality
+of parameter space from N=9 (e.g. 3 parameters per Gaussian) to N=7.
+"""
+model_list = []
+
+for model_index in range(len(data_list)):
+ gaussian = af.Model(af.ex.Gaussian)
+
+ gaussian.centre = centre_shared_prior # This prior is used by all 3 Gaussians!
+ gaussian.normalization = af.LogUniformPrior(lower_limit=1e-6, upper_limit=1e6)
+ gaussian.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=25.0)
+
+ model_list.append(gaussian)
+
+"""
+__Analysis Factors__
+
+Above, we composed a model consisting of three `Gaussian`'s with a shared `centre` prior. We also loaded three
+datasets which we intend to fit with each of these `Gaussians`, setting up each in an `Analysis` class that defines
+how the model is used to fit the data.
+
+We now simply need to pair each model-component to each `Analysis` class, so that:
+
+- `prior_model_0` fits `data_0` via `analysis_0`.
+- `prior_model_1` fits `data_1` via `analysis_1`.
+- `prior_model_2` fits `data_2` via `analysis_2`.
+
+The point where a `Model` and `Analysis` class meet is called a `AnalysisFactor`.
+
+This term is used to denote that we are composing a graphical model, which is commonly termed a 'factor graph'. A
+factor defines a node on this graph where we have some data, a model, and we fit the two together. The 'links' between
+these different nodes then define the global model we are fitting.
+"""
+analysis_factor_list = []
+
+for model, analysis in zip(model_list, analysis_list):
+ analysis_factor = af.AnalysisFactor(prior_model=model, analysis=analysis)
+
+ analysis_factor_list.append(analysis_factor)
+
+"""
+__Factor Graph__
+
+We combine our `AnalysisFactors` into one, to compose the factor graph.
+
+So, what is a factor graph?
+
+A factor graph defines the graphical model we have composed. For example, it defines the different model components
+that make up our model (e.g. the three `Gaussian` classes) and how their parameters are linked or shared (e.g. that
+each `Gaussian` has its own unique `normalization` and `sigma`, but a shared `centre` parameter.
+
+This is what our factor graph looks like:
+
+The factor graph above is made up of two components:
+
+- Nodes: these are points on the graph where we have a unique set of data and a model that is made up of a subset of
+our overall graphical model. This is effectively the `AnalysisFactor` objects we created above.
+
+- Links: these define the model components and parameters that are shared across different nodes and thus retain the
+same values when fitting different datasets.
+"""
+factor_graph = af.FactorGraphModel(*analysis_factor_list)
+
+"""
+__Search__
+
+We can now create a non-linear search and used it to the fit the factor graph, using its `global_prior_model` property.
+"""
+search = af.DynestyStatic(
+ path_prefix="features", name="graphical_model", sample="rwalk"
+)
+
+result = search.fit(model=factor_graph.global_prior_model, analysis=factor_graph)
+
+"""
+This will fit the N=7 dimension parameter space where every Gaussian has a shared centre!
+
+This is all expanded upon in the **HowToFit** chapter on graphical models, where we will give a more detailed
+description of why this approach to model-fitting extracts a lot more information than fitting each dataset
+one-by-one.
+
+__Hierarchical Models__
+
+A specific type of graphical model is a hierarchical model, where the shared parameter(s) of a graph are assumed
+to be drawn from a common parent distribution.
+
+Fitting the datasets simultaneously enables better estimate of this global hierarchical distribution.
+
+__Expectation Propagation__
+
+For large datasets, a graphical model may have hundreds, thousands, or *hundreds of thousands* of parameters. The
+high dimensionality of such a parameter space can make it inefficient or impossible to fit the model.
+
+Fitting high dimensionality graphical models in **PyAutoFit** can use an Expectation Propagation (EP) framework to
+make scaling up feasible. This framework fits every dataset individually and pass messages throughout the graph to
+inform every fit the expected
+values of each parameter.
+
+The following paper describes the EP framework in formal Bayesian notation:
+
+https://arxiv.org/pdf/1412.4869.pdf
+
+Hierarchical models can also be scaled up to large datasets via Expectation Propagation.
+"""
diff --git a/scripts/features/interpolate.py b/scripts/features/interpolate.py
index fd36da56..b772701f 100644
--- a/scripts/features/interpolate.py
+++ b/scripts/features/interpolate.py
@@ -1,278 +1,274 @@
-"""
-Feature: Interpolate
-====================
-
-It is common to fit a model to many similar datasets, where it is anticipated that one or more model parameters vary
-smoothly across the datasets.
-
-For example, the datasets may be taken at different times, where the signal in the data and therefore model parameters
-vary smoothly as a function of time. Alternatively, the datasets may be taken at different wavelengths, with the signal
-varying smoothly as a function of wavelength.
-
-It may be desirable to fit the datasets one-by-one and then interpolate the results in order to determine the most
-likely model parameters at any point in time (or at any wavelength).
-
-This example illustrates model interpolation functionality in **PyAutoFit** using the example of fitting 3 noisy
-1D Gaussians, where these data are assumed to have been taken at 3 different times. The `centre` of each `Gaussian`
-varies smoothly over time. The interpolation is therefore used to estimate the `centre` of each `Gaussian` at any time
-outside of the times the data were observed.
-
-__Example Source Code (`af.ex`)__
-
-The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
-
- - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
- `visualize` functions.
-
- - `Gaussian`: a model component representing a 1D Gaussian profile.
-
-These are functionally identical to the `Analysis` and `Gaussian` objects you have seen elsewhere in the workspace.
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Example Source Code (`af.ex`)**: The example objects used in this script.
-- **Dataset**: Load 3 noisy 1D Gaussian datasets taken at different times.
-- **Fit**: Fit each dataset individually, storing the maximum likelihood instances for interpolation.
-- **Interpolation**: Use a LinearInterpolator to interpolate model parameters as a function of time.
-- **Serialization**: Serialize the interpolator to a JSON file for reuse.
-- **Database**: Load results from hard disk using the Aggregator for interpolation.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import numpy as np
-import matplotlib.pyplot as plt
-from os import path
-
-import autofit as af
-
-"""
-__Dataset__
-
-We illustrate model interpolation using 3 noisy 1D Gaussian datasets taken at 3 different times, where the `centre` of
-each `Gaussian` varies smoothly over time.
-
-The datasets are taken at 3 times, t=0, t=1 and t=2, which defines the name of the folder we load the data from.
-
-We load each data and noise map and store them in lists, so we can plot them next.
-"""
-total_datasets = 3
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(path.join("dataset", "example_1d", "gaussian_x1_time", "time_0")):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data_list = []
-noise_map_list = []
-time_list = []
-
-for time in range(3):
- dataset_name = f"time_{time}"
-
- dataset_prefix_path = path.join("dataset", "example_1d", "gaussian_x1_time")
-
- dataset_path = path.join(dataset_prefix_path, dataset_name)
-
- data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
- noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
- )
-
- data_list.append(data)
- noise_map_list.append(noise_map)
- time_list.append(time)
-
-"""
-Visual comparison of the datasets shows that the `centre` of each `Gaussian` varies smoothly over time, with it moving
-from pixel 40 at t=0 to pixel 60 at t=2.
-"""
-for time in range(3):
- xvalues = range(data_list[time].shape[0])
-
- plt.errorbar(
- x=xvalues,
- y=data_list[time],
- yerr=noise_map_list[time],
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
- )
- plt.title("1D Gaussian Data #1.")
- plt.xlabel("x values of profile")
- plt.ylabel("Profile normalization")
- plt.show()
- plt.close()
-
-
-"""
-__Fit__
-
-We now fit each of the 3 datasets.
-
-The fits are performed in a for loop, with the docstrings inside the loop explaining the code.
-
-The interpolate at the end of the fits uses the maximum log likelihood model of each fit, which we store in a list.
-"""
-ml_instances_list = []
-
-for data, noise_map, time in zip(data_list, noise_map_list, time_list):
- """
- __Analysis__
-
- For each dataset we create an `Analysis` class, which includes the `log_likelihood_function` we fit the data with.
- """
- analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
- """
- __Time__
-
- The model composed below has an input not seen in other examples, the parameter `time`.
-
- This is the time that the simulated data was acquired and is not a free parameter in the fit.
-
- For interpolation it plays a crucial role, as the model is interpolated to the time of every dataset input
- into the model below. If the `time` input were missing, interpolation could not be performed.
-
- Over the iterations of the for loop, the `time` input will therefore be the values 0.0, 1.0 and 2.0.
-
- __Model__
-
- We now compose our model, which is a single `Gaussian`.
-
- The `centre` of the `Gaussian` is a free parameter with a `UniformPrior` that ranges between 0.0 and 100.0.
-
- We expect the inferred `centre` inferred from the fit to each dataset to vary smoothly as a function of time.
- """
- model = af.Collection(gaussian=af.Model(af.ex.Gaussian), time=time)
-
- """
- __Search__
-
- The model is fitted to the data using the nested sampling algorithm
- Dynesty (https://johannesbuchner.github.io/UltraNest/readme.html).
- """
- search = af.DynestyStatic(
- path_prefix=path.join("interpolate"),
- name=f"time_{time}",
- nlive=100,
- )
-
- """
- __Model-Fit__
-
- We can now begin the model-fit by passing the model and analysis object to the search, which performs a non-linear
- search to find which models fit the data with the highest likelihood.
- """
- result = search.fit(model=model, analysis=analysis)
-
- """
- __Instances__
-
- Interpolation uses the maximum log likelihood model of each fit to build an interpolation model of the model as a
- function of time.
-
- We therefore store the maximum log likelihood model of every fit in a list, which is used below.
- """
- ml_instances_list.append(result.instance)
-
-
-"""
-__Interpolation__
-
-Now all fits are complete, we use the `ml_instances_list` to build an interpolation model of the model as a function
-of time.
-
-This is performed using the `LinearInterpolator` object, which interpolates the model parameters as a function of
-time linearly between the values computed by the model-fits above.
-
-More advanced interpolation schemes are available and described in the `interpolation.py` example.
-"""
-interpolator = af.LinearInterpolator(instances=ml_instances_list)
-
-"""
-The model can be interpolated to any time, for example time=1.5.
-
-This returns a new `instance` of the model, as an instance of the `Gaussian` object, where the parameters are computed
-by interpolating between the values computed above.
-"""
-instance = interpolator[interpolator.time == 1.5]
-
-"""
-The `centre` of the `Gaussian` at time 1.5 is between the value inferred for the first and second fits taken
-at times 1.0 and 2.0.
-
-This is a `centre` close to a value of 55.0.
-"""
-print(f"Gaussian centre of fit 1 (t = 1): {ml_instances_list[0].gaussian.centre}")
-print(f"Gaussian centre of fit 2 (t = 2): {ml_instances_list[1].gaussian.centre}")
-
-print(f"Gaussian centre interpolated at t = 1.5 {instance.gaussian.centre}")
-
-
-"""
-__Serialization__
-
-The interpolator and model can be serialized to a .json file using **PyAutoConf**'s dedicated serialization methods.
-
-This means an interpolator can easily be loaded into other scripts.
-"""
-from autoconf.dictable import output_to_json, from_json
-
-json_file = path.join(dataset_prefix_path, "interpolator.json")
-
-output_to_json(obj=interpolator, file_path=json_file)
-
-interpolator = from_json(file_path=json_file)
-
-"""
-__Database__
-
-It may be inconvenient to fit all the models in a single Python script (e.g. the model-fits take a long time and you
-are fitting many datasets).
-
-PyAutoFit's allows you to store the results of model-fits from hard-disk.
-
-Database functionality then allows you to load the results of the fit above, set up the interpolator and perform the
-interpolation.
-
-If you are not familiar with the database API, you should checkout the `cookbook/database.ipynb` example.
-"""
-from autofit.aggregator.aggregator import Aggregator
-
-agg = Aggregator.from_directory(
- directory=path.join("output", "interpolate"), completed_only=False
-)
-
-ml_instances_list = [samps.max_log_likelihood() for samps in agg.values("samples")]
-
-interpolator = af.LinearInterpolator(instances=ml_instances_list)
-
-instance = interpolator[interpolator.time == 1.5]
-
-print(f"Gaussian centre of fit 1 (t = 1): {ml_instances_list[0].gaussian.centre}")
-print(f"Gaussian centre of fit 2 (t = 2): {ml_instances_list[1].gaussian.centre}")
-
-print(f"Gaussian centre interpolated at t = 1.5 {instance.gaussian.centre}")
-
-"""
-Finish.
-"""
+"""
+Feature: Interpolate
+====================
+
+It is common to fit a model to many similar datasets, where it is anticipated that one or more model parameters vary
+smoothly across the datasets.
+
+For example, the datasets may be taken at different times, where the signal in the data and therefore model parameters
+vary smoothly as a function of time. Alternatively, the datasets may be taken at different wavelengths, with the signal
+varying smoothly as a function of wavelength.
+
+It may be desirable to fit the datasets one-by-one and then interpolate the results in order to determine the most
+likely model parameters at any point in time (or at any wavelength).
+
+This example illustrates model interpolation functionality in **PyAutoFit** using the example of fitting 3 noisy
+1D Gaussians, where these data are assumed to have been taken at 3 different times. The `centre` of each `Gaussian`
+varies smoothly over time. The interpolation is therefore used to estimate the `centre` of each `Gaussian` at any time
+outside of the times the data were observed.
+
+__Example Source Code (`af.ex`)__
+
+The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
+
+ - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
+ `visualize` functions.
+
+ - `Gaussian`: a model component representing a 1D Gaussian profile.
+
+These are functionally identical to the `Analysis` and `Gaussian` objects you have seen elsewhere in the workspace.
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Example Source Code (`af.ex`)**: The example objects used in this script.
+- **Dataset**: Load 3 noisy 1D Gaussian datasets taken at different times.
+- **Fit**: Fit each dataset individually, storing the maximum likelihood instances for interpolation.
+- **Interpolation**: Use a LinearInterpolator to interpolate model parameters as a function of time.
+- **Serialization**: Serialize the interpolator to a JSON file for reuse.
+- **Database**: Load results from hard disk using the Aggregator for interpolation.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import numpy as np
+import matplotlib.pyplot as plt
+from os import path
+
+import autofit as af
+
+"""
+__Dataset__
+
+We illustrate model interpolation using 3 noisy 1D Gaussian datasets taken at 3 different times, where the `centre` of
+each `Gaussian` varies smoothly over time.
+
+The datasets are taken at 3 times, t=0, t=1 and t=2, which defines the name of the folder we load the data from.
+
+We load each data and noise map and store them in lists, so we can plot them next.
+"""
+total_datasets = 3
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(path.join("dataset", "example_1d", "gaussian_x1_time", "time_0")):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data_list = []
+noise_map_list = []
+time_list = []
+
+for time in range(3):
+ dataset_name = f"time_{time}"
+
+ dataset_prefix_path = path.join("dataset", "example_1d", "gaussian_x1_time")
+
+ dataset_path = path.join(dataset_prefix_path, dataset_name)
+
+ data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+ noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+ )
+
+ data_list.append(data)
+ noise_map_list.append(noise_map)
+ time_list.append(time)
+
+"""
+Visual comparison of the datasets shows that the `centre` of each `Gaussian` varies smoothly over time, with it moving
+from pixel 40 at t=0 to pixel 60 at t=2.
+"""
+for time in range(3):
+ xvalues = range(data_list[time].shape[0])
+
+ plt.errorbar(
+ x=xvalues,
+ y=data_list[time],
+ yerr=noise_map_list[time],
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+ )
+ plt.title("1D Gaussian Data #1.")
+ plt.xlabel("x values of profile")
+ plt.ylabel("Profile normalization")
+ plt.show()
+ plt.close()
+
+
+"""
+__Fit__
+
+We now fit each of the 3 datasets.
+
+The fits are performed in a for loop, with the docstrings inside the loop explaining the code.
+
+The interpolate at the end of the fits uses the maximum log likelihood model of each fit, which we store in a list.
+"""
+ml_instances_list = []
+
+for data, noise_map, time in zip(data_list, noise_map_list, time_list):
+ """
+ __Analysis__
+
+ For each dataset we create an `Analysis` class, which includes the `log_likelihood_function` we fit the data with.
+ """
+ analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+ """
+ __Time__
+
+ The model composed below has an input not seen in other examples, the parameter `time`.
+
+ This is the time that the simulated data was acquired and is not a free parameter in the fit.
+
+ For interpolation it plays a crucial role, as the model is interpolated to the time of every dataset input
+ into the model below. If the `time` input were missing, interpolation could not be performed.
+
+ Over the iterations of the for loop, the `time` input will therefore be the values 0.0, 1.0 and 2.0.
+
+ __Model__
+
+ We now compose our model, which is a single `Gaussian`.
+
+ The `centre` of the `Gaussian` is a free parameter with a `UniformPrior` that ranges between 0.0 and 100.0.
+
+ We expect the inferred `centre` inferred from the fit to each dataset to vary smoothly as a function of time.
+ """
+ model = af.Collection(gaussian=af.Model(af.ex.Gaussian), time=time)
+
+ """
+ __Search__
+
+ The model is fitted to the data using the nested sampling algorithm
+ Dynesty (https://johannesbuchner.github.io/UltraNest/readme.html).
+ """
+ search = af.DynestyStatic(
+ path_prefix=path.join("interpolate"),
+ name=f"time_{time}",
+ nlive=100,
+ )
+
+ """
+ __Model-Fit__
+
+ We can now begin the model-fit by passing the model and analysis object to the search, which performs a non-linear
+ search to find which models fit the data with the highest likelihood.
+ """
+ result = search.fit(model=model, analysis=analysis)
+
+ """
+ __Instances__
+
+ Interpolation uses the maximum log likelihood model of each fit to build an interpolation model of the model as a
+ function of time.
+
+ We therefore store the maximum log likelihood model of every fit in a list, which is used below.
+ """
+ ml_instances_list.append(result.instance)
+
+
+"""
+__Interpolation__
+
+Now all fits are complete, we use the `ml_instances_list` to build an interpolation model of the model as a function
+of time.
+
+This is performed using the `LinearInterpolator` object, which interpolates the model parameters as a function of
+time linearly between the values computed by the model-fits above.
+
+More advanced interpolation schemes are available and described in the `interpolation.py` example.
+"""
+interpolator = af.LinearInterpolator(instances=ml_instances_list)
+
+"""
+The model can be interpolated to any time, for example time=1.5.
+
+This returns a new `instance` of the model, as an instance of the `Gaussian` object, where the parameters are computed
+by interpolating between the values computed above.
+"""
+instance = interpolator[interpolator.time == 1.5]
+
+"""
+The `centre` of the `Gaussian` at time 1.5 is between the value inferred for the first and second fits taken
+at times 1.0 and 2.0.
+
+This is a `centre` close to a value of 55.0.
+"""
+print(f"Gaussian centre of fit 1 (t = 1): {ml_instances_list[0].gaussian.centre}")
+print(f"Gaussian centre of fit 2 (t = 2): {ml_instances_list[1].gaussian.centre}")
+
+print(f"Gaussian centre interpolated at t = 1.5 {instance.gaussian.centre}")
+
+
+"""
+__Serialization__
+
+The interpolator and model can be serialized to a .json file using **PyAutoConf**'s dedicated serialization methods.
+
+This means an interpolator can easily be loaded into other scripts.
+"""
+from autoconf.dictable import output_to_json, from_json
+
+json_file = path.join(dataset_prefix_path, "interpolator.json")
+
+output_to_json(obj=interpolator, file_path=json_file)
+
+interpolator = from_json(file_path=json_file)
+
+"""
+__Database__
+
+It may be inconvenient to fit all the models in a single Python script (e.g. the model-fits take a long time and you
+are fitting many datasets).
+
+PyAutoFit's allows you to store the results of model-fits from hard-disk.
+
+Database functionality then allows you to load the results of the fit above, set up the interpolator and perform the
+interpolation.
+
+If you are not familiar with the database API, you should checkout the `cookbook/database.ipynb` example.
+"""
+from autofit.aggregator.aggregator import Aggregator
+
+agg = Aggregator.from_directory(
+ directory=path.join("output", "interpolate"), completed_only=False
+)
+
+ml_instances_list = [samps.max_log_likelihood() for samps in agg.values("samples")]
+
+interpolator = af.LinearInterpolator(instances=ml_instances_list)
+
+instance = interpolator[interpolator.time == 1.5]
+
+print(f"Gaussian centre of fit 1 (t = 1): {ml_instances_list[0].gaussian.centre}")
+print(f"Gaussian centre of fit 2 (t = 2): {ml_instances_list[1].gaussian.centre}")
+
+print(f"Gaussian centre interpolated at t = 1.5 {instance.gaussian.centre}")
+
+"""
+Finish.
+"""
diff --git a/scripts/features/model_comparison.py b/scripts/features/model_comparison.py
index ea8c4622..3c906141 100644
--- a/scripts/features/model_comparison.py
+++ b/scripts/features/model_comparison.py
@@ -1,371 +1,367 @@
-"""
-Feature: Model Comparison
-=========================
-
-Common questions when fitting a model to data are: what model should I use? How many parameters should the model have?
-Is the model too complex or too simple?
-
-Model comparison answers to these questions. It amounts to composing and fitting many different models to the data
-and comparing how well they fit the data.
-
-This example illustrates model comparison using the noisy 1D Gaussian example. We fit a dataset consisting of two
-Gaussians and fit it with three models comprised of 1, 2 and 3 Gaussian's respectively. Using the Bayesian evidence to
-compare the models, we favour the model with 2 Gaussians, which is the "correct" model given that it was the model used
-to simulate the dataset in the first place.
-
-__Metrics__
-
-Different metrics can be used compare models and quantify their goodness-of-fit.
-
-In this example we show the results of using two different metrics:
-
- - `log_likelihood`: The value returned by the `log_likelihood_function` of an `Analysis` object. which is directly
- related to the sum of the residuals squared (e.g. the `chi_squared`). The log likelihood does not change when more
- or less parameters are included in the model, therefore it does not account for over-fitting and will often favour
- more complex models irrespective of whether they fit the data better.
-
- - `log_evidence`: The Bayesian evidence, which is closely related to the log likelihood but utilizes additional
- information which penalizes models based on their complexity. The Bayesian evidence will therefore favour simpler
- models over more complex models, unless the more complex model provides a much better fit to the data.
-
-__Example Source Code (`af.ex`)__
-
-The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
-
- - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
- `visualize` functions.
-
- - `Gaussian`: a model component representing a 1D Gaussian profile.
-
-These are functionally identical to the `Analysis` and `Gaussian` objects you have seen elsewhere in the workspace.
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Metrics**: Describe the log likelihood and Bayesian evidence metrics used for model comparison.
-- **Example Source Code (`af.ex`)**: The example objects used in this script.
-- **Data**: Load the 1D Gaussian data consisting of two Gaussians.
-- **Model x1 Gaussian**: Create and fit a model with a single Gaussian.
-- **Model x2 Gaussian**: Create and fit a model with two Gaussians.
-- **Model x3 Gaussian**: Create and fit a model with three Gaussians.
-- **Wrap Up**: Summarize the model comparison results using log likelihood and Bayesian evidence.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import matplotlib.pyplot as plt
-import numpy as np
-from os import path
-
-import autofit as af
-
-"""
-__Data__
-
-Load data of a 1D Gaussian from a .json file in the directory `autofit_workspace/dataset/gaussian_x2`.
-
-This 1D data was created using two 1D Gaussians, therefore model comparison should favor a model with two Gaussians over
-a models with 1 or 3 Gaussians.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x2")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-"""
-Plot the data.
-"""
-xvalues = range(data.shape[0])
-
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- color="k",
- ecolor="k",
- linestyle="",
- elinewidth=1,
- capsize=2,
-)
-plt.title("1D Gaussian Dataset Used For Model Comparison.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-__Model x1 Gaussian__
-
-Create a model to fit the data, starting with a model where the data is fitted with 1 Gaussian.
-"""
-model = af.Collection(gaussian_0=af.ex.Gaussian)
-
-"""
-The `info` attribute shows the model in a readable format, showing it contains one `Gaussian`.
-"""
-print(model.info)
-
-"""
-Create the analysis which fits the model to the data.
-"""
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-"""
-Fit the data using a non-linear search, to determine the goodness of fit of this model.
-
-We use the nested sampling algorithm Dynesty, noting that the Bayesian evidence (`log_evidence`) of a model can only
-be estimated using a nested sampling algorithm.
-"""
-search = af.DynestyStatic(
- path_prefix=path.join("features", "model_comparison"),
- name="gaussian_x1",
- nlive=50,
- iterations_per_full_update=3000,
-)
-
-"""
-Perform the fit.
-"""
-result_x1_gaussian = search.fit(model=model, analysis=analysis)
-
-"""
-The results are concisely summarised using the `result.info` property.
-
-These show that the parameters of the Gaussian are well constrained, with small errors on their inferred values.
-However, it does not inform us of whether the model provides a good fit to the data overall.
-"""
-print(result_x1_gaussian.info)
-
-"""
-The maximum log likelihood model is used to visualize the fit.
-
-For 1 Gaussian, residuals are visible, whereby the model Gaussian cannot fit the highest central data-point and
-there is a mismatch at the edges of the profile around pixels 40 and 60.
-
-Based on visual inspection, the model therefore provides a poor fit to the data.
-"""
-instance = result_x1_gaussian.max_log_likelihood_instance
-
-gaussian_0 = instance.gaussian_0.model_data_from(xvalues=np.arange(data.shape[0]))
-model_data = gaussian_0
-
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- color="k",
- ecolor="k",
- linestyle="",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(range(data.shape[0]), model_data, color="r")
-plt.plot(range(data.shape[0]), gaussian_0, "--")
-plt.title("Model fit using 1 Gaussian.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-Print the `log_likelihood` and `log_evidence` of this model-fit, which we will compare to more complex models in order
-to determine which model provides the best fit to the data.
-"""
-print("1 Gaussian:")
-print(f"Log Likelihood: {result_x1_gaussian.samples.max_log_likelihood()}")
-print(f"Log Evidence: {result_x1_gaussian.samples.log_evidence}")
-
-"""
-__Model x2 Gaussian__
-
-We now create a model to fit the data which consists of 2 Gaussians.
-"""
-model = af.Collection(gaussian_0=af.ex.Gaussian, gaussian_1=af.ex.Gaussian)
-
-"""
-The `info` attribute shows the model now consists of two `Gaussian`'s.
-"""
-print(model.info)
-
-"""
-We repeat the steps above to create the non-linear search and perform the model-fit.
-"""
-search = af.DynestyStatic(
- path_prefix=path.join("features", "model_comparison"),
- name="gaussian_x2",
- nlive=50,
- iterations_per_full_update=3000,
-)
-
-result_x2_gaussian = search.fit(model=model, analysis=analysis)
-
-"""
-The results show that two Gaussians have now been fitted to the data.
-"""
-print(result_x2_gaussian.info)
-
-"""
-Visualizing the fit, we see that the problems with the previous fit have been addressed. The central data-point at the
-highest normalization is fitted correctly and the residuals at the edges of the profile around pixels 40 and 60 are
-significantly reduced.
-
-There are effectively no residuals, indicating that the model provides a good fit to the data.
-
-The residuals are so small that they are consistent with noise in the data. One therefore should not expect that
-a more complex model than one with 2 Gaussians can provide a better fit.
-"""
-instance = result_x2_gaussian.max_log_likelihood_instance
-
-gaussian_0 = instance.gaussian_0.model_data_from(xvalues=np.arange(data.shape[0]))
-gaussian_1 = instance.gaussian_0.model_data_from(xvalues=np.arange(data.shape[0]))
-model_data = gaussian_0 + gaussian_1
-
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- color="k",
- ecolor="k",
- linestyle="",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(range(data.shape[0]), model_data, color="r")
-plt.plot(range(data.shape[0]), gaussian_0, "--")
-plt.plot(range(data.shape[0]), gaussian_1, "--")
-plt.title("Model fit using 2 Gaussian.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-Print the `log_likelihood` and `log_evidence` of this model-fit, and compare these values to the previous model-fit
-which used 1 Gaussian.
-"""
-print("1 Gaussian:")
-print(f"Log Likelihood: {max(result_x1_gaussian.samples.log_likelihood_list)}")
-print(f"Log Evidence: {result_x1_gaussian.samples.log_evidence}")
-
-print("2 Gaussians:")
-print(f"Log Likelihood: {max(result_x2_gaussian.samples.log_likelihood_list)}")
-print(f"Log Evidence: {result_x2_gaussian.samples.log_evidence}")
-
-"""
-Both the `log_likelihood` and `log_evidence` have increased significantly, indicating that the model with 2 Gaussians
-is favored over the model with 1 Gaussian.
-
-This is expected, as we know the data was generated using 2 Gaussians!
-
-__Model x3 Gaussian__
-
-We now create a model to fit the data which consists of 3 Gaussians.
-"""
-model = af.Collection(
- gaussian_0=af.ex.Gaussian, gaussian_1=af.ex.Gaussian, gaussian_2=af.ex.Gaussian
-)
-
-"""
-The `info` attribute shows the model consists of three `Gaussian`'s.
-"""
-print(model.info)
-
-"""
-We repeat the steps above to create the non-linear search and perform the model-fit.
-"""
-search = af.DynestyStatic(
- path_prefix=path.join("features", "model_comparison"),
- name="gaussian_x3",
- nlive=50,
- iterations_per_full_update=3000,
-)
-
-result_x3_gaussian = search.fit(model=model, analysis=analysis)
-
-"""
-The results show that three Gaussians have now been fitted to the data.
-"""
-print(result_x3_gaussian.info)
-
-"""
-Visualizing the fit, we see that there are effectively no residuals, indicating that the model provides a good fit.
-
-By eye, this fit is as good as the 2 Gaussian model above.
-"""
-instance = result_x3_gaussian.max_log_likelihood_instance
-
-gaussian_0 = instance.gaussian_0.model_data_from(xvalues=np.arange(data.shape[0]))
-gaussian_1 = instance.gaussian_0.model_data_from(xvalues=np.arange(data.shape[0]))
-gaussian_2 = instance.gaussian_0.model_data_from(xvalues=np.arange(data.shape[0]))
-model_data = gaussian_0 + gaussian_1 + gaussian_2
-
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- color="k",
- ecolor="k",
- linestyle="",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(range(data.shape[0]), model_data, color="r")
-plt.plot(range(data.shape[0]), gaussian_0, "--")
-plt.plot(range(data.shape[0]), gaussian_1, "--")
-plt.plot(range(data.shape[0]), gaussian_2, "--")
-plt.title("Model fit using 3 Gaussian.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-We print the `log_likelihood` and `log_evidence` of this model-fit, and compare these values to the previous model-fit
-which used 1 and 2 Gaussian.
-"""
-print("1 Gaussian:")
-print(f"Log Likelihood: {max(result_x1_gaussian.samples.log_likelihood_list)}")
-print(f"Log Evidence: {result_x1_gaussian.samples.log_evidence}")
-
-print("2 Gaussians:")
-print(f"Log Likelihood: {max(result_x2_gaussian.samples.log_likelihood_list)}")
-print(f"Log Evidence: {result_x2_gaussian.samples.log_evidence}")
-
-print("3 Gaussians:")
-print(f"Log Likelihood: {max(result_x3_gaussian.samples.log_likelihood_list)}")
-print(f"Log Evidence: {result_x3_gaussian.samples.log_evidence}")
-
-"""
-We now see an interesting result. The `log_likelihood` of the 3 Gaussian model is higher than the 2 Gaussian model
-(albeit, only slightly higher). However, the `log_evidence` is lower than the 2 Gaussian model.
-
-This confirms the behavior discussed at the start of the tutorial. The Bayesian evidence penalizes models with more
-freedom to fit the data, unless they provide a significantly better fit to the data. Using the evidence we favor the
-2 Gaussian model over the 3 Gaussian model for this reason, whereas using the likelihood we favor the 3 Gaussian model.
-
-__Wrap Up__
-
-Discuss Priors. Discuss unique id and benefits of autofit / science workflow.
-"""
+"""
+Feature: Model Comparison
+=========================
+
+Common questions when fitting a model to data are: what model should I use? How many parameters should the model have?
+Is the model too complex or too simple?
+
+Model comparison answers to these questions. It amounts to composing and fitting many different models to the data
+and comparing how well they fit the data.
+
+This example illustrates model comparison using the noisy 1D Gaussian example. We fit a dataset consisting of two
+Gaussians and fit it with three models comprised of 1, 2 and 3 Gaussian's respectively. Using the Bayesian evidence to
+compare the models, we favour the model with 2 Gaussians, which is the "correct" model given that it was the model used
+to simulate the dataset in the first place.
+
+__Metrics__
+
+Different metrics can be used compare models and quantify their goodness-of-fit.
+
+In this example we show the results of using two different metrics:
+
+ - `log_likelihood`: The value returned by the `log_likelihood_function` of an `Analysis` object. which is directly
+ related to the sum of the residuals squared (e.g. the `chi_squared`). The log likelihood does not change when more
+ or less parameters are included in the model, therefore it does not account for over-fitting and will often favour
+ more complex models irrespective of whether they fit the data better.
+
+ - `log_evidence`: The Bayesian evidence, which is closely related to the log likelihood but utilizes additional
+ information which penalizes models based on their complexity. The Bayesian evidence will therefore favour simpler
+ models over more complex models, unless the more complex model provides a much better fit to the data.
+
+__Example Source Code (`af.ex`)__
+
+The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
+
+ - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
+ `visualize` functions.
+
+ - `Gaussian`: a model component representing a 1D Gaussian profile.
+
+These are functionally identical to the `Analysis` and `Gaussian` objects you have seen elsewhere in the workspace.
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Metrics**: Describe the log likelihood and Bayesian evidence metrics used for model comparison.
+- **Example Source Code (`af.ex`)**: The example objects used in this script.
+- **Data**: Load the 1D Gaussian data consisting of two Gaussians.
+- **Model x1 Gaussian**: Create and fit a model with a single Gaussian.
+- **Model x2 Gaussian**: Create and fit a model with two Gaussians.
+- **Model x3 Gaussian**: Create and fit a model with three Gaussians.
+- **Wrap Up**: Summarize the model comparison results using log likelihood and Bayesian evidence.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import matplotlib.pyplot as plt
+import numpy as np
+from os import path
+
+import autofit as af
+
+"""
+__Data__
+
+Load data of a 1D Gaussian from a .json file in the directory `autofit_workspace/dataset/gaussian_x2`.
+
+This 1D data was created using two 1D Gaussians, therefore model comparison should favor a model with two Gaussians over
+a models with 1 or 3 Gaussians.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x2")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+"""
+Plot the data.
+"""
+xvalues = range(data.shape[0])
+
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ color="k",
+ ecolor="k",
+ linestyle="",
+ elinewidth=1,
+ capsize=2,
+)
+plt.title("1D Gaussian Dataset Used For Model Comparison.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+__Model x1 Gaussian__
+
+Create a model to fit the data, starting with a model where the data is fitted with 1 Gaussian.
+"""
+model = af.Collection(gaussian_0=af.ex.Gaussian)
+
+"""
+The `info` attribute shows the model in a readable format, showing it contains one `Gaussian`.
+"""
+print(model.info)
+
+"""
+Create the analysis which fits the model to the data.
+"""
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+"""
+Fit the data using a non-linear search, to determine the goodness of fit of this model.
+
+We use the nested sampling algorithm Dynesty, noting that the Bayesian evidence (`log_evidence`) of a model can only
+be estimated using a nested sampling algorithm.
+"""
+search = af.DynestyStatic(
+ path_prefix=path.join("features", "model_comparison"),
+ name="gaussian_x1",
+ nlive=50,
+ iterations_per_full_update=3000,
+)
+
+"""
+Perform the fit.
+"""
+result_x1_gaussian = search.fit(model=model, analysis=analysis)
+
+"""
+The results are concisely summarised using the `result.info` property.
+
+These show that the parameters of the Gaussian are well constrained, with small errors on their inferred values.
+However, it does not inform us of whether the model provides a good fit to the data overall.
+"""
+print(result_x1_gaussian.info)
+
+"""
+The maximum log likelihood model is used to visualize the fit.
+
+For 1 Gaussian, residuals are visible, whereby the model Gaussian cannot fit the highest central data-point and
+there is a mismatch at the edges of the profile around pixels 40 and 60.
+
+Based on visual inspection, the model therefore provides a poor fit to the data.
+"""
+instance = result_x1_gaussian.max_log_likelihood_instance
+
+gaussian_0 = instance.gaussian_0.model_data_from(xvalues=np.arange(data.shape[0]))
+model_data = gaussian_0
+
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ color="k",
+ ecolor="k",
+ linestyle="",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(range(data.shape[0]), model_data, color="r")
+plt.plot(range(data.shape[0]), gaussian_0, "--")
+plt.title("Model fit using 1 Gaussian.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+Print the `log_likelihood` and `log_evidence` of this model-fit, which we will compare to more complex models in order
+to determine which model provides the best fit to the data.
+"""
+print("1 Gaussian:")
+print(f"Log Likelihood: {result_x1_gaussian.samples.max_log_likelihood()}")
+print(f"Log Evidence: {result_x1_gaussian.samples.log_evidence}")
+
+"""
+__Model x2 Gaussian__
+
+We now create a model to fit the data which consists of 2 Gaussians.
+"""
+model = af.Collection(gaussian_0=af.ex.Gaussian, gaussian_1=af.ex.Gaussian)
+
+"""
+The `info` attribute shows the model now consists of two `Gaussian`'s.
+"""
+print(model.info)
+
+"""
+We repeat the steps above to create the non-linear search and perform the model-fit.
+"""
+search = af.DynestyStatic(
+ path_prefix=path.join("features", "model_comparison"),
+ name="gaussian_x2",
+ nlive=50,
+ iterations_per_full_update=3000,
+)
+
+result_x2_gaussian = search.fit(model=model, analysis=analysis)
+
+"""
+The results show that two Gaussians have now been fitted to the data.
+"""
+print(result_x2_gaussian.info)
+
+"""
+Visualizing the fit, we see that the problems with the previous fit have been addressed. The central data-point at the
+highest normalization is fitted correctly and the residuals at the edges of the profile around pixels 40 and 60 are
+significantly reduced.
+
+There are effectively no residuals, indicating that the model provides a good fit to the data.
+
+The residuals are so small that they are consistent with noise in the data. One therefore should not expect that
+a more complex model than one with 2 Gaussians can provide a better fit.
+"""
+instance = result_x2_gaussian.max_log_likelihood_instance
+
+gaussian_0 = instance.gaussian_0.model_data_from(xvalues=np.arange(data.shape[0]))
+gaussian_1 = instance.gaussian_0.model_data_from(xvalues=np.arange(data.shape[0]))
+model_data = gaussian_0 + gaussian_1
+
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ color="k",
+ ecolor="k",
+ linestyle="",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(range(data.shape[0]), model_data, color="r")
+plt.plot(range(data.shape[0]), gaussian_0, "--")
+plt.plot(range(data.shape[0]), gaussian_1, "--")
+plt.title("Model fit using 2 Gaussian.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+Print the `log_likelihood` and `log_evidence` of this model-fit, and compare these values to the previous model-fit
+which used 1 Gaussian.
+"""
+print("1 Gaussian:")
+print(f"Log Likelihood: {max(result_x1_gaussian.samples.log_likelihood_list)}")
+print(f"Log Evidence: {result_x1_gaussian.samples.log_evidence}")
+
+print("2 Gaussians:")
+print(f"Log Likelihood: {max(result_x2_gaussian.samples.log_likelihood_list)}")
+print(f"Log Evidence: {result_x2_gaussian.samples.log_evidence}")
+
+"""
+Both the `log_likelihood` and `log_evidence` have increased significantly, indicating that the model with 2 Gaussians
+is favored over the model with 1 Gaussian.
+
+This is expected, as we know the data was generated using 2 Gaussians!
+
+__Model x3 Gaussian__
+
+We now create a model to fit the data which consists of 3 Gaussians.
+"""
+model = af.Collection(
+ gaussian_0=af.ex.Gaussian, gaussian_1=af.ex.Gaussian, gaussian_2=af.ex.Gaussian
+)
+
+"""
+The `info` attribute shows the model consists of three `Gaussian`'s.
+"""
+print(model.info)
+
+"""
+We repeat the steps above to create the non-linear search and perform the model-fit.
+"""
+search = af.DynestyStatic(
+ path_prefix=path.join("features", "model_comparison"),
+ name="gaussian_x3",
+ nlive=50,
+ iterations_per_full_update=3000,
+)
+
+result_x3_gaussian = search.fit(model=model, analysis=analysis)
+
+"""
+The results show that three Gaussians have now been fitted to the data.
+"""
+print(result_x3_gaussian.info)
+
+"""
+Visualizing the fit, we see that there are effectively no residuals, indicating that the model provides a good fit.
+
+By eye, this fit is as good as the 2 Gaussian model above.
+"""
+instance = result_x3_gaussian.max_log_likelihood_instance
+
+gaussian_0 = instance.gaussian_0.model_data_from(xvalues=np.arange(data.shape[0]))
+gaussian_1 = instance.gaussian_0.model_data_from(xvalues=np.arange(data.shape[0]))
+gaussian_2 = instance.gaussian_0.model_data_from(xvalues=np.arange(data.shape[0]))
+model_data = gaussian_0 + gaussian_1 + gaussian_2
+
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ color="k",
+ ecolor="k",
+ linestyle="",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(range(data.shape[0]), model_data, color="r")
+plt.plot(range(data.shape[0]), gaussian_0, "--")
+plt.plot(range(data.shape[0]), gaussian_1, "--")
+plt.plot(range(data.shape[0]), gaussian_2, "--")
+plt.title("Model fit using 3 Gaussian.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+We print the `log_likelihood` and `log_evidence` of this model-fit, and compare these values to the previous model-fit
+which used 1 and 2 Gaussian.
+"""
+print("1 Gaussian:")
+print(f"Log Likelihood: {max(result_x1_gaussian.samples.log_likelihood_list)}")
+print(f"Log Evidence: {result_x1_gaussian.samples.log_evidence}")
+
+print("2 Gaussians:")
+print(f"Log Likelihood: {max(result_x2_gaussian.samples.log_likelihood_list)}")
+print(f"Log Evidence: {result_x2_gaussian.samples.log_evidence}")
+
+print("3 Gaussians:")
+print(f"Log Likelihood: {max(result_x3_gaussian.samples.log_likelihood_list)}")
+print(f"Log Evidence: {result_x3_gaussian.samples.log_evidence}")
+
+"""
+We now see an interesting result. The `log_likelihood` of the 3 Gaussian model is higher than the 2 Gaussian model
+(albeit, only slightly higher). However, the `log_evidence` is lower than the 2 Gaussian model.
+
+This confirms the behavior discussed at the start of the tutorial. The Bayesian evidence penalizes models with more
+freedom to fit the data, unless they provide a significantly better fit to the data. Using the evidence we favor the
+2 Gaussian model over the 3 Gaussian model for this reason, whereas using the likelihood we favor the 3 Gaussian model.
+
+__Wrap Up__
+
+Discuss Priors. Discuss unique id and benefits of autofit / science workflow.
+"""
diff --git a/scripts/features/search_chaining.py b/scripts/features/search_chaining.py
index f39aec91..db973a53 100644
--- a/scripts/features/search_chaining.py
+++ b/scripts/features/search_chaining.py
@@ -1,665 +1,657 @@
-"""
-Feature: Search Chaining
-========================
-
-To perform a model-fit, we typically compose one model and fit it to our data using one non-linear search.
-
-Search chaining fits many different models to a dataset using a chained sequence of non-linear searches. Initial
-fits are performed using simplified model parameterizations and faster non-linear fitting techniques. The results of
-these simplified fits can then be used to initialize fits using a higher dimensionality model with more detailed
-non-linear search.
-
-To fit highly complex models our aim is therefore to granularize the fitting procedure into a series of **bite-sized**
-searches which are faster and more reliable than fitting the more complex model straight away.
-
-Our ability to construct chained non-linear searches that perform model fitting more accurately and efficiently relies
-on our **domain specific knowledge** of the model fitting task. For example, we may know that our dataset contains
-multiple features that can be accurately fitted separately before performing a joint fit, or that certain parameter
-share minimal covariance such that certain parameters can be fixed before fitting both with a more complex model
-parameterization.
-
-We may also know tricks that can speed up the fitting of the initial model, for example reducing the size of the data
-or changing the likelihood evaluations in a way that makes them quicker (likely at the expense of the overall
-quality of the fit itself). By using chained searches these speed-ups can be relaxed towards the end of the
-model-fitting sequence when we want the most precise, most accurate model that best fits the dataset available.
-
-In this example we demonstrate search chaining using the example data where there are two `Gaussians` that are visibly
-split. Instead of fitting them simultaneously using a single non-linear search consisting of N=6 parameters, we break
-this model-fit into a chained of three searches where:
-
- 1) The first model fits just the left `Gaussian` where N=3.
- 2) The first model fits just the right `Gaussian` where again N=3.
- 3) The final model is fitted with both `Gaussians` where N=6. Crucially, the results of the first two searches
- are used to initialize the search and tell it the highest likelihood regions of parameter space.
-
-By initially fitting parameter spaces of reduced complexity we can achieve a more efficient and reliable model-fitting
-procedure.
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Example Source Code (`af.ex`)**: The example objects used in this script.
-- **Data**: Load the 1D data containing two split Gaussians.
-- **Analysis**: Create the Analysis class for fitting the model to data.
-- **Model**: Define the model for the left Gaussian.
-- **Search 1**: Perform the first search fitting the left Gaussian (N=3).
-- **Result 1**: Examine the results of the first search.
-- **Search 2**: Perform the second search fitting the right Gaussian (N=3).
-- **Model**: Define the model for the right Gaussian using the result of Search 1.
-- **Result 2**: Examine the results of the second search.
-- **Search 3**: Perform the final search fitting both Gaussians (N=6) using prior results.
-- **Prior Passing**: Explain how priors are passed between chained searches.
-- **EXAMPLE**: A concrete example of prior passing with numerical values.
-- **Prerequisites**: Prerequisites for prior passing concepts.
-- **Overview**: Overview of search chaining and prior passing.
-- **Model-Fit**: The model-fit used in prior passing examples.
-- **Instance & Model**: Explain how results are passed as instances and models.
-- **Component Specification**: Specify which model component priors to pass.
-- **Take Attributes**: Use the take_attributes method to pass priors between different model components.
-- **As Model**: Use the as_model method to create a model from a result instance.
-
-__Example Source Code (`af.ex`)__
-
-The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
-
- - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
- `visualize` functions.
-
- - `Gaussian`: a model component representing a 1D Gaussian profile.
-
-These are functionally identical to the `Analysis` and `Gaussian` objects you have seen elsewhere in the workspace.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import matplotlib.pyplot as plt
-import numpy as np
-from os import path
-
-import autofit as af
-
-"""
-__Data__
-
-Load data of two 1D Gaussians, by loading it from a .json file in the directory
-`autofit_workspace/dataset/gaussian_x1_with_feature`.
-
-This 1D data includes two `Gaussians` that are split from one another.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x2_split")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-"""
-Lets plot the data.
-
-Two separate `Gaussians` are clearly visible.
-"""
-xvalues = range(data.shape[0])
-
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.title("1D Gaussian Data With two Gaussians split apart.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-__Analysis__
-
-Create the analysis which fits the model to the data.
-
-It fits the data as the sum of as many `Gaussian`'s as are in the model.
-
-To better fit the left gaussian, we remove all data points in the right-half of the data. Note that for more
-computationally demanding model-fitting problems this would give a significant speed-up in log likelihood function.
-"""
-analysis_1 = af.ex.Analysis(data=data[0:50], noise_map=noise_map[0:50])
-
-"""
-__Model__
-
-We are now going to fit the left `Gaussian` in this split dataset.
-"""
-model_1 = af.Collection(gaussian_left=af.ex.Gaussian)
-
-"""
-The `info` attribute shows the model in a readable format.
-"""
-print(model_1.info)
-
-"""
-__Search 1__
-
-Fit the data with the `left_gaussian` using a single non-linear search.
-
-Given the simplicity of the model, we can use a low number of live points to achieve a fast model-fit.
-"""
-search_1 = af.DynestyStatic(
- name="search[1]__left_gaussian",
- path_prefix=path.join("features", "search_chaining"),
- nlive=30,
-)
-
-result_1 = search_1.fit(model=model_1, analysis=analysis_1)
-
-"""
-__Result 1__
-
-The `info` attribute shows the result in a readable format.
-"""
-print(result_1.info)
-
-"""
-By plotting the result we can see we have fitted the left `Gaussian` reasonably well.
-"""
-instance = result_1.max_log_likelihood_instance
-
-model_data = instance.gaussian_left.model_data_from(xvalues=np.arange(data.shape[0]))
-
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(range(data.shape[0]), model_data, color="r")
-plt.title("Search 1 fit to left Gaussian.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-__Search 2 __
-
-We now repeat the above process for the right `Gaussian`.
-
-We could remove the data on the left like we did the `Gaussian` above. However, we are instead going to fit the full
-dataset.
-"""
-analysis_2 = af.ex.Analysis(data=data, noise_map=noise_map)
-
-"""
-__Model__
-
-Because the left Gaussian is now again in the data, we need to fit it. We can do this, without increasing the
-dimensionality of our parameter space to N=6, by using the maximum log likelihood model of the `gaussian_left` in
-search[1].
-
-For search chaining, **PyAutoFit** has many convenient methods for passing the results of a search to a subsequence
-search. Below, we achieve this by passing the result of the search above as an `instance.
-"""
-model_2 = af.Collection(
- gaussian_left=result_1.instance.gaussian_left, gaussian_right=af.ex.Gaussian
-)
-
-"""
-The `info` attribute shows the model, including how parameters and priors were passed from `result_1`.
-"""
-print(model_2.info)
-
-"""
-We now run our second Dynesty search to fit the right `Gaussian`.
-
-Given the simplicity of the model, we can again use a low number of live points to achieve a fast model-fit.
-"""
-search_2 = af.DynestyStatic(
- name="search[2]__right_gaussian",
- path_prefix=path.join("features", "search_chaining"),
- nlive=30,
-)
-
-result_2 = search_2.fit(model=model_2, analysis=analysis_2)
-
-"""
-__Result 2__
-
-The `info` attribute shows the result in a readable format.
-"""
-print(result_2.info)
-
-"""
-By plotting the result we can see we have fitted the right `Gaussian` reasonably well and that the model includes the
-`left_gaussian` from the first search.
-"""
-instance = result_2.max_log_likelihood_instance
-
-gaussian_left = instance.gaussian_left.model_data_from(xvalues=np.arange(data.shape[0]))
-gaussian_right = instance.gaussian_right.model_data_from(
- xvalues=np.arange(data.shape[0])
-)
-
-model_data = gaussian_left + gaussian_right
-
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(range(data.shape[0]), model_data, color="r")
-plt.plot(range(data.shape[0]), gaussian_left, "--")
-plt.plot(range(data.shape[0]), gaussian_right, "--")
-plt.title("Search 2 fit to right Gaussian.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-__Search 3__
-
-We now fit both `Gaussians`'s simultaneously, using the results of the previous two searches to initialize where
-the non-linear searches parameter space.
-
-To pass the result in this way we use the command `result.model`, which in contrast to `result.instance` above passes
-the parameters not as the maximum log likelihood values but as `TruncatedGaussianPrior`'s that are fitted for by the
-non-linear search. We discuss below how this `TruncatedGaussianPrior` is passed.
-"""
-
-model_3 = af.Collection(
- gaussian_left=result_1.model.gaussian_left,
- gaussian_right=result_2.model.gaussian_right,
-)
-
-"""
-The `info` attribute shows the model, including how parameters and priors were passed from `result_1` and `result_2`.
-"""
-print(model_3.info)
-
-"""
-We now perform the search.
-"""
-analysis_3 = af.ex.Analysis(data=data, noise_map=noise_map)
-
-search_3 = af.DynestyStatic(
- name="search[3]__both_gaussians",
- path_prefix=path.join("features", "search_chaining"),
- nlive=100,
-)
-
-result_3 = search_3.fit(model=model_3, analysis=analysis_3)
-
-"""
-By plotting the result we can now see we have fitted both `Gaussian`'s accurately.
-"""
-instance = result_3.max_log_likelihood_instance
-
-gaussian_left = instance.gaussian_left.model_data_from(xvalues=np.arange(data.shape[0]))
-gaussian_right = instance.gaussian_right.model_data_from(
- xvalues=np.arange(data.shape[0])
-)
-
-model_data = gaussian_left + gaussian_right
-
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(range(data.shape[0]), model_data, color="r")
-plt.plot(range(data.shape[0]), gaussian_left, "--")
-plt.plot(range(data.shape[0]), gaussian_right, "--")
-plt.title("Search 3 fit to both Gaussian.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-__Prior Passing__
-
-Now search 3 is complete, you should checkout its `model.info` file. The parameters do not use the default priors of
-the `Gaussian` model component. Instead, they use TruncatedGaussianPrior`s where:
-
- - The mean values are the median PDF results of every parameter inferred by the fits performed in searches 1 and 2.
- - They sigma values are the errors computed by these searches, or they are values higher than these errors.
-
-The sigma values uses the errors of searches 1 and 2 for an obvious reason, this is a reasonable estimate of where in
-parameter space the model-fit can be expected to provide a good fit to the data. However, we may want to specify
-even larger sigma values on certain parameters, if for example we anticipate that our earlier searches may under
-estimate the errors.
-
-The `width_modifier` term in the `Gaussian` section of the `config/priors/gaussian.yaml` is used instead of the errors
-of a search, when the errors estimated are smaller than the `width_modifier` value. This ensure that the sigma
-values used in later searches do not assume extremely small values if earlier searches risk under estimating the errors.
-
-Thus, search 3 used the results of searches 1 and 2 to inform it where to search non-linear parameter space!
-
-The `prior_passer` settings in the `general.yaml` config customizes how priors are passed from a search as follows:
-
- - sigma: The sigma value of the errors passed to set the sigma values in the previous search are estimated at.
- - use_widths: If False, the "width_modifier" values in the json_prior configs are not used to override a passed
- error value.
- - use_errors: If False, errors are not passed from search 1 to set up the priors and only the `width_modifier`
- entries in the configs are used.
-
-There are two ways a value is specified using the priors/width file:
-
- 1) Absolute: In this case, the error assumed on the parameter is the value given in the config file.
- For example, if for the width on `centre` the width modifier reads "Absolute" with a value 20.0, this means if the
- error on the parameter `centre` was less than 20.0 in the previous search, the sigma of its `TruncatedGaussianPrior` in
- the next search will be 20.0.
-
- 2) Relative: In this case, the error assumed on the parameter is the % of the value of the estimate value given in
- the config file. For example, if the normalization estimated in the previous search was 2.0, and the relative error in
- the config file reads "Relative" with a value 0.5, then the sigma of the `TruncatedGaussianPrior`
- will be 50% of this value, i.e. sigma = 0.5 * 2.0 = 1.0.
-
-We use absolute and relative values for different parameters, depending on their properties. For example, using the
-relative value of a parameter like the `centre` makes no sense as the profile could be centred at 0.0, making
-the relative error tiny and poorly defined.
-
-However, there are parameters where using an absolute value does not make sense. Normalization is a good example of this.
-The normalization of an image depends on its units and S/N. There is no single absolute value that one can use to
-generically chain the normalization of any two proflies. Thus, it makes more sense to chain them using the relative value
-from a previous search.
-
-We can customize how priors are passed from the results of a search editing the `prior_passer` settings in
-the `general.yaml` config.
-
-__EXAMPLE__
-
-Lets go through an example using a real parameter. Lets say in search 1 we fit a `Gaussian` and we estimate that
-its normalization is equal to 4.0 +- 2.0 where the error value of 2.0 was computed at 3.0 sigma confidence. To pass this
-as a prior to search 2, we would write:
-
- gaussian.normalization = search_1_result.model.gaussian.normalization
-
-The prior on the `Gaussian` `normalization` in search 2 would thus be a `TruncatedGaussianPrior`, with mean=4.0 and
-sigma=2.0. If we had used a sigma value of 1.0 to compute the error, which reduced the estimate from 4.0 +- 2.0 to
-4.0 +- 1.0, the sigma of the `TruncatedGaussianPrior` would instead be 1.0.
-
-If the error on the normalization in search 1 had been really small, lets say, 0.01, we would instead use the value of the
-normalization width in the priors config file to set sigma instead. In this case, the prior config file specifies
-that we use an "Relative" value of 0.5 to chain this prior. Thus, the TruncatedGaussianPrior in search 2 would have a mean=4.0
-and sigma=2.0.
-
-And with that, we`re done. Chaining searches is a bit of an art form, but for certain problems can be extremely
-powerful.
-"""
-
-"""
-Cookbook 5: Model Linking
-=========================
-
-__Prerequisites__
-
-You should be familiar with the search chaining API detailed in the following scripts and docs:
-
-__Overview__
-
-Search chaining allows one to perform back-to-back non-linear searches to fit a dataset, where the model complexity
-increases after each fit.
-
-To perform search chaining, **PyAutoFit** has tools for passing the results of one model-fit from one fit to the next,
-and change its parameterization between each fit.
-
-This cookbook is a concise reference to the model linking API.
-"""
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import json
-import os
-from os import path
-
-import autofit as af
-
-"""
-__Model-Fit__
-
-We perform a quick model-fit, to create a `Result` object which has the attributes necessary to illustrate the model
-linking API.
-"""
-model = af.Collection(gaussian=af.ex.Gaussian, exponential=af.ex.Exponential)
-
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1__exponential_x1")
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-dynesty = af.DynestyStatic(name="cookbook_5_model_linking", nlive=50, sample="rwalk")
-
-result = dynesty.fit(model=model, analysis=analysis)
-
-"""
-__Instance & Model__
-
-The result object has two key attributes for model linking:
-
- - `instance`: The maximum log likelihood instance of the model-fit, where every parameter is therefore a float.
-
- - `model`: An attribute which represents how the result can be passed as a model-component to the next fit (the
- details of how its priors are passed are given in full below).
-
-We create a new model using both of these attributes, where:
-
- - All of the `gaussian` model components parameters are passed via the `instance` attribute and therefore fixed to
- the inferred maximum log likelihood values (and are not free parameters in the model).
-
- - All of the `exponential` model components parameters are passed via the `model` attribute and therefore are free
- parameters in the model.
-
-The new model therefore has 3 free parameters and 3 fixed parameters.
-"""
-model = af.Collection(gaussian=af.ex.Gaussian, exponential=af.ex.Exponential)
-
-model.gaussian = result.instance.gaussian
-model.exponential = result.model.exponential
-
-"""
-The `model.info` attribute shows that the parameter and prior passing has occurred as described above.
-"""
-print(model.info)
-
-"""
-We can print the priors of the exponenital:
-"""
-print("Exponential Model Priors \n")
-print("centre = ", model.exponential.centre)
-print("normalization = ", model.exponential.normalization)
-print("rate = ", model.exponential.rate)
-
-"""
-How are the priors set via model linking? The full description is quite long, therefore it is attatched to the
-bottom of this script so that we can focus on the model linking API.
-
-__Component Specification__
-
-Model linking can be performed on any component of a model, for example to only pass specific parameters as
-an `instance` or `model`.
-"""
-gaussian = af.Model(af.ex.Gaussian)
-
-gaussian.centre = result.instance.gaussian.centre
-gaussian.normalization = result.model.gaussian.normalization
-gaussian.sigma = result.instance.gaussian.sigma
-
-exponential = af.Model(af.ex.Exponential)
-
-exponential.centre = result.model.exponential.centre
-exponential.normalization = result.model.exponential.normalization
-exponential.rate = result.instance.exponential.rate
-
-model = af.Collection(gaussian=gaussian, exponential=exponential)
-
-"""
-The `model.info` attribute shows that the parameter and prior passing has occurred on individual components.
-"""
-print(model.info)
-
-"""
-__Take Attributes__
-
-The examples above linked models where the individual model components that were passed stayed the same.
-
-We can link two related models, where only a subset of parameters are shared, by using the `take_attributes()` method.
-
-For example, lets define a `GaussianKurtosis` which is a `Gaussian` with an extra parameter for its kurtosis.
-"""
-
-
-class GaussianKurtosis:
- def __init__(
- self,
- centre=30.0, # <- **PyAutoFit** recognises these constructor arguments
- normalization=1.0, # <- are the Gaussian``s model parameters.
- sigma=5.0,
- kurtosis=1.0,
- ):
- self.centre = centre
- self.normalization = normalization
- self.sigma = sigma
- self.kurtosis = kurtosis
-
-
-"""
-The `take_attributes()` method takes a `source` model component, and inspects the names of all its parameters.
-
-For the `Gaussian` model result input below, it finds the parameters `centre`, `normalization` and `sigma`.
-
-It then finds all parameters in the new `model` which have the same names, which for the `GaussianKurtosis` is
-`centre`, `normalization` and `sigma`.
-
-For all parameters which have the same name, the parameter is passed.
-"""
-model = af.Collection(gaussian=af.Model(GaussianKurtosis))
-model.kurtosis = af.UniformPrior(lower_limit=-1.0, upper_limit=1.0)
-
-model.gaussian.take_attributes(source=result.model.gaussian)
-
-"""
-Because the result was passed using `model` we see the priors on the `GaussianKurtosis` `centre`,
-`normalization` and `sigma` have been updated, whereas its `kurtosis` has not.
-"""
-print("GaussianKurtosis Model Priors After Take Attributes via Model \n")
-print("centre = ", model.gaussian.centre)
-print("normalization = ", model.gaussian.normalization)
-print("sigma = ", model.gaussian.sigma)
-print("kurtosis = ", model.gaussian.kurtosis)
-
-"""
-If we pass `result.instance` to take_attributes the same name linking is used, however parameters are passed as
-floats.
-"""
-model = af.Collection(gaussian=af.Model(GaussianKurtosis))
-model.kurtosis = af.UniformPrior(lower_limit=-1.0, upper_limit=1.0)
-
-model.gaussian.take_attributes(source=result.instance.gaussian)
-
-print("Gaussian Model Priors After Take Attributes via Instance \n")
-print("centre = ", model.gaussian.centre)
-print("normalization = ", model.gaussian.normalization)
-print("sigma = ", model.gaussian.sigma)
-print("kurtosis = ", model.gaussian.kurtosis)
-
-"""
-__As Model__
-
-A common problem is when we have an `instance` (e.g. from a previous fit where we fixed the parameters)
-but now wish to make its parameters free parameters again.
-
-Furthermore, we may want to do this for specific model components.
-
-The `as_model` method allows us to do this. Below, we pass the entire result (e.g. both the `gaussian`
-and `exponential` components), however we pass the `Gaussian` class to `as_model`, meaning that any model
-component in the `instance` which is a `Gaussian` will be converted to a model with free parameters.
-"""
-model = result.instance.as_model((af.ex.Gaussian,))
-
-print("Gaussian Model Priors After via as_model: \n")
-print("centre = ", model.gaussian.centre)
-print("normalization = ", model.gaussian.normalization)
-print("sigma = ", model.gaussian.sigma)
-print("centre = ", model.exponential.centre)
-print("normalization = ", model.exponential.normalization)
-print("rate= ", model.exponential.rate)
-
-"""
-The `as_model()` method does not have too much utility for the simple model used in this cookbook.
-
-However, for multi-level models with many components, it is a powerful tool to compose custom models.
-"""
-
-
-class MultiLevelProfiles:
- def __init__(
- self,
- higher_level_centre=50.0, # This is the centre of all Gaussians in this multi level component.
- profile_list=None, # This will contain a list of model-components
- ):
- self.higher_level_centre = higher_level_centre
-
- self.profile_list = profile_list
-
-
-group_level_0 = af.Model(
- MultiLevelProfiles, profile_list=[af.ex.Gaussian, af.ex.Exponential, af.ex.Gaussian]
-)
-
-group_level_1 = af.Model(
- MultiLevelProfiles,
- profile_list=[af.ex.Gaussian, af.ex.Exponential, af.ex.Exponential],
-)
-
-model = af.Collection(group_level_0=group_level_0, group_level_1=group_level_1)
-
-"""
-This means every `Gaussian` in the complex multi-level model above would have parameters set via the result of our
-model-fit, if the model above was fitted such that it was contained in the result.
-"""
-model = result.instance.as_model((af.ex.Gaussian,))
+"""
+Feature: Search Chaining
+========================
+
+To perform a model-fit, we typically compose one model and fit it to our data using one non-linear search.
+
+Search chaining fits many different models to a dataset using a chained sequence of non-linear searches. Initial
+fits are performed using simplified model parameterizations and faster non-linear fitting techniques. The results of
+these simplified fits can then be used to initialize fits using a higher dimensionality model with more detailed
+non-linear search.
+
+To fit highly complex models our aim is therefore to granularize the fitting procedure into a series of **bite-sized**
+searches which are faster and more reliable than fitting the more complex model straight away.
+
+Our ability to construct chained non-linear searches that perform model fitting more accurately and efficiently relies
+on our **domain specific knowledge** of the model fitting task. For example, we may know that our dataset contains
+multiple features that can be accurately fitted separately before performing a joint fit, or that certain parameter
+share minimal covariance such that certain parameters can be fixed before fitting both with a more complex model
+parameterization.
+
+We may also know tricks that can speed up the fitting of the initial model, for example reducing the size of the data
+or changing the likelihood evaluations in a way that makes them quicker (likely at the expense of the overall
+quality of the fit itself). By using chained searches these speed-ups can be relaxed towards the end of the
+model-fitting sequence when we want the most precise, most accurate model that best fits the dataset available.
+
+In this example we demonstrate search chaining using the example data where there are two `Gaussians` that are visibly
+split. Instead of fitting them simultaneously using a single non-linear search consisting of N=6 parameters, we break
+this model-fit into a chained of three searches where:
+
+ 1) The first model fits just the left `Gaussian` where N=3.
+ 2) The first model fits just the right `Gaussian` where again N=3.
+ 3) The final model is fitted with both `Gaussians` where N=6. Crucially, the results of the first two searches
+ are used to initialize the search and tell it the highest likelihood regions of parameter space.
+
+By initially fitting parameter spaces of reduced complexity we can achieve a more efficient and reliable model-fitting
+procedure.
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Example Source Code (`af.ex`)**: The example objects used in this script.
+- **Data**: Load the 1D data containing two split Gaussians.
+- **Analysis**: Create the Analysis class for fitting the model to data.
+- **Model**: Define the model for the left Gaussian.
+- **Search 1**: Perform the first search fitting the left Gaussian (N=3).
+- **Result 1**: Examine the results of the first search.
+- **Search 2**: Perform the second search fitting the right Gaussian (N=3).
+- **Model**: Define the model for the right Gaussian using the result of Search 1.
+- **Result 2**: Examine the results of the second search.
+- **Search 3**: Perform the final search fitting both Gaussians (N=6) using prior results.
+- **Prior Passing**: Explain how priors are passed between chained searches.
+- **EXAMPLE**: A concrete example of prior passing with numerical values.
+- **Prerequisites**: Prerequisites for prior passing concepts.
+- **Overview**: Overview of search chaining and prior passing.
+- **Model-Fit**: The model-fit used in prior passing examples.
+- **Instance & Model**: Explain how results are passed as instances and models.
+- **Component Specification**: Specify which model component priors to pass.
+- **Take Attributes**: Use the take_attributes method to pass priors between different model components.
+- **As Model**: Use the as_model method to create a model from a result instance.
+
+__Example Source Code (`af.ex`)__
+
+The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
+
+ - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
+ `visualize` functions.
+
+ - `Gaussian`: a model component representing a 1D Gaussian profile.
+
+These are functionally identical to the `Analysis` and `Gaussian` objects you have seen elsewhere in the workspace.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import matplotlib.pyplot as plt
+import numpy as np
+from os import path
+
+import autofit as af
+
+"""
+__Data__
+
+Load data of two 1D Gaussians, by loading it from a .json file in the directory
+`autofit_workspace/dataset/gaussian_x1_with_feature`.
+
+This 1D data includes two `Gaussians` that are split from one another.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x2_split")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+"""
+Lets plot the data.
+
+Two separate `Gaussians` are clearly visible.
+"""
+xvalues = range(data.shape[0])
+
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.title("1D Gaussian Data With two Gaussians split apart.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+__Analysis__
+
+Create the analysis which fits the model to the data.
+
+It fits the data as the sum of as many `Gaussian`'s as are in the model.
+
+To better fit the left gaussian, we remove all data points in the right-half of the data. Note that for more
+computationally demanding model-fitting problems this would give a significant speed-up in log likelihood function.
+"""
+analysis_1 = af.ex.Analysis(data=data[0:50], noise_map=noise_map[0:50])
+
+"""
+__Model__
+
+We are now going to fit the left `Gaussian` in this split dataset.
+"""
+model_1 = af.Collection(gaussian_left=af.ex.Gaussian)
+
+"""
+The `info` attribute shows the model in a readable format.
+"""
+print(model_1.info)
+
+"""
+__Search 1__
+
+Fit the data with the `left_gaussian` using a single non-linear search.
+
+Given the simplicity of the model, we can use a low number of live points to achieve a fast model-fit.
+"""
+search_1 = af.DynestyStatic(
+ name="search[1]__left_gaussian",
+ path_prefix=path.join("features", "search_chaining"),
+ nlive=30,
+)
+
+result_1 = search_1.fit(model=model_1, analysis=analysis_1)
+
+"""
+__Result 1__
+
+The `info` attribute shows the result in a readable format.
+"""
+print(result_1.info)
+
+"""
+By plotting the result we can see we have fitted the left `Gaussian` reasonably well.
+"""
+instance = result_1.max_log_likelihood_instance
+
+model_data = instance.gaussian_left.model_data_from(xvalues=np.arange(data.shape[0]))
+
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(range(data.shape[0]), model_data, color="r")
+plt.title("Search 1 fit to left Gaussian.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+__Search 2 __
+
+We now repeat the above process for the right `Gaussian`.
+
+We could remove the data on the left like we did the `Gaussian` above. However, we are instead going to fit the full
+dataset.
+"""
+analysis_2 = af.ex.Analysis(data=data, noise_map=noise_map)
+
+"""
+__Model__
+
+Because the left Gaussian is now again in the data, we need to fit it. We can do this, without increasing the
+dimensionality of our parameter space to N=6, by using the maximum log likelihood model of the `gaussian_left` in
+search[1].
+
+For search chaining, **PyAutoFit** has many convenient methods for passing the results of a search to a subsequence
+search. Below, we achieve this by passing the result of the search above as an `instance.
+"""
+model_2 = af.Collection(
+ gaussian_left=result_1.instance.gaussian_left, gaussian_right=af.ex.Gaussian
+)
+
+"""
+The `info` attribute shows the model, including how parameters and priors were passed from `result_1`.
+"""
+print(model_2.info)
+
+"""
+We now run our second Dynesty search to fit the right `Gaussian`.
+
+Given the simplicity of the model, we can again use a low number of live points to achieve a fast model-fit.
+"""
+search_2 = af.DynestyStatic(
+ name="search[2]__right_gaussian",
+ path_prefix=path.join("features", "search_chaining"),
+ nlive=30,
+)
+
+result_2 = search_2.fit(model=model_2, analysis=analysis_2)
+
+"""
+__Result 2__
+
+The `info` attribute shows the result in a readable format.
+"""
+print(result_2.info)
+
+"""
+By plotting the result we can see we have fitted the right `Gaussian` reasonably well and that the model includes the
+`left_gaussian` from the first search.
+"""
+instance = result_2.max_log_likelihood_instance
+
+gaussian_left = instance.gaussian_left.model_data_from(xvalues=np.arange(data.shape[0]))
+gaussian_right = instance.gaussian_right.model_data_from(
+ xvalues=np.arange(data.shape[0])
+)
+
+model_data = gaussian_left + gaussian_right
+
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(range(data.shape[0]), model_data, color="r")
+plt.plot(range(data.shape[0]), gaussian_left, "--")
+plt.plot(range(data.shape[0]), gaussian_right, "--")
+plt.title("Search 2 fit to right Gaussian.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+__Search 3__
+
+We now fit both `Gaussians`'s simultaneously, using the results of the previous two searches to initialize where
+the non-linear searches parameter space.
+
+To pass the result in this way we use the command `result.model`, which in contrast to `result.instance` above passes
+the parameters not as the maximum log likelihood values but as `TruncatedGaussianPrior`'s that are fitted for by the
+non-linear search. We discuss below how this `TruncatedGaussianPrior` is passed.
+"""
+
+model_3 = af.Collection(
+ gaussian_left=result_1.model.gaussian_left,
+ gaussian_right=result_2.model.gaussian_right,
+)
+
+"""
+The `info` attribute shows the model, including how parameters and priors were passed from `result_1` and `result_2`.
+"""
+print(model_3.info)
+
+"""
+We now perform the search.
+"""
+analysis_3 = af.ex.Analysis(data=data, noise_map=noise_map)
+
+search_3 = af.DynestyStatic(
+ name="search[3]__both_gaussians",
+ path_prefix=path.join("features", "search_chaining"),
+ nlive=100,
+)
+
+result_3 = search_3.fit(model=model_3, analysis=analysis_3)
+
+"""
+By plotting the result we can now see we have fitted both `Gaussian`'s accurately.
+"""
+instance = result_3.max_log_likelihood_instance
+
+gaussian_left = instance.gaussian_left.model_data_from(xvalues=np.arange(data.shape[0]))
+gaussian_right = instance.gaussian_right.model_data_from(
+ xvalues=np.arange(data.shape[0])
+)
+
+model_data = gaussian_left + gaussian_right
+
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(range(data.shape[0]), model_data, color="r")
+plt.plot(range(data.shape[0]), gaussian_left, "--")
+plt.plot(range(data.shape[0]), gaussian_right, "--")
+plt.title("Search 3 fit to both Gaussian.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+__Prior Passing__
+
+Now search 3 is complete, you should checkout its `model.info` file. The parameters do not use the default priors of
+the `Gaussian` model component. Instead, they use TruncatedGaussianPrior`s where:
+
+ - The mean values are the median PDF results of every parameter inferred by the fits performed in searches 1 and 2.
+ - They sigma values are the errors computed by these searches, or they are values higher than these errors.
+
+The sigma values uses the errors of searches 1 and 2 for an obvious reason, this is a reasonable estimate of where in
+parameter space the model-fit can be expected to provide a good fit to the data. However, we may want to specify
+even larger sigma values on certain parameters, if for example we anticipate that our earlier searches may under
+estimate the errors.
+
+The `width_modifier` term in the `Gaussian` section of the `config/priors/gaussian.yaml` is used instead of the errors
+of a search, when the errors estimated are smaller than the `width_modifier` value. This ensure that the sigma
+values used in later searches do not assume extremely small values if earlier searches risk under estimating the errors.
+
+Thus, search 3 used the results of searches 1 and 2 to inform it where to search non-linear parameter space!
+
+The `prior_passer` settings in the `general.yaml` config customizes how priors are passed from a search as follows:
+
+ - sigma: The sigma value of the errors passed to set the sigma values in the previous search are estimated at.
+ - use_widths: If False, the "width_modifier" values in the json_prior configs are not used to override a passed
+ error value.
+ - use_errors: If False, errors are not passed from search 1 to set up the priors and only the `width_modifier`
+ entries in the configs are used.
+
+There are two ways a value is specified using the priors/width file:
+
+ 1) Absolute: In this case, the error assumed on the parameter is the value given in the config file.
+ For example, if for the width on `centre` the width modifier reads "Absolute" with a value 20.0, this means if the
+ error on the parameter `centre` was less than 20.0 in the previous search, the sigma of its `TruncatedGaussianPrior` in
+ the next search will be 20.0.
+
+ 2) Relative: In this case, the error assumed on the parameter is the % of the value of the estimate value given in
+ the config file. For example, if the normalization estimated in the previous search was 2.0, and the relative error in
+ the config file reads "Relative" with a value 0.5, then the sigma of the `TruncatedGaussianPrior`
+ will be 50% of this value, i.e. sigma = 0.5 * 2.0 = 1.0.
+
+We use absolute and relative values for different parameters, depending on their properties. For example, using the
+relative value of a parameter like the `centre` makes no sense as the profile could be centred at 0.0, making
+the relative error tiny and poorly defined.
+
+However, there are parameters where using an absolute value does not make sense. Normalization is a good example of this.
+The normalization of an image depends on its units and S/N. There is no single absolute value that one can use to
+generically chain the normalization of any two proflies. Thus, it makes more sense to chain them using the relative value
+from a previous search.
+
+We can customize how priors are passed from the results of a search editing the `prior_passer` settings in
+the `general.yaml` config.
+
+__EXAMPLE__
+
+Lets go through an example using a real parameter. Lets say in search 1 we fit a `Gaussian` and we estimate that
+its normalization is equal to 4.0 +- 2.0 where the error value of 2.0 was computed at 3.0 sigma confidence. To pass this
+as a prior to search 2, we would write:
+
+ gaussian.normalization = search_1_result.model.gaussian.normalization
+
+The prior on the `Gaussian` `normalization` in search 2 would thus be a `TruncatedGaussianPrior`, with mean=4.0 and
+sigma=2.0. If we had used a sigma value of 1.0 to compute the error, which reduced the estimate from 4.0 +- 2.0 to
+4.0 +- 1.0, the sigma of the `TruncatedGaussianPrior` would instead be 1.0.
+
+If the error on the normalization in search 1 had been really small, lets say, 0.01, we would instead use the value of the
+normalization width in the priors config file to set sigma instead. In this case, the prior config file specifies
+that we use an "Relative" value of 0.5 to chain this prior. Thus, the TruncatedGaussianPrior in search 2 would have a mean=4.0
+and sigma=2.0.
+
+And with that, we`re done. Chaining searches is a bit of an art form, but for certain problems can be extremely
+powerful.
+"""
+
+"""
+Cookbook 5: Model Linking
+=========================
+
+__Prerequisites__
+
+You should be familiar with the search chaining API detailed in the following scripts and docs:
+
+__Overview__
+
+Search chaining allows one to perform back-to-back non-linear searches to fit a dataset, where the model complexity
+increases after each fit.
+
+To perform search chaining, **PyAutoFit** has tools for passing the results of one model-fit from one fit to the next,
+and change its parameterization between each fit.
+
+This cookbook is a concise reference to the model linking API.
+"""
+# from autoconf import setup_notebook; setup_notebook()
+
+import json
+import os
+from os import path
+
+import autofit as af
+
+"""
+__Model-Fit__
+
+We perform a quick model-fit, to create a `Result` object which has the attributes necessary to illustrate the model
+linking API.
+"""
+model = af.Collection(gaussian=af.ex.Gaussian, exponential=af.ex.Exponential)
+
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1__exponential_x1")
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+dynesty = af.DynestyStatic(name="cookbook_5_model_linking", nlive=50, sample="rwalk")
+
+result = dynesty.fit(model=model, analysis=analysis)
+
+"""
+__Instance & Model__
+
+The result object has two key attributes for model linking:
+
+ - `instance`: The maximum log likelihood instance of the model-fit, where every parameter is therefore a float.
+
+ - `model`: An attribute which represents how the result can be passed as a model-component to the next fit (the
+ details of how its priors are passed are given in full below).
+
+We create a new model using both of these attributes, where:
+
+ - All of the `gaussian` model components parameters are passed via the `instance` attribute and therefore fixed to
+ the inferred maximum log likelihood values (and are not free parameters in the model).
+
+ - All of the `exponential` model components parameters are passed via the `model` attribute and therefore are free
+ parameters in the model.
+
+The new model therefore has 3 free parameters and 3 fixed parameters.
+"""
+model = af.Collection(gaussian=af.ex.Gaussian, exponential=af.ex.Exponential)
+
+model.gaussian = result.instance.gaussian
+model.exponential = result.model.exponential
+
+"""
+The `model.info` attribute shows that the parameter and prior passing has occurred as described above.
+"""
+print(model.info)
+
+"""
+We can print the priors of the exponenital:
+"""
+print("Exponential Model Priors \n")
+print("centre = ", model.exponential.centre)
+print("normalization = ", model.exponential.normalization)
+print("rate = ", model.exponential.rate)
+
+"""
+How are the priors set via model linking? The full description is quite long, therefore it is attatched to the
+bottom of this script so that we can focus on the model linking API.
+
+__Component Specification__
+
+Model linking can be performed on any component of a model, for example to only pass specific parameters as
+an `instance` or `model`.
+"""
+gaussian = af.Model(af.ex.Gaussian)
+
+gaussian.centre = result.instance.gaussian.centre
+gaussian.normalization = result.model.gaussian.normalization
+gaussian.sigma = result.instance.gaussian.sigma
+
+exponential = af.Model(af.ex.Exponential)
+
+exponential.centre = result.model.exponential.centre
+exponential.normalization = result.model.exponential.normalization
+exponential.rate = result.instance.exponential.rate
+
+model = af.Collection(gaussian=gaussian, exponential=exponential)
+
+"""
+The `model.info` attribute shows that the parameter and prior passing has occurred on individual components.
+"""
+print(model.info)
+
+"""
+__Take Attributes__
+
+The examples above linked models where the individual model components that were passed stayed the same.
+
+We can link two related models, where only a subset of parameters are shared, by using the `take_attributes()` method.
+
+For example, lets define a `GaussianKurtosis` which is a `Gaussian` with an extra parameter for its kurtosis.
+"""
+
+
+class GaussianKurtosis:
+ def __init__(
+ self,
+ centre=30.0, # <- **PyAutoFit** recognises these constructor arguments
+ normalization=1.0, # <- are the Gaussian``s model parameters.
+ sigma=5.0,
+ kurtosis=1.0,
+ ):
+ self.centre = centre
+ self.normalization = normalization
+ self.sigma = sigma
+ self.kurtosis = kurtosis
+
+
+"""
+The `take_attributes()` method takes a `source` model component, and inspects the names of all its parameters.
+
+For the `Gaussian` model result input below, it finds the parameters `centre`, `normalization` and `sigma`.
+
+It then finds all parameters in the new `model` which have the same names, which for the `GaussianKurtosis` is
+`centre`, `normalization` and `sigma`.
+
+For all parameters which have the same name, the parameter is passed.
+"""
+model = af.Collection(gaussian=af.Model(GaussianKurtosis))
+model.kurtosis = af.UniformPrior(lower_limit=-1.0, upper_limit=1.0)
+
+model.gaussian.take_attributes(source=result.model.gaussian)
+
+"""
+Because the result was passed using `model` we see the priors on the `GaussianKurtosis` `centre`,
+`normalization` and `sigma` have been updated, whereas its `kurtosis` has not.
+"""
+print("GaussianKurtosis Model Priors After Take Attributes via Model \n")
+print("centre = ", model.gaussian.centre)
+print("normalization = ", model.gaussian.normalization)
+print("sigma = ", model.gaussian.sigma)
+print("kurtosis = ", model.gaussian.kurtosis)
+
+"""
+If we pass `result.instance` to take_attributes the same name linking is used, however parameters are passed as
+floats.
+"""
+model = af.Collection(gaussian=af.Model(GaussianKurtosis))
+model.kurtosis = af.UniformPrior(lower_limit=-1.0, upper_limit=1.0)
+
+model.gaussian.take_attributes(source=result.instance.gaussian)
+
+print("Gaussian Model Priors After Take Attributes via Instance \n")
+print("centre = ", model.gaussian.centre)
+print("normalization = ", model.gaussian.normalization)
+print("sigma = ", model.gaussian.sigma)
+print("kurtosis = ", model.gaussian.kurtosis)
+
+"""
+__As Model__
+
+A common problem is when we have an `instance` (e.g. from a previous fit where we fixed the parameters)
+but now wish to make its parameters free parameters again.
+
+Furthermore, we may want to do this for specific model components.
+
+The `as_model` method allows us to do this. Below, we pass the entire result (e.g. both the `gaussian`
+and `exponential` components), however we pass the `Gaussian` class to `as_model`, meaning that any model
+component in the `instance` which is a `Gaussian` will be converted to a model with free parameters.
+"""
+model = result.instance.as_model((af.ex.Gaussian,))
+
+print("Gaussian Model Priors After via as_model: \n")
+print("centre = ", model.gaussian.centre)
+print("normalization = ", model.gaussian.normalization)
+print("sigma = ", model.gaussian.sigma)
+print("centre = ", model.exponential.centre)
+print("normalization = ", model.exponential.normalization)
+print("rate= ", model.exponential.rate)
+
+"""
+The `as_model()` method does not have too much utility for the simple model used in this cookbook.
+
+However, for multi-level models with many components, it is a powerful tool to compose custom models.
+"""
+
+
+class MultiLevelProfiles:
+ def __init__(
+ self,
+ higher_level_centre=50.0, # This is the centre of all Gaussians in this multi level component.
+ profile_list=None, # This will contain a list of model-components
+ ):
+ self.higher_level_centre = higher_level_centre
+
+ self.profile_list = profile_list
+
+
+group_level_0 = af.Model(
+ MultiLevelProfiles, profile_list=[af.ex.Gaussian, af.ex.Exponential, af.ex.Gaussian]
+)
+
+group_level_1 = af.Model(
+ MultiLevelProfiles,
+ profile_list=[af.ex.Gaussian, af.ex.Exponential, af.ex.Exponential],
+)
+
+model = af.Collection(group_level_0=group_level_0, group_level_1=group_level_1)
+
+"""
+This means every `Gaussian` in the complex multi-level model above would have parameters set via the result of our
+model-fit, if the model above was fitted such that it was contained in the result.
+"""
+model = result.instance.as_model((af.ex.Gaussian,))
diff --git a/scripts/features/search_grid_search.py b/scripts/features/search_grid_search.py
index 7d52dd8b..d014a4b3 100644
--- a/scripts/features/search_grid_search.py
+++ b/scripts/features/search_grid_search.py
@@ -1,344 +1,340 @@
-"""
-Feature: Search Grid Search
-===========================
-
-A classic method to perform model-fitting is a grid search, where the parameters of a model are divided on to a grid of
-values and the likelihood of each set of parameters on this grid is sampled. For low dimensionality problems this
-simple approach can be sufficient to locate high likelihood solutions, however it scales poorly to higher dimensional
-problems.
-
-**PyAutoFit** can perform a search grid search, which allows one to perform a grid-search over a subset of parameters
-within a model, but use a non-linear search to fit for the other parameters. The parameters over which the grid-search
-is performed are also included in the model fit and their values are simply confined to the boundaries of their grid
-cell by setting these as the lower and upper limits of a `UniformPrior`.
-
-The benefits of using a search grid search are:
-
- - For problems with complex and multi-model parameters spaces it can be difficult to robustly and efficiently perform
- model-fitting. If specific parameters are known to drive the multi-modality then sampling over a grid can ensure the
- parameter space of each individual model-fit is not multi-modal and therefore sampled more accurately and efficiently.
-
- - It can provide a goodness-of-fit measure (e.g. the Bayesian evidence) of many model-fits over the grid. This can
- provide additional insight into where the model does and does not fit the data well, in a way that a standard
- non-linear search does not.
-
- - The search grid search is embarrassingly parallel, and if sufficient computing facilities are available one can
- perform model-fitting faster in real-time than a single non-linear search. The **PyAutoFit** search grid search
- includes an option for parallel model-fitting via the Python `multiprocessing` module.
-
-In this example we will demonstrate the search grid search feature, again using the example of fitting 1D Gaussian's
-in noisy data.
-
-__Example Source Code (`af.ex`)__
-
-The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
-
- - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
- `visualize` functions.
-
- - `Gaussian`: a model component representing a 1D Gaussian profile.
-
-These are functionally identical to the `Analysis` and `Gaussian` objects you have seen elsewhere in the workspace.
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Example Source Code (`af.ex`)**: The example objects used in this script.
-- **Data**: Load 1D Gaussian data with a small feature at pixel 70.
-- **Model**: Create a model with two Gaussians (main signal + feature).
-- **Analysis**: Create the Analysis class for fitting the model to data.
-- **Search**: Configure a non-linear search for a single fit.
-- **Result**: Plot and visualize the results from the single fit.
-- **Search Grid Search**: Set up and perform a grid search over a parameter subset.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import matplotlib.pyplot as plt
-import numpy as np
-from os import path
-
-import autofit as af
-
-"""
-__Data__
-
-Load data of a 1D Gaussian from a .json file in the directory
-`autofit_workspace/dataset/gaussian_x1_with_feature`.
-
-This 1D data includes a small feature to the right of the central `Gaussian`. This feature is a second `Gaussian`
-centred on pixel 70.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1_with_feature")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-"""
-Lets plot the data.
-
-The feature on pixel 70 is clearly visible.
-"""
-xvalues = range(data.shape[0])
-
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.title("1D Gaussian Data With Feature at pixel 70.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-__Model__
-
-Next, we create the model, which in this case corresponds to two `Gaussian`'s, one for the main signal seen in the
-data and one for the feature on pixel 70.
-"""
-model = af.Collection(gaussian_main=af.ex.Gaussian, gaussian_feature=af.ex.Gaussian)
-
-"""
-The `info` attribute shows the model in a readable format, showing it contains two `Gaussian`'s.
-"""
-print(model.info)
-
-"""
-__Analysis__
-
-Create the analysis which fits the model to the data.
-
-It fits the data as the sum of the two `Gaussian`'s in the model.
-"""
-analysis = af.ex.Analysis(data=data, noise_map=noise_map, use_jax=True)
-
-"""
-__Search__
-
-Fit the data using a single non-linear search, to demonstrate the behaviour of the fit before we invoke
-the search grid search.
-"""
-search = af.DynestyStatic(
- path_prefix=path.join("features", "search_grid_search"),
- name="single_fit",
- nlive=100,
- maxcall=30000,
-)
-
-search = af.Nautilus(
- path_prefix=path.join("features", "search_grid_search"),
- name="single_fit",
- n_live=100,
- # maxcall=30000,
-)
-
-"""
-To perform the fit with Dynesty, we pass it our model and analysis and we`re good to go!
-
-Checkout the folder `autofit_workspace/output/features.search_grid_search/single_fit`, where the `NonLinearSearch`
-results, visualization and information can be found.
-
-For test runs on my laptop it is 'hit or miss' whether the feature is fitted correctly. This is because although models
-including the feature corresponds to the highest likelihood solutions, they occupy a small volume in parameter space
-which the non linear search may miss. Furthemore, it is common for the model-fit to get stuck in local maxima where
-both `Gaussian`'s go to a centre value of 50.0.
-
-The fit can also take a very long time to run, therefore I limited `Dynesty` to 30000 iterations above.
-"""
-result = search.fit(model=model, analysis=analysis)
-
-"""
-__Result__
-
-If you ran the fit above, you can now plot the result.
-"""
-instance = result.max_log_likelihood_instance
-
-gaussian_main = instance.gaussian_main.model_data_from(xvalues=np.arange(data.shape[0]))
-gaussian_feature = instance.gaussian_feature.model_data_from(
- xvalues=np.arange(data.shape[0])
-)
-model_data = gaussian_main + gaussian_feature
-
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(range(data.shape[0]), model_data, color="r")
-plt.plot(range(data.shape[0]), gaussian_main, "--")
-plt.plot(range(data.shape[0]), gaussian_feature, "--")
-plt.title("Dynesty model fit to 1D Gaussian with feature dataset.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-__Search Grid Search__
-
-We will now perform the search grid search.
-
-We will use the same `Dynesty` settings, but change its `name`.
-"""
-# search = af.DynestyStatic(
-# name="grid_fit",
-# path_prefix=path.join("features", "search_grid_search"),
-# nlive=100,
-# maxcall=30000,
-# number_of_cores=2,
-# # force_x1_cpu=True, # ensures parallelizing over grid search works.
-# )
-
-search = af.Nautilus(
- name="grid_fit",
- path_prefix=path.join("features", "search_grid_search"),
- n_live=100,
- # maxcall=30000,
- # number_of_cores=2,
- # force_x1_cpu=True, # ensures parallelizing over grid search works.
-)
-
-"""
-To set up the search grid search we specify two additional settings:
-
-`number_of_steps`: The number of steps in the grid search that are performedm which is set to 5 below.
-
-Because the prior on the parameter `centre` is a `UniformPrior` from 0.0 -> 100.0, this means the first grid search
-will set the prior on the centre to be a `UniformPrior` from 0.0 -> 20.0. The second will run from 20.0 -> 40.0,
-the third 40.0 -> 60.0, and so on.
-
-`parallel`: If `True`, each grid search is performed in parallel on your laptop.
-
-`number_of_cores`: The number of cores the grid search will parallelize the run over. If `number_of_cores=1`, the
-search is run in serial. For > 1 core, 1 core is reserved as a farmer, e.g., if `number_of_cores=4` then up to 3
-searches will be run in parallel. In case your laptop has limited hardware resources we do not run in parallel in
-this example by default, but feel free to change the option to `True` if you have a lot of CPUs and memory!
-"""
-grid_search = af.SearchGridSearch(search=search, number_of_steps=5, number_of_cores=1)
-
-"""
-We can now run the grid search.
-
-This is where we specify the parameter over which the grid search is performed, in this case the `centre` of the
-`gaussian_feature` in our model.
-
-On my laptop, each model fit performed by the grid search takes ~15000 iterations, whereas the fit above
-required ~ 40000 iterations. Thus, in this simple example, the grid search did not speed up the overall analysis
-(unless it is run in parallel). However, more complex and realistic model-fitting problems, the grid search has the
-potential to give huge performance improvements if used effectively.
-"""
-grid_search_result = grid_search.fit(
- model=model, analysis=analysis, grid_priors=[model.gaussian_feature.centre]
-)
-
-"""
-This returns a `GridSearchResult`, which includes information on every model-fit performed on the grid.
-
-Below, we print:
-
- - The central value of the `UniformPrior` on the `centre` of the gaussian_feature` for each fit performed on the
- grid search.
-
- - The maximum log likelihood value of each of the 5 fits.
-
- - The Bayesian evidence of each (this is accessible because we used a nested sampling algorithm).
-
-You should see that the highest likelihood and evidence values correspond to run 4, where the `UniformPrior` on the
-centre parameter ran from 60 -> 80 and therefore captured the true value of 70.0.
-"""
-print(grid_search_result.physical_centres_lists)
-print(grid_search_result.log_likelihoods().native)
-print(grid_search_result.log_evidences().native)
-
-"""
-We can also access the `best_samples` and their maximum likelihood instance.
-"""
-print(grid_search_result.best_samples)
-
-instance = grid_search_result.best_samples.instance
-
-print(instance.gaussian_main.centre)
-print(instance.gaussian_main.normalization)
-print(instance.gaussian_main.sigma)
-print(instance.gaussian_feature.centre)
-print(instance.gaussian_feature.normalization)
-print(instance.gaussian_feature.sigma)
-
-"""
-By plotting the `best` instance we can confirm the grid search fitted the feature at pixel 70.
-"""
-gaussian_main = instance.gaussian_main.model_data_from(xvalues=np.arange(data.shape[0]))
-gaussian_feature = instance.gaussian_feature.model_data_from(
- xvalues=np.arange(data.shape[0])
-)
-model_data = gaussian_main + gaussian_feature
-
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(range(data.shape[0]), model_data, color="r")
-plt.plot(range(data.shape[0]), gaussian_main, "--")
-plt.plot(range(data.shape[0]), gaussian_feature, "--")
-plt.title("Dynesty model fit to 1D Gaussian with feature dataset.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-A multi-dimensional grid search can be easily performed by adding more parameters to the `grid_priors` input.
-
-The fit below belows performs a 5x5 grid search over the `centres` of both `Gaussians`. This would take quite a long
-time to run, so I've commented it out, but feel free to run it!
-"""
-# grid_search_result = grid_search.fit(
-# model=model,
-# analysis=analysis,
-# grid_priors=[model.gaussian_feature.centre, model.gaussian_main.centre],
-# )
-
-"""
-Finish.
-"""
+"""
+Feature: Search Grid Search
+===========================
+
+A classic method to perform model-fitting is a grid search, where the parameters of a model are divided on to a grid of
+values and the likelihood of each set of parameters on this grid is sampled. For low dimensionality problems this
+simple approach can be sufficient to locate high likelihood solutions, however it scales poorly to higher dimensional
+problems.
+
+**PyAutoFit** can perform a search grid search, which allows one to perform a grid-search over a subset of parameters
+within a model, but use a non-linear search to fit for the other parameters. The parameters over which the grid-search
+is performed are also included in the model fit and their values are simply confined to the boundaries of their grid
+cell by setting these as the lower and upper limits of a `UniformPrior`.
+
+The benefits of using a search grid search are:
+
+ - For problems with complex and multi-model parameters spaces it can be difficult to robustly and efficiently perform
+ model-fitting. If specific parameters are known to drive the multi-modality then sampling over a grid can ensure the
+ parameter space of each individual model-fit is not multi-modal and therefore sampled more accurately and efficiently.
+
+ - It can provide a goodness-of-fit measure (e.g. the Bayesian evidence) of many model-fits over the grid. This can
+ provide additional insight into where the model does and does not fit the data well, in a way that a standard
+ non-linear search does not.
+
+ - The search grid search is embarrassingly parallel, and if sufficient computing facilities are available one can
+ perform model-fitting faster in real-time than a single non-linear search. The **PyAutoFit** search grid search
+ includes an option for parallel model-fitting via the Python `multiprocessing` module.
+
+In this example we will demonstrate the search grid search feature, again using the example of fitting 1D Gaussian's
+in noisy data.
+
+__Example Source Code (`af.ex`)__
+
+The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
+
+ - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
+ `visualize` functions.
+
+ - `Gaussian`: a model component representing a 1D Gaussian profile.
+
+These are functionally identical to the `Analysis` and `Gaussian` objects you have seen elsewhere in the workspace.
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Example Source Code (`af.ex`)**: The example objects used in this script.
+- **Data**: Load 1D Gaussian data with a small feature at pixel 70.
+- **Model**: Create a model with two Gaussians (main signal + feature).
+- **Analysis**: Create the Analysis class for fitting the model to data.
+- **Search**: Configure a non-linear search for a single fit.
+- **Result**: Plot and visualize the results from the single fit.
+- **Search Grid Search**: Set up and perform a grid search over a parameter subset.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import matplotlib.pyplot as plt
+import numpy as np
+from os import path
+
+import autofit as af
+
+"""
+__Data__
+
+Load data of a 1D Gaussian from a .json file in the directory
+`autofit_workspace/dataset/gaussian_x1_with_feature`.
+
+This 1D data includes a small feature to the right of the central `Gaussian`. This feature is a second `Gaussian`
+centred on pixel 70.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1_with_feature")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+"""
+Lets plot the data.
+
+The feature on pixel 70 is clearly visible.
+"""
+xvalues = range(data.shape[0])
+
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.title("1D Gaussian Data With Feature at pixel 70.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+__Model__
+
+Next, we create the model, which in this case corresponds to two `Gaussian`'s, one for the main signal seen in the
+data and one for the feature on pixel 70.
+"""
+model = af.Collection(gaussian_main=af.ex.Gaussian, gaussian_feature=af.ex.Gaussian)
+
+"""
+The `info` attribute shows the model in a readable format, showing it contains two `Gaussian`'s.
+"""
+print(model.info)
+
+"""
+__Analysis__
+
+Create the analysis which fits the model to the data.
+
+It fits the data as the sum of the two `Gaussian`'s in the model.
+"""
+analysis = af.ex.Analysis(data=data, noise_map=noise_map, use_jax=True)
+
+"""
+__Search__
+
+Fit the data using a single non-linear search, to demonstrate the behaviour of the fit before we invoke
+the search grid search.
+"""
+search = af.DynestyStatic(
+ path_prefix=path.join("features", "search_grid_search"),
+ name="single_fit",
+ nlive=100,
+ maxcall=30000,
+)
+
+search = af.Nautilus(
+ path_prefix=path.join("features", "search_grid_search"),
+ name="single_fit",
+ n_live=100,
+ # maxcall=30000,
+)
+
+"""
+To perform the fit with Dynesty, we pass it our model and analysis and we`re good to go!
+
+Checkout the folder `autofit_workspace/output/features.search_grid_search/single_fit`, where the `NonLinearSearch`
+results, visualization and information can be found.
+
+For test runs on my laptop it is 'hit or miss' whether the feature is fitted correctly. This is because although models
+including the feature corresponds to the highest likelihood solutions, they occupy a small volume in parameter space
+which the non linear search may miss. Furthemore, it is common for the model-fit to get stuck in local maxima where
+both `Gaussian`'s go to a centre value of 50.0.
+
+The fit can also take a very long time to run, therefore I limited `Dynesty` to 30000 iterations above.
+"""
+result = search.fit(model=model, analysis=analysis)
+
+"""
+__Result__
+
+If you ran the fit above, you can now plot the result.
+"""
+instance = result.max_log_likelihood_instance
+
+gaussian_main = instance.gaussian_main.model_data_from(xvalues=np.arange(data.shape[0]))
+gaussian_feature = instance.gaussian_feature.model_data_from(
+ xvalues=np.arange(data.shape[0])
+)
+model_data = gaussian_main + gaussian_feature
+
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(range(data.shape[0]), model_data, color="r")
+plt.plot(range(data.shape[0]), gaussian_main, "--")
+plt.plot(range(data.shape[0]), gaussian_feature, "--")
+plt.title("Dynesty model fit to 1D Gaussian with feature dataset.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+__Search Grid Search__
+
+We will now perform the search grid search.
+
+We will use the same `Dynesty` settings, but change its `name`.
+"""
+# search = af.DynestyStatic(
+# name="grid_fit",
+# path_prefix=path.join("features", "search_grid_search"),
+# nlive=100,
+# maxcall=30000,
+# number_of_cores=2,
+# # force_x1_cpu=True, # ensures parallelizing over grid search works.
+# )
+
+search = af.Nautilus(
+ name="grid_fit",
+ path_prefix=path.join("features", "search_grid_search"),
+ n_live=100,
+ # maxcall=30000,
+ # number_of_cores=2,
+ # force_x1_cpu=True, # ensures parallelizing over grid search works.
+)
+
+"""
+To set up the search grid search we specify two additional settings:
+
+`number_of_steps`: The number of steps in the grid search that are performedm which is set to 5 below.
+
+Because the prior on the parameter `centre` is a `UniformPrior` from 0.0 -> 100.0, this means the first grid search
+will set the prior on the centre to be a `UniformPrior` from 0.0 -> 20.0. The second will run from 20.0 -> 40.0,
+the third 40.0 -> 60.0, and so on.
+
+`parallel`: If `True`, each grid search is performed in parallel on your laptop.
+
+`number_of_cores`: The number of cores the grid search will parallelize the run over. If `number_of_cores=1`, the
+search is run in serial. For > 1 core, 1 core is reserved as a farmer, e.g., if `number_of_cores=4` then up to 3
+searches will be run in parallel. In case your laptop has limited hardware resources we do not run in parallel in
+this example by default, but feel free to change the option to `True` if you have a lot of CPUs and memory!
+"""
+grid_search = af.SearchGridSearch(search=search, number_of_steps=5, number_of_cores=1)
+
+"""
+We can now run the grid search.
+
+This is where we specify the parameter over which the grid search is performed, in this case the `centre` of the
+`gaussian_feature` in our model.
+
+On my laptop, each model fit performed by the grid search takes ~15000 iterations, whereas the fit above
+required ~ 40000 iterations. Thus, in this simple example, the grid search did not speed up the overall analysis
+(unless it is run in parallel). However, more complex and realistic model-fitting problems, the grid search has the
+potential to give huge performance improvements if used effectively.
+"""
+grid_search_result = grid_search.fit(
+ model=model, analysis=analysis, grid_priors=[model.gaussian_feature.centre]
+)
+
+"""
+This returns a `GridSearchResult`, which includes information on every model-fit performed on the grid.
+
+Below, we print:
+
+ - The central value of the `UniformPrior` on the `centre` of the gaussian_feature` for each fit performed on the
+ grid search.
+
+ - The maximum log likelihood value of each of the 5 fits.
+
+ - The Bayesian evidence of each (this is accessible because we used a nested sampling algorithm).
+
+You should see that the highest likelihood and evidence values correspond to run 4, where the `UniformPrior` on the
+centre parameter ran from 60 -> 80 and therefore captured the true value of 70.0.
+"""
+print(grid_search_result.physical_centres_lists)
+print(grid_search_result.log_likelihoods().native)
+print(grid_search_result.log_evidences().native)
+
+"""
+We can also access the `best_samples` and their maximum likelihood instance.
+"""
+print(grid_search_result.best_samples)
+
+instance = grid_search_result.best_samples.instance
+
+print(instance.gaussian_main.centre)
+print(instance.gaussian_main.normalization)
+print(instance.gaussian_main.sigma)
+print(instance.gaussian_feature.centre)
+print(instance.gaussian_feature.normalization)
+print(instance.gaussian_feature.sigma)
+
+"""
+By plotting the `best` instance we can confirm the grid search fitted the feature at pixel 70.
+"""
+gaussian_main = instance.gaussian_main.model_data_from(xvalues=np.arange(data.shape[0]))
+gaussian_feature = instance.gaussian_feature.model_data_from(
+ xvalues=np.arange(data.shape[0])
+)
+model_data = gaussian_main + gaussian_feature
+
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(range(data.shape[0]), model_data, color="r")
+plt.plot(range(data.shape[0]), gaussian_main, "--")
+plt.plot(range(data.shape[0]), gaussian_feature, "--")
+plt.title("Dynesty model fit to 1D Gaussian with feature dataset.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+A multi-dimensional grid search can be easily performed by adding more parameters to the `grid_priors` input.
+
+The fit below belows performs a 5x5 grid search over the `centres` of both `Gaussians`. This would take quite a long
+time to run, so I've commented it out, but feel free to run it!
+"""
+# grid_search_result = grid_search.fit(
+# model=model,
+# analysis=analysis,
+# grid_priors=[model.gaussian_feature.centre, model.gaussian_main.centre],
+# )
+
+"""
+Finish.
+"""
diff --git a/scripts/features/sensitivity_mapping.py b/scripts/features/sensitivity_mapping.py
index b7b2aead..c15144cf 100644
--- a/scripts/features/sensitivity_mapping.py
+++ b/scripts/features/sensitivity_mapping.py
@@ -1,583 +1,579 @@
-"""
-Feature: Sensitivity Mapping
-============================
-
-Bayesian model comparison allows us to take a dataset, fit it with multiple models and use the Bayesian evidence to
-quantify which model objectively gives the best-fit following the principles of Occam's Razor.
-
-However, a complex model may not be favoured by model comparison not because it is the 'wrong' model, but simply
-because the dataset being fitted is not of a sufficient quality for the more complex model to be favoured. Sensitivity
-mapping addresses what quality of data would be needed for the more complex model to be favoured.
-
-In order to do this, sensitivity mapping involves us writing a function that uses the model(s) to simulate a dataset.
-We then use this function to simulate many datasets, for different models, and fit each dataset to quantify
-how much the change in the model led to a measurable change in the data. This is called computing the sensitivity.
-
-How we compute the sensitivity is chosen by us, the user. In this example, we will perform multiple model-fits
-with a nested sampling search, and therefore perform Bayesian model comparison to compute the sensitivity. This allows
-us to infer how much of a Bayesian evidence increase we should expect for datasets of varying quality and / or models
-with different parameters.
-
-__Example Source Code (`af.ex`)__
-
-The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
-
- - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
- `visualize` functions.
-
- - `Gaussian`: a model component representing a 1D Gaussian profile.
-
-These are functionally identical to the `Analysis` and `Gaussian` objects you have seen elsewhere in the workspace.
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Example Source Code (`af.ex`)**: The example objects used in this script.
-- **Data**: Load 1D Gaussian data with a small feature at pixel 70.
-- **Analysis**: Create the Analysis class for fitting the model to data.
-- **Model Comparison**: Perform Bayesian model comparison on the original data.
-- **Sensitivity Mapping**: Introduce the sensitivity mapping procedure.
-- **Base Model**: Define the simpler model used to simulate datasets.
-- **Perturb Model**: Define the model component used to perturb the base model during sensitivity mapping.
-- **Mapping Grid**: Specify the grid of model parameters for sensitivity mapping.
-- **Simulation Instance**: Provide the instance used for dataset simulation.
-- **Simulate Function Class**: Define the Dataset and Analysis classes for sensitivity mapping.
-- **Base Fit**: Define how the base model is fitted to simulated datasets.
-- **Perturb Fit**: Define how the perturbed model is fitted to simulated datasets.
-- **Results**: Interpret the sensitivity mapping results.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import matplotlib.pyplot as plt
-import numpy as np
-from os import path
-
-import autofit as af
-
-"""
-__Data__
-
-Load data of a 1D Gaussian from a .json file in the directory
-`autofit_workspace/dataset/gaussian_x1_with_feature`.
-
-This 1D data includes a small feature to the right of the central `Gaussian`. This feature is a second `Gaussian`
-centred on pixel 70.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1_with_feature")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-"""
-Lets plot the data.
-
-The feature on pixel 70 is clearly visible.
-"""
-xvalues = range(data.shape[0])
-
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.title("1D Gaussian Data With Feature at pixel 70.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-__Analysis__
-
-Create the analysis which fits the model to the data.
-
-It fits the data as the sum of the two `Gaussian`'s in the model.
-"""
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-"""
-__Model Comparison__
-
-Before performing sensitivity mapping, we will quickly perform Bayesian model comparison on this data to get a sense
-for whether the `Gaussian` feature is detectable and how much the Bayesian evidence increases when it is included in
-the model.
-
-We therefore fit the data using two models, one where the model is a single `Gaussian` and one where it is
-two `Gaussians`.
-
-To avoid slow model-fitting and more clearly prounce the results of model comparison, we restrict the centre of
-the`gaussian_feature` to its true centre of 70 and sigma value of 0.5.
-"""
-model = af.Collection(gaussian_main=af.ex.Gaussian)
-
-search = af.DynestyStatic(
- path_prefix=path.join("features", "sensitivity_mapping"),
- name="single_gaussian",
- unique_tag="hello",
- nlive=100,
-)
-
-result_single = search.fit(model=model, analysis=analysis)
-
-model = af.Collection(gaussian_main=af.ex.Gaussian, gaussian_feature=af.ex.Gaussian)
-model.gaussian_feature.centre = 70.0
-model.gaussian_feature.sigma = 0.5
-
-search = af.DynestyStatic(
- path_prefix=path.join("features", "sensitivity_mapping", "two_gaussians"), nlive=100
-)
-
-result_multiple = search.fit(model=model, analysis=analysis)
-
-"""
-We can now print the `log_evidence` of each fit and confirm the model with two `Gaussians` was preferred to the model
-with just one `Gaussian`.
-"""
-print(result_single.samples.log_evidence)
-print(result_multiple.samples.log_evidence)
-
-"""
-__Sensitivity Mapping__
-
-The model comparison above shows that in this dataset, the `Gaussian` feature was detectable and that it increased the
-Bayesian evidence by ~25. Furthermore, the normalization of this `Gaussian` was ~0.3.
-
-A lower value of normalization makes the `Gaussian` fainter and harder to detect. We will demonstrate sensitivity
-mapping by answering the following question, at what value of normalization does the `Gaussian` feature become
-undetectable and not provide us with a noticeable increase in Bayesian evidence?
-
-__Base Model__
-
-To begin, we define the `base_model` that we use to perform sensitivity mapping. This model is used to simulate every
-dataset. It is also fitted to every simulated dataset without the extra model component below, to give us the Bayesian
-evidence of the every simpler model to compare to the more complex model.
-
-The `base_model` corresponds to the `gaussian_main` above.
-"""
-base_model = af.Collection(gaussian_main=af.ex.Gaussian)
-
-"""
-__Perturb Model__
-
-We now define the `perturb_model`, which is the model component whose parameters we iterate over to perform
-sensitivity mapping. Many instances of the `perturb_model` are created and used to simulate the many datasets
-that we fit. However, it is only included in half of the sensitivity mapping models, corresponding to the more complex
-models whose Bayesian evidence we compare to the simpler model-fits consisting of just the `base_model`.
-
-The `perturb_model` is therefore another `Gaussian` but now corresponds to the `gaussian_feature` above.
-
-By fitting both of these models to every simulated dataset, we will therefore infer the Bayesian evidence of every
-model to every dataset. Sensitivity mapping therefore maps out for what values of `normalization` in
-the `gaussian_feature` does the more complex model-fit provide higher values of Bayesian evidence than the simpler
-model-fit.
-"""
-perturb_model = af.Model(af.ex.Gaussian)
-
-"""
-__Mapping Grid__
-
-Sensitivity mapping is performed over a large grid of model parameters. To make this demonstration quick and clear we
-are going to fix the `centre` and `sigma` values to the true values of the `gaussian_feature`. We will also iterate
-over just two `normalization` values corresponding to 0.01 and 100.0, which will exhaggerate the difference in
-sensitivity between the models at these two values.
-"""
-perturb_model.centre = 70.0
-perturb_model.sigma = 0.5
-perturb_model.normalization = af.UniformPrior(lower_limit=0.01, upper_limit=100.0)
-
-"""
-__Simulation Instance__
-
-We are performing sensitivity mapping to determine how bright the `gaussian_feature` needs to be in order to be
-detectable. However, every simulated dataset must include the `main_gaussian`, as its presence in the data will effect
-the detectability of the `gaussian_feature`.
-
-We can pass the `main_gaussian` into the sensitivity mapping as the `simulation_instance`, meaning that it will be used
-in the simulation of every dataset. For this example we use the inferred `main_gaussian` from one of the model-fits
-performed above.
-"""
-simulation_instance = result_single.instance
-
-"""
-__Simulate Function Class__
-
-We are about to write a `simulate_cls` that simulates examples of 1D `Gaussian` datasets that are fitted to
-perform sensitivity mapping.
-
-To pass each simulated data through **PyAutoFit**'s sensitivity mapping tools, the function must return a single
-Python object. We therefore define a `Dataset` class that combines the `data` and `noise_map` that are to be
-output by this `simulate_cls`.
-
-It is also convenient to define a `Analysis` class, which behaves analogously to the `Analysis` class used in
-PyAutoFit to fit a model to data. In this example it makes it easy to define how we fit each simulated dataset.
-"""
-
-
-class Dataset:
- def __init__(self, data, noise_map):
- self.data = data
- self.noise_map = noise_map
-
-
-class Analysis(af.ex.Analysis):
- def __init__(self, dataset):
- super().__init__(data=dataset.data, noise_map=dataset.noise_map)
-
-
-"""
-We now write the `simulate_cls`, which takes the `simulation_instance` of our model (defined above) and uses it to
-simulate a dataset which is subsequently fitted.
-
-Additional attributes required to simulate the data can be passed to the `__init__` method, and the simulation is
-performed in the `__call__` method.
-
-Note that when this dataset is simulated, the quantity `instance.perturb` is used in `__call__`.
-This is an instance of the `gaussian_feature`, and it is different every time the `simulate_cls` is called.
-
-In this example, this `instance.perturb` corresponds to two different `gaussian_feature` with values of
-`normalization` of 0.01 and 100.0, such that our simulated datasets correspond to a very faint and very bright gaussian
-features .
-"""
-
-
-class Simulate:
- def __init__(self):
- """
- Class used to simulate every dataset used for sensitivity mapping.
-
- This `__init__` constructor can be extended with new inputs which can be used to control how the dataset is
- simulated in the `__call__` simulate_function below.
-
- In this example we leave it empty as our `simulate_function` does not require any additional information.
- """
- pass
-
- def __call__(self, instance, simulate_path):
- """
- The `simulate_function` called by the `Sensitivity` class which simulates each dataset fitted
- by the sensitivity mapper.
-
- The simulation procedure is as follows:
-
- 1) Use the input sensitivity `instance` to simulate the data with the small Gaussian feature.
-
- 2) Output information about the simulation to hard-disk.
-
- 3) Return the data for the sensitivity mapper to fit.
-
- Parameters
- ----------
- instance
- The sensitivity instance, which includes the Gaussian feature parameters are varied to perform sensitivity.
- The Gaussian feature in this instance changes for every iteration of the sensitivity mapping.
- simulate_path
- The path where the simulated dataset is output, contained within each sub-folder of the sensitivity
- mapping.
-
- Returns
- -------
- A simulated image of a Gaussian, which i input into the fits of the sensitivity mapper.
- """
-
- """
- Specify the number of pixels used to create the xvalues on which the 1D line of the profile is generated
- using and thus defining the number of data-points in our data.
- """
- pixels = 100
- xvalues = np.arange(pixels)
-
- """
- Evaluate the `Gaussian` and Exponential model instances at every xvalues to create their model profile
- and sum them together to create the overall model profile.
-
- This print statement will show that, when you run `Sensitivity` below the values of the perturbation
- use fixed values of `centre=70` and `sigma=0.5`, whereas the normalization varies over the `number_of_steps`
- based on its prior.
- """
-
- print(instance.perturb.centre)
- print(instance.perturb.normalization)
- print(instance.perturb.sigma)
-
- model_line = instance.gaussian_main.model_data_from(
- xvalues=xvalues
- ) + instance.perturb.model_data_from(xvalues=xvalues)
-
- """
- Determine the noise (at a specified signal to noise level) in every pixel of our model profile.
- """
- signal_to_noise_ratio = 25.0
- noise = np.random.normal(0.0, 1.0 / signal_to_noise_ratio, pixels)
-
- """
- Add this noise to the model line to create the line data that is fitted, using the signal-to-noise ratio
- to compute noise-map of our data which is required when evaluating the chi-squared value of the likelihood.
- """
- data = model_line + noise
- noise_map = (1.0 / signal_to_noise_ratio) * np.ones(pixels)
-
- return Dataset(data=data, noise_map=noise_map)
-
-
-"""
-__Base Fit__
-
-We have defined a `Simulate` class that will be used to simulate every dataset simulated by the sensitivity mapper.
-Each simulated dataset will have a unique set of parameters for the `gaussian_feature` (e.g. due to different values of
-`perturb_model`.
-
-We will fit each simulated dataset using the `base_model`, which quantifies whether not including the Gaussian feature
-in the model changes the goodness-of-fit and therefore indicates if we are sensitive to the Gaussian feature.
-
-We now write a `BaseFit` class, defining how the `base_model` is fitted to each simulated dataset and
-the goodness-of-fit used to quantify whether the model fits the data well. As above, the `__init__` method can be
-extended with new inputs to control how the model is fitted and the `__call__` method performs the fit.
-
-In this example, we use a full non-linear search to fit the `base_model` to the simulated data and return
-the `log_evidence` of the model fit as the goodness-of-fit. This fit could easily be something much simpler and
-more computationally efficient, for example performing a single log likelihood evaluation of the `base_model` fit
-to the simulated data.
-"""
-
-
-class BaseFit:
- def __init__(self, analysis_cls):
- """
- Class used to fit every dataset used for sensitivity mapping with the base model (the model without the
- perturbed feature sensitivity mapping maps out).
-
- In this example, the base model therefore does not include the extra Gaussian feature, but the simulated
- dataset includes one.
-
- The base fit is repeated for every parameter on the sensitivity grid and compared to the perturbed fit. This
- maps out the sensitivity of every parameter is (e.g. the sensitivity of the normalization of the Gaussian
- feature).
-
- The `__init__` constructor can be extended with new inputs which can be used to control how the dataset is
- fitted, below we include an input `analysis_cls` which is the `Analysis` class used to fit the model to the
- dataset.
-
- Parameters
- ----------
- analysis_cls
- The `Analysis` class used to fit the model to the dataset.
- """
- self.analysis_cls = analysis_cls
-
- def __call__(self, dataset, model, paths, instance):
- """
- The base fitting function which fits every dataset used for sensitivity mapping with the base model.
-
- This function receives as input each simulated dataset of the sensitivity map and fits it, in order to
- quantify how sensitive the model is to the perturbed feature.
-
- In this example, a full non-linear search is performed to determine how well the model fits the dataset.
- The `log_evidence` of the fit is returned which acts as the sensitivity map figure of merit.
-
- Parameters
- ----------
- dataset
- The dataset which is simulated with the perturbed model and which is fitted.
- model
- The model instance which is fitted to the dataset, which does not include the perturbed feature.
- paths
- The `Paths` instance which contains the path to the folder where the results of the fit are written to.
- instance
- The simulation instance, which includes the perturbed feature that is used to simulate the dataset.
- This is often not used, but may be useful for certain sensitivity mapping tasks, for example using
- true values of the simulated instance to set up aspects of the model-fit (e.g. the priors).
- """
-
- search = af.DynestyStatic(
- paths=paths.for_sub_analysis(analysis_name="[base]"),
- nlive=50,
- iterations_per_full_update=50000,
- )
-
- analysis = self.analysis_cls(dataset=dataset)
-
- return search.fit(model=model, analysis=analysis)
-
-
-"""
-__Perturb Fit__
-
-We now define a `PerturbFit` class, which defines how the `perturb_model` is fitted to each simulated dataset. This
-behaves analogously to the `BaseFit` class above, but now fits the `perturb_model` to the simulated data (as
-opposed to the `base_model`).
-
-Again, in this example we use a full non-linear search to fit the `perturb_model` to the simulated data and return
-the `log_evidence` of the model fit as the goodness-of-fit. This fit could easily be something much simpler and
-more computationally efficient, for example performing a single log likelihood evaluation of the `perturb_model` fit
-to the simulated data.
-"""
-
-
-class PerturbFit:
- def __init__(self, analysis_cls):
- """
- Class used to fit every dataset used for sensitivity mapping with the perturbed model (the model with the
- perturbed feature sensitivity mapping maps out).
-
- In this example, the perturbed model therefore includes the extra Gaussian feature, which is also in the
- simulated dataset.
-
- The perturbed fit is repeated for every parameter on the sensitivity grid and compared to the base fit. This
- maps out the sensitivity of every parameter is (e.g. the sensitivity of the normalization of the Gaussian
- feature).
-
- The `__init__` constructor can be extended with new inputs which can be used to control how the dataset is
- fitted, below we include an input `analysis_cls` which is the `Analysis` class used to fit the model to the
- dataset.
-
- Parameters
- ----------
- analysis_cls
- The `Analysis` class used to fit the model to the dataset.
- """
- self.analysis_cls = analysis_cls
-
- def __call__(self, dataset, model, paths, instance):
- """
- The perturbed fitting function which fits every dataset used for sensitivity mapping with the perturbed model.
-
- This function receives as input each simulated dataset of the sensitivity map and fits it, in order to
- quantify how sensitive the model is to the perturbed feature.
-
- In this example, a full non-linear search is performed to determine how well the model fits the dataset.
- The `log_evidence` of the fit is returned which acts as the sensitivity map figure of merit.
-
- Parameters
- ----------
- dataset
- The dataset which is simulated with the perturbed model and which is fitted.
- model
- The model instance which is fitted to the dataset, which includes the perturbed feature.
- paths
- The `Paths` instance which contains the path to the folder where the results of the fit are written to.
- instance
- The simulation instance, which includes the perturbed feature that is used to simulate the dataset.
- This is often not used, but may be useful for certain sensitivity mapping tasks, for example using
- true values of the simulated instance to set up aspects of the model-fit (e.g. the priors).
- """
-
- search = af.DynestyStatic(
- paths=paths.for_sub_analysis(analysis_name="[perturbed]"),
- nlive=50,
- iterations_per_full_update=50000,
- )
-
- analysis = self.analysis_cls(dataset=dataset)
-
- return search.fit(model=model, analysis=analysis)
-
-
-"""
-We can now combine all of the objects created above and perform sensitivity mapping. The inputs to the `Sensitivity`
-object below are:
-
-- `simulation_instance`: This is an instance of the model used to simulate every dataset that is fitted. In this
-example it contains an instance of the `gaussian_main` model component.
-
-- `base_model`: This is the simpler model that is fitted to every simulated dataset, which in this example is composed
-of a single `Gaussian` called the `gaussian_main`.
-
-- `perturb_model`: This is the extra model component that has two roles: (i) based on the sensitivity grid parameters
-it is added to the `simulation_instance` to simulate each dataset ; (ii) it is added to the`base_model` and fitted to
-every simulated dataset (in this example every `simulation_instance` and `perturb_model` there has two `Gaussians`
-called the `gaussian_main` and `gaussian_feature`).
-
-- `simulate_cls`: This is the function that uses the `simulation_instance` and many instances of the `perturb_model`
-to simulate many datasets which are fitted with the `base_model` and `base_model` + `perturb_model`.
-
-- `base_fit_cls`: This is the function that fits the `base_model` to every simulated dataset and returns the
-goodness-of-fit of the model to the data.
-
-- `perturb_fit_cls`: This is the function that fits the `base_model` + `perturb_model` to every simulated dataset and
-returns the goodness-of-fit of the model to the data.
-
-- `number_of_steps`: The number of steps over which the parameters in the `perturb_model` are iterated. In this
-example, normalization has a `LogUniformPrior` with lower limit 1e-4 and upper limit 1e2, therefore the `number_of_steps`
-of 2 wills imulate and fit just 2 datasets where the intensities between 1e-4 and 1e2.
-
-- `number_of_cores`: The number of cores over which the sensitivity mapping is performed, enabling parallel processing
-if set above 1.
-"""
-paths = af.DirectoryPaths(
- path_prefix=path.join("features"),
- unique_tag="hello",
- name="sensitivity_mapping",
-)
-
-sensitivity = af.Sensitivity(
- paths=paths,
- simulation_instance=simulation_instance,
- base_model=base_model,
- perturb_model=perturb_model,
- simulate_cls=Simulate(),
- base_fit_cls=BaseFit(analysis_cls=Analysis),
- perturb_fit_cls=PerturbFit(analysis_cls=Analysis),
- number_of_steps=2,
- number_of_cores=2,
-)
-sensitivity_result = sensitivity.run()
-
-"""
-__Results__
-
-You should now look at the results of the sensitivity mapping in the folder `output/features/sensitivity_mapping`.
-
-You will note the following 4 model-fits have been performed:
-
- - The `base_model` is fitted to a simulated dataset where the `simulation_instance` and
- a `perturb` with `normalization=0.01` are used.
-
- - The `base_model` + `perturb_model` is fitted to a simulated dataset where the `simulation_instance` and
- a `perturb` with `normalization=0.01` are used.
-
- - The `base_model` is fitted to a simulated dataset where the `simulation_instance` and
- a `perturb` with `normalization=100.0` are used.
-
- - The `base_model` + `perturb_model` is fitted to a simulated dataset where the `simulation_instance` and
- a `perturb` with `normalization=100.0` are used.
-
-The fit produced a `sensitivity_result`.
-
-We are still developing the `SensitivityResult` class to provide a data structure that better streamlines the analysis
-of results. If you intend to use sensitivity mapping, the best way to interpret the resutls is currently via
-**PyAutoFit**'s database and `Aggregator` tools.
-"""
-print(sensitivity_result.samples)
-print(sensitivity_result.log_evidences_base)
-
-"""
-Finish.
-"""
+"""
+Feature: Sensitivity Mapping
+============================
+
+Bayesian model comparison allows us to take a dataset, fit it with multiple models and use the Bayesian evidence to
+quantify which model objectively gives the best-fit following the principles of Occam's Razor.
+
+However, a complex model may not be favoured by model comparison not because it is the 'wrong' model, but simply
+because the dataset being fitted is not of a sufficient quality for the more complex model to be favoured. Sensitivity
+mapping addresses what quality of data would be needed for the more complex model to be favoured.
+
+In order to do this, sensitivity mapping involves us writing a function that uses the model(s) to simulate a dataset.
+We then use this function to simulate many datasets, for different models, and fit each dataset to quantify
+how much the change in the model led to a measurable change in the data. This is called computing the sensitivity.
+
+How we compute the sensitivity is chosen by us, the user. In this example, we will perform multiple model-fits
+with a nested sampling search, and therefore perform Bayesian model comparison to compute the sensitivity. This allows
+us to infer how much of a Bayesian evidence increase we should expect for datasets of varying quality and / or models
+with different parameters.
+
+__Example Source Code (`af.ex`)__
+
+The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
+
+ - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
+ `visualize` functions.
+
+ - `Gaussian`: a model component representing a 1D Gaussian profile.
+
+These are functionally identical to the `Analysis` and `Gaussian` objects you have seen elsewhere in the workspace.
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Example Source Code (`af.ex`)**: The example objects used in this script.
+- **Data**: Load 1D Gaussian data with a small feature at pixel 70.
+- **Analysis**: Create the Analysis class for fitting the model to data.
+- **Model Comparison**: Perform Bayesian model comparison on the original data.
+- **Sensitivity Mapping**: Introduce the sensitivity mapping procedure.
+- **Base Model**: Define the simpler model used to simulate datasets.
+- **Perturb Model**: Define the model component used to perturb the base model during sensitivity mapping.
+- **Mapping Grid**: Specify the grid of model parameters for sensitivity mapping.
+- **Simulation Instance**: Provide the instance used for dataset simulation.
+- **Simulate Function Class**: Define the Dataset and Analysis classes for sensitivity mapping.
+- **Base Fit**: Define how the base model is fitted to simulated datasets.
+- **Perturb Fit**: Define how the perturbed model is fitted to simulated datasets.
+- **Results**: Interpret the sensitivity mapping results.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import matplotlib.pyplot as plt
+import numpy as np
+from os import path
+
+import autofit as af
+
+"""
+__Data__
+
+Load data of a 1D Gaussian from a .json file in the directory
+`autofit_workspace/dataset/gaussian_x1_with_feature`.
+
+This 1D data includes a small feature to the right of the central `Gaussian`. This feature is a second `Gaussian`
+centred on pixel 70.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1_with_feature")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+"""
+Lets plot the data.
+
+The feature on pixel 70 is clearly visible.
+"""
+xvalues = range(data.shape[0])
+
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.title("1D Gaussian Data With Feature at pixel 70.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+__Analysis__
+
+Create the analysis which fits the model to the data.
+
+It fits the data as the sum of the two `Gaussian`'s in the model.
+"""
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+"""
+__Model Comparison__
+
+Before performing sensitivity mapping, we will quickly perform Bayesian model comparison on this data to get a sense
+for whether the `Gaussian` feature is detectable and how much the Bayesian evidence increases when it is included in
+the model.
+
+We therefore fit the data using two models, one where the model is a single `Gaussian` and one where it is
+two `Gaussians`.
+
+To avoid slow model-fitting and more clearly prounce the results of model comparison, we restrict the centre of
+the`gaussian_feature` to its true centre of 70 and sigma value of 0.5.
+"""
+model = af.Collection(gaussian_main=af.ex.Gaussian)
+
+search = af.DynestyStatic(
+ path_prefix=path.join("features", "sensitivity_mapping"),
+ name="single_gaussian",
+ unique_tag="hello",
+ nlive=100,
+)
+
+result_single = search.fit(model=model, analysis=analysis)
+
+model = af.Collection(gaussian_main=af.ex.Gaussian, gaussian_feature=af.ex.Gaussian)
+model.gaussian_feature.centre = 70.0
+model.gaussian_feature.sigma = 0.5
+
+search = af.DynestyStatic(
+ path_prefix=path.join("features", "sensitivity_mapping", "two_gaussians"), nlive=100
+)
+
+result_multiple = search.fit(model=model, analysis=analysis)
+
+"""
+We can now print the `log_evidence` of each fit and confirm the model with two `Gaussians` was preferred to the model
+with just one `Gaussian`.
+"""
+print(result_single.samples.log_evidence)
+print(result_multiple.samples.log_evidence)
+
+"""
+__Sensitivity Mapping__
+
+The model comparison above shows that in this dataset, the `Gaussian` feature was detectable and that it increased the
+Bayesian evidence by ~25. Furthermore, the normalization of this `Gaussian` was ~0.3.
+
+A lower value of normalization makes the `Gaussian` fainter and harder to detect. We will demonstrate sensitivity
+mapping by answering the following question, at what value of normalization does the `Gaussian` feature become
+undetectable and not provide us with a noticeable increase in Bayesian evidence?
+
+__Base Model__
+
+To begin, we define the `base_model` that we use to perform sensitivity mapping. This model is used to simulate every
+dataset. It is also fitted to every simulated dataset without the extra model component below, to give us the Bayesian
+evidence of the every simpler model to compare to the more complex model.
+
+The `base_model` corresponds to the `gaussian_main` above.
+"""
+base_model = af.Collection(gaussian_main=af.ex.Gaussian)
+
+"""
+__Perturb Model__
+
+We now define the `perturb_model`, which is the model component whose parameters we iterate over to perform
+sensitivity mapping. Many instances of the `perturb_model` are created and used to simulate the many datasets
+that we fit. However, it is only included in half of the sensitivity mapping models, corresponding to the more complex
+models whose Bayesian evidence we compare to the simpler model-fits consisting of just the `base_model`.
+
+The `perturb_model` is therefore another `Gaussian` but now corresponds to the `gaussian_feature` above.
+
+By fitting both of these models to every simulated dataset, we will therefore infer the Bayesian evidence of every
+model to every dataset. Sensitivity mapping therefore maps out for what values of `normalization` in
+the `gaussian_feature` does the more complex model-fit provide higher values of Bayesian evidence than the simpler
+model-fit.
+"""
+perturb_model = af.Model(af.ex.Gaussian)
+
+"""
+__Mapping Grid__
+
+Sensitivity mapping is performed over a large grid of model parameters. To make this demonstration quick and clear we
+are going to fix the `centre` and `sigma` values to the true values of the `gaussian_feature`. We will also iterate
+over just two `normalization` values corresponding to 0.01 and 100.0, which will exhaggerate the difference in
+sensitivity between the models at these two values.
+"""
+perturb_model.centre = 70.0
+perturb_model.sigma = 0.5
+perturb_model.normalization = af.UniformPrior(lower_limit=0.01, upper_limit=100.0)
+
+"""
+__Simulation Instance__
+
+We are performing sensitivity mapping to determine how bright the `gaussian_feature` needs to be in order to be
+detectable. However, every simulated dataset must include the `main_gaussian`, as its presence in the data will effect
+the detectability of the `gaussian_feature`.
+
+We can pass the `main_gaussian` into the sensitivity mapping as the `simulation_instance`, meaning that it will be used
+in the simulation of every dataset. For this example we use the inferred `main_gaussian` from one of the model-fits
+performed above.
+"""
+simulation_instance = result_single.instance
+
+"""
+__Simulate Function Class__
+
+We are about to write a `simulate_cls` that simulates examples of 1D `Gaussian` datasets that are fitted to
+perform sensitivity mapping.
+
+To pass each simulated data through **PyAutoFit**'s sensitivity mapping tools, the function must return a single
+Python object. We therefore define a `Dataset` class that combines the `data` and `noise_map` that are to be
+output by this `simulate_cls`.
+
+It is also convenient to define a `Analysis` class, which behaves analogously to the `Analysis` class used in
+PyAutoFit to fit a model to data. In this example it makes it easy to define how we fit each simulated dataset.
+"""
+
+
+class Dataset:
+ def __init__(self, data, noise_map):
+ self.data = data
+ self.noise_map = noise_map
+
+
+class Analysis(af.ex.Analysis):
+ def __init__(self, dataset):
+ super().__init__(data=dataset.data, noise_map=dataset.noise_map)
+
+
+"""
+We now write the `simulate_cls`, which takes the `simulation_instance` of our model (defined above) and uses it to
+simulate a dataset which is subsequently fitted.
+
+Additional attributes required to simulate the data can be passed to the `__init__` method, and the simulation is
+performed in the `__call__` method.
+
+Note that when this dataset is simulated, the quantity `instance.perturb` is used in `__call__`.
+This is an instance of the `gaussian_feature`, and it is different every time the `simulate_cls` is called.
+
+In this example, this `instance.perturb` corresponds to two different `gaussian_feature` with values of
+`normalization` of 0.01 and 100.0, such that our simulated datasets correspond to a very faint and very bright gaussian
+features .
+"""
+
+
+class Simulate:
+ def __init__(self):
+ """
+ Class used to simulate every dataset used for sensitivity mapping.
+
+ This `__init__` constructor can be extended with new inputs which can be used to control how the dataset is
+ simulated in the `__call__` simulate_function below.
+
+ In this example we leave it empty as our `simulate_function` does not require any additional information.
+ """
+ pass
+
+ def __call__(self, instance, simulate_path):
+ """
+ The `simulate_function` called by the `Sensitivity` class which simulates each dataset fitted
+ by the sensitivity mapper.
+
+ The simulation procedure is as follows:
+
+ 1) Use the input sensitivity `instance` to simulate the data with the small Gaussian feature.
+
+ 2) Output information about the simulation to hard-disk.
+
+ 3) Return the data for the sensitivity mapper to fit.
+
+ Parameters
+ ----------
+ instance
+ The sensitivity instance, which includes the Gaussian feature parameters are varied to perform sensitivity.
+ The Gaussian feature in this instance changes for every iteration of the sensitivity mapping.
+ simulate_path
+ The path where the simulated dataset is output, contained within each sub-folder of the sensitivity
+ mapping.
+
+ Returns
+ -------
+ A simulated image of a Gaussian, which i input into the fits of the sensitivity mapper.
+ """
+
+ """
+ Specify the number of pixels used to create the xvalues on which the 1D line of the profile is generated
+ using and thus defining the number of data-points in our data.
+ """
+ pixels = 100
+ xvalues = np.arange(pixels)
+
+ """
+ Evaluate the `Gaussian` and Exponential model instances at every xvalues to create their model profile
+ and sum them together to create the overall model profile.
+
+ This print statement will show that, when you run `Sensitivity` below the values of the perturbation
+ use fixed values of `centre=70` and `sigma=0.5`, whereas the normalization varies over the `number_of_steps`
+ based on its prior.
+ """
+
+ print(instance.perturb.centre)
+ print(instance.perturb.normalization)
+ print(instance.perturb.sigma)
+
+ model_line = instance.gaussian_main.model_data_from(
+ xvalues=xvalues
+ ) + instance.perturb.model_data_from(xvalues=xvalues)
+
+ """
+ Determine the noise (at a specified signal to noise level) in every pixel of our model profile.
+ """
+ signal_to_noise_ratio = 25.0
+ noise = np.random.normal(0.0, 1.0 / signal_to_noise_ratio, pixels)
+
+ """
+ Add this noise to the model line to create the line data that is fitted, using the signal-to-noise ratio
+ to compute noise-map of our data which is required when evaluating the chi-squared value of the likelihood.
+ """
+ data = model_line + noise
+ noise_map = (1.0 / signal_to_noise_ratio) * np.ones(pixels)
+
+ return Dataset(data=data, noise_map=noise_map)
+
+
+"""
+__Base Fit__
+
+We have defined a `Simulate` class that will be used to simulate every dataset simulated by the sensitivity mapper.
+Each simulated dataset will have a unique set of parameters for the `gaussian_feature` (e.g. due to different values of
+`perturb_model`.
+
+We will fit each simulated dataset using the `base_model`, which quantifies whether not including the Gaussian feature
+in the model changes the goodness-of-fit and therefore indicates if we are sensitive to the Gaussian feature.
+
+We now write a `BaseFit` class, defining how the `base_model` is fitted to each simulated dataset and
+the goodness-of-fit used to quantify whether the model fits the data well. As above, the `__init__` method can be
+extended with new inputs to control how the model is fitted and the `__call__` method performs the fit.
+
+In this example, we use a full non-linear search to fit the `base_model` to the simulated data and return
+the `log_evidence` of the model fit as the goodness-of-fit. This fit could easily be something much simpler and
+more computationally efficient, for example performing a single log likelihood evaluation of the `base_model` fit
+to the simulated data.
+"""
+
+
+class BaseFit:
+ def __init__(self, analysis_cls):
+ """
+ Class used to fit every dataset used for sensitivity mapping with the base model (the model without the
+ perturbed feature sensitivity mapping maps out).
+
+ In this example, the base model therefore does not include the extra Gaussian feature, but the simulated
+ dataset includes one.
+
+ The base fit is repeated for every parameter on the sensitivity grid and compared to the perturbed fit. This
+ maps out the sensitivity of every parameter is (e.g. the sensitivity of the normalization of the Gaussian
+ feature).
+
+ The `__init__` constructor can be extended with new inputs which can be used to control how the dataset is
+ fitted, below we include an input `analysis_cls` which is the `Analysis` class used to fit the model to the
+ dataset.
+
+ Parameters
+ ----------
+ analysis_cls
+ The `Analysis` class used to fit the model to the dataset.
+ """
+ self.analysis_cls = analysis_cls
+
+ def __call__(self, dataset, model, paths, instance):
+ """
+ The base fitting function which fits every dataset used for sensitivity mapping with the base model.
+
+ This function receives as input each simulated dataset of the sensitivity map and fits it, in order to
+ quantify how sensitive the model is to the perturbed feature.
+
+ In this example, a full non-linear search is performed to determine how well the model fits the dataset.
+ The `log_evidence` of the fit is returned which acts as the sensitivity map figure of merit.
+
+ Parameters
+ ----------
+ dataset
+ The dataset which is simulated with the perturbed model and which is fitted.
+ model
+ The model instance which is fitted to the dataset, which does not include the perturbed feature.
+ paths
+ The `Paths` instance which contains the path to the folder where the results of the fit are written to.
+ instance
+ The simulation instance, which includes the perturbed feature that is used to simulate the dataset.
+ This is often not used, but may be useful for certain sensitivity mapping tasks, for example using
+ true values of the simulated instance to set up aspects of the model-fit (e.g. the priors).
+ """
+
+ search = af.DynestyStatic(
+ paths=paths.for_sub_analysis(analysis_name="[base]"),
+ nlive=50,
+ iterations_per_full_update=50000,
+ )
+
+ analysis = self.analysis_cls(dataset=dataset)
+
+ return search.fit(model=model, analysis=analysis)
+
+
+"""
+__Perturb Fit__
+
+We now define a `PerturbFit` class, which defines how the `perturb_model` is fitted to each simulated dataset. This
+behaves analogously to the `BaseFit` class above, but now fits the `perturb_model` to the simulated data (as
+opposed to the `base_model`).
+
+Again, in this example we use a full non-linear search to fit the `perturb_model` to the simulated data and return
+the `log_evidence` of the model fit as the goodness-of-fit. This fit could easily be something much simpler and
+more computationally efficient, for example performing a single log likelihood evaluation of the `perturb_model` fit
+to the simulated data.
+"""
+
+
+class PerturbFit:
+ def __init__(self, analysis_cls):
+ """
+ Class used to fit every dataset used for sensitivity mapping with the perturbed model (the model with the
+ perturbed feature sensitivity mapping maps out).
+
+ In this example, the perturbed model therefore includes the extra Gaussian feature, which is also in the
+ simulated dataset.
+
+ The perturbed fit is repeated for every parameter on the sensitivity grid and compared to the base fit. This
+ maps out the sensitivity of every parameter is (e.g. the sensitivity of the normalization of the Gaussian
+ feature).
+
+ The `__init__` constructor can be extended with new inputs which can be used to control how the dataset is
+ fitted, below we include an input `analysis_cls` which is the `Analysis` class used to fit the model to the
+ dataset.
+
+ Parameters
+ ----------
+ analysis_cls
+ The `Analysis` class used to fit the model to the dataset.
+ """
+ self.analysis_cls = analysis_cls
+
+ def __call__(self, dataset, model, paths, instance):
+ """
+ The perturbed fitting function which fits every dataset used for sensitivity mapping with the perturbed model.
+
+ This function receives as input each simulated dataset of the sensitivity map and fits it, in order to
+ quantify how sensitive the model is to the perturbed feature.
+
+ In this example, a full non-linear search is performed to determine how well the model fits the dataset.
+ The `log_evidence` of the fit is returned which acts as the sensitivity map figure of merit.
+
+ Parameters
+ ----------
+ dataset
+ The dataset which is simulated with the perturbed model and which is fitted.
+ model
+ The model instance which is fitted to the dataset, which includes the perturbed feature.
+ paths
+ The `Paths` instance which contains the path to the folder where the results of the fit are written to.
+ instance
+ The simulation instance, which includes the perturbed feature that is used to simulate the dataset.
+ This is often not used, but may be useful for certain sensitivity mapping tasks, for example using
+ true values of the simulated instance to set up aspects of the model-fit (e.g. the priors).
+ """
+
+ search = af.DynestyStatic(
+ paths=paths.for_sub_analysis(analysis_name="[perturbed]"),
+ nlive=50,
+ iterations_per_full_update=50000,
+ )
+
+ analysis = self.analysis_cls(dataset=dataset)
+
+ return search.fit(model=model, analysis=analysis)
+
+
+"""
+We can now combine all of the objects created above and perform sensitivity mapping. The inputs to the `Sensitivity`
+object below are:
+
+- `simulation_instance`: This is an instance of the model used to simulate every dataset that is fitted. In this
+example it contains an instance of the `gaussian_main` model component.
+
+- `base_model`: This is the simpler model that is fitted to every simulated dataset, which in this example is composed
+of a single `Gaussian` called the `gaussian_main`.
+
+- `perturb_model`: This is the extra model component that has two roles: (i) based on the sensitivity grid parameters
+it is added to the `simulation_instance` to simulate each dataset ; (ii) it is added to the`base_model` and fitted to
+every simulated dataset (in this example every `simulation_instance` and `perturb_model` there has two `Gaussians`
+called the `gaussian_main` and `gaussian_feature`).
+
+- `simulate_cls`: This is the function that uses the `simulation_instance` and many instances of the `perturb_model`
+to simulate many datasets which are fitted with the `base_model` and `base_model` + `perturb_model`.
+
+- `base_fit_cls`: This is the function that fits the `base_model` to every simulated dataset and returns the
+goodness-of-fit of the model to the data.
+
+- `perturb_fit_cls`: This is the function that fits the `base_model` + `perturb_model` to every simulated dataset and
+returns the goodness-of-fit of the model to the data.
+
+- `number_of_steps`: The number of steps over which the parameters in the `perturb_model` are iterated. In this
+example, normalization has a `LogUniformPrior` with lower limit 1e-4 and upper limit 1e2, therefore the `number_of_steps`
+of 2 wills imulate and fit just 2 datasets where the intensities between 1e-4 and 1e2.
+
+- `number_of_cores`: The number of cores over which the sensitivity mapping is performed, enabling parallel processing
+if set above 1.
+"""
+paths = af.DirectoryPaths(
+ path_prefix=path.join("features"),
+ unique_tag="hello",
+ name="sensitivity_mapping",
+)
+
+sensitivity = af.Sensitivity(
+ paths=paths,
+ simulation_instance=simulation_instance,
+ base_model=base_model,
+ perturb_model=perturb_model,
+ simulate_cls=Simulate(),
+ base_fit_cls=BaseFit(analysis_cls=Analysis),
+ perturb_fit_cls=PerturbFit(analysis_cls=Analysis),
+ number_of_steps=2,
+ number_of_cores=2,
+)
+sensitivity_result = sensitivity.run()
+
+"""
+__Results__
+
+You should now look at the results of the sensitivity mapping in the folder `output/features/sensitivity_mapping`.
+
+You will note the following 4 model-fits have been performed:
+
+ - The `base_model` is fitted to a simulated dataset where the `simulation_instance` and
+ a `perturb` with `normalization=0.01` are used.
+
+ - The `base_model` + `perturb_model` is fitted to a simulated dataset where the `simulation_instance` and
+ a `perturb` with `normalization=0.01` are used.
+
+ - The `base_model` is fitted to a simulated dataset where the `simulation_instance` and
+ a `perturb` with `normalization=100.0` are used.
+
+ - The `base_model` + `perturb_model` is fitted to a simulated dataset where the `simulation_instance` and
+ a `perturb` with `normalization=100.0` are used.
+
+The fit produced a `sensitivity_result`.
+
+We are still developing the `SensitivityResult` class to provide a data structure that better streamlines the analysis
+of results. If you intend to use sensitivity mapping, the best way to interpret the resutls is currently via
+**PyAutoFit**'s database and `Aggregator` tools.
+"""
+print(sensitivity_result.samples)
+print(sensitivity_result.log_evidences_base)
+
+"""
+Finish.
+"""
diff --git a/scripts/howtofit/chapter_1_introduction/tutorial_1_models.py b/scripts/howtofit/chapter_1_introduction/tutorial_1_models.py
index dfa91577..33866c3f 100644
--- a/scripts/howtofit/chapter_1_introduction/tutorial_1_models.py
+++ b/scripts/howtofit/chapter_1_introduction/tutorial_1_models.py
@@ -1,560 +1,556 @@
-"""
-Tutorial 1: Models
-==================
-
-At the heart of model-fitting is the model: a set of equations, numerical processes, and assumptions describing a
-physical system of interest. The goal of model-fitting is to better understand this physical system and develop
-predictive models that describe it more accurately.
-
-In astronomy, a model might describe the distribution of stars within a galaxy. In biology, it might represent the
-interaction of proteins within a cell. In finance, it could describe the evolution of stock prices in a market.
-Regardless of the field, the model acts as a mathematical description of the physical system, aiming to enhance
-understanding and enable new predictions.
-
-Whatever your model, its equations are defined by "free parameters." Changing these parameters alters the
-behavior and predictions of the model.
-
-Once the model is defined and parameter values are chosen, the model creates "model data"—a realization of how the
-physical system appears given those parameters. This process, often referred to as "forward modeling," describes the
-physical system from its starting point and predicts the data we observe.
-
-By varying the model parameters, we can generate numerous model datasets. The ultimate goal of model-fitting, which
-you will learn by the end of this chapter, is to determine the model parameters and corresponding dataset that best
-fit the observed data.
-
-__Astronomy Example__
-
-For instance, in astronomy, we might model the distribution of stars, including:
-
-- A parameter describing the brightness of the stars.
-
-- Multiple parameters defining their distribution.
-
-- Several parameters describing their colors.
-
-If our model pertains to the distribution of stars within a galaxy, the forward model will produce an image of what
-that galaxy looks like when observed with a telescope. This forward model might account for physical effects such as
-the blurring of light due to diffraction in the telescope optics.
-
-By altering the parameters describing the stars, we can generate many different model images via this forward model.
-
-At the end of this chapter, we will use a real-world astronomy example to illustrate everything you have learned,
-including fitting a real galaxy observed with the Hubble Space Telescope.
-
-__Overview__
-
-In tutorial 1, we will cover the basics of defining a model, specifically:
-
-- Defining a simple model described by a few simple equations.
-
-- Showing that this model is characterized by three or more free parameters.
-
-- Using the model, with different sets of parameters, to generate model data.
-
-__Contents__
-
-This tutorial is split into the following sections:
-
-- **Paths**: Setting up the working directory path so the tutorial runs correctly on your computer.
-- **PyProjRoot**: A brief introduction to the PyProjRoot package, which sets the working directory for Jupiter notebooks.
-- **Model Parameterization**: An example of how a model is parameterized and is made up of free parameters.
-- **Model Composition**: Composing a model using PyAutoFit's model composition API.
-- **Model Creation**: Creating an instance of the model using PyAutoFit's `Model` python object.
-- **Model Mapping**: Mapping an input vector of parameters to the model to create an instance of the model.
-- **Complex Models**: Composing a more complex model with multiple model components and more free parameters.
-- **Tuple Parameters**: Defining a model component with tuple parameters.
-- **Extensibility**: Discussing how PyAutoFit's model composition API is scalable and extensible.
-- **Wrap Up**: Concluding the tutorial and considering how to apply the concepts to your own scientific problem.
-
-This tutorial introduces the PyAutoFit API for model composition, which forms the foundation of all model-fitting
-performed by PyAutoFit.
-"""
-
-import numpy as np
-import matplotlib.pyplot as plt
-
-import autofit as af
-
-"""
-__Paths__
-
-PyAutoFit assumes the current working directory is /path/to/autofit_workspace/ on your hard-disk (or in Binder).
-This setup allows PyAutoFit to:
-
-- Load configuration settings from config files in the autofit_workspace/config folder.
-
-- Load example data from the autofit_workspace/dataset folder.
-
-- Output the results of model fits to your hard disk in the autofit/output folder.
-
-If you don't have an autofit_workspace, you can download it here:
-
- https://github.com/Jammy2211/autofit_workspace
-
-__PyProjRoot__
-
-At the top of every tutorial notebook, you will see the following cell. This cell uses the project pyprojroot to
-locate the path to the workspace on your computer and set it as the working directory of the notebook.
-"""
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-"""
-__Model Parameterization__
-
-A model is a set of equations, numerical processes, and assumptions that describe a physical system and dataset.
-
-In this example, our model is one or more 1-dimensional Gaussians, defined by the following equation:
-
-\begin{equation*}
-g(x, I, \sigma) = \frac{N}{\sigma\sqrt{2\pi}} \exp{(-0.5 (x / \sigma)^2)}
-\end{equation*}
-
-Where:
-
-- `x`: The x-axis coordinate where the Gaussian is evaluated.
-
-- `N`: The overall normalization of the Gaussian.
-
-
-- `\sigma`: The size of the Gaussian (Full Width Half Maximum, $\mathrm{FWHM}$, is $2{\sqrt{2\ln 2}}\;\sigma$).
-
-While a 1D Gaussian might seem like a rudimentary model, it has many real-world applications in signal processing.
-For example, 1D Gaussians are fitted to datasets to measure the size of an observed signal. Thus, this model has
-practical real-world applications.
-
-We now have a model, expressed as a simple 1D Gaussian. The model has three parameters, $(x, N, \sigma)$. Using
-different combinations of these parameters creates different realizations of the model, which we illustrate below.
-
-__Model Composition__
-
-We now define the 1D Gaussian as a "model component" in PyAutoFit. We use the term "model component" because the model
-can be extended to include multiple components, each related to different equations and numerical processes.
-
-We first illustrate a model composed of a single model component, the 1D Gaussian. We then show a model made of
-multiple model components.
-
-To define a "model component" in PyAutoFit, we simply write it as a Python class using the format shown below:
-"""
-
-
-class Gaussian:
- def __init__(
- self,
- centre: float = 30.0, # <- **PyAutoFit** recognises these constructor arguments
- normalization: float = 1.0, # <- are the Gaussian`s model parameters.
- sigma: float = 5.0,
- ):
- """
- Represents a 1D Gaussian profile.
-
- This is a model-component of example models in the **HowToFit** lectures and is used to perform model-fitting
- of example datasets.
-
- Parameters
- ----------
- centre
- The x coordinate of the profile centre.
- normalization
- Overall normalization of the profile.
- sigma
- The sigma value controlling the size of the Gaussian.
- """
- self.centre = centre
- self.normalization = normalization
- self.sigma = sigma
-
- def model_data_from(self, xvalues: np.ndarray) -> np.ndarray:
- """
- Returns a 1D Gaussian on an input list of Cartesian x coordinates.
-
- The input xvalues are translated to a coordinate system centred on the Gaussian, via its `centre`.
-
- The output is referred to as the `model_data` to signify that it is a representation of the data from the
- model.
-
- Parameters
- ----------
- xvalues
- The x coordinates in the original reference frame of the data.
-
- Returns
- -------
- np.array
- The Gaussian values at the input x coordinates.
- """
- transformed_xvalues = np.subtract(xvalues, self.centre)
- return np.multiply(
- np.divide(self.normalization, self.sigma * np.sqrt(2.0 * np.pi)),
- np.exp(-0.5 * np.square(np.divide(transformed_xvalues, self.sigma))),
- )
-
-
-"""
-The format of this Python class defines how PyAutoFit composes the Gaussian as a model component, where:
-
-- The name of the class is the name of the model component, in this case, "Gaussian".
-
-- The input arguments of the constructor (the `__init__` method) are the parameters of the model, in the example
-above `centre`, `normalization`, and `sigma`.
-
-- The default values and typing of the input arguments define whether a parameter is a single-valued float or a
-multi-valued tuple. For the `Gaussian` class above, no input parameters are tuples, but later examples use tuples.
-
-- It includes functions associated with that model component, specifically the model_data function. When we create
-instances of a `Gaussian` below, this function is used to generate a 1D representation of it as a NumPy array.
-
-__Model Creation__
-
-The `Gaussian` class above is a standard Python class. It does not yet act as a model component that can be used
-for model fitting with PyAutoFit.
-
-To transform the Gaussian class into a model component that can be used for model fitting with PyAutoFit, we use
-the `af.Model` object. This tells PyAutoFit to treat the input Python class as a model component.
-"""
-model = af.Model(Gaussian)
-print("Model `Gaussian` object: \n")
-print(model)
-
-"""
-In PyAutoFit, a Model object encapsulates a model component that can be used for model fitting. It provides several
-attributes that describe the model component, such as the `total_free_parameters` attribute, which indicates the
-number of free parameters in the model:
-"""
-print(model.total_free_parameters)
-
-"""
-In PyAutoFit, you can retrieve comprehensive information about a model by accessing its `info` attribute.
-
-When you print the model info, it displays detailed information about each parameter in the model, including its name,
-type, and associated prior distribution. Priors define the expected range or distribution of values for each
-parameter during the model fitting process. If you're unfamiliar with priors, they are covered in tutorial 3 of
-this chapter, which explains their role in model fitting.
-
-[The `info` below may not display optimally on your computer screen, for example the whitespace between parameter
-names on the left and parameter priors on the right may lead them to appear across multiple lines. This is a
-common issue in Jupyter notebooks.
-
-The`info_whitespace_length` parameter in the file `config/general.yaml` in the "output" section can be changed to
-increase or decrease the amount of whitespace (The Jupyter notebook kernel will need to be reset for this change to
-appear in a notebook).]
-"""
-print(model.info)
-
-"""
-__Model Mapping__
-
-In PyAutoFit, instances of model components created via the af.Model object can be instantiated by mapping an input
-vector of parameters to the Python class that the model object represents. The order of parameters in the model is
-crucial for correctly defining the input vector.
-
-To determine the order of parameters in the model, PyAutoFit provides the paths attribute of the model object.
-This attribute contains information about the parameter paths within the model.
-
-Here's how you can access the paths attribute to understand the order of parameters in the model:
-"""
-print(model.paths)
-
-"""
-To create an instance of the Gaussian model component using PyAutoFit, following the order of parameters defined by
-the paths attribute (`centre`, `normalization`, and `sigma`), you can initialize the instance as follows:
-"""
-instance = model.instance_from_vector(vector=[30.0, 2.0, 3.0])
-
-"""
-This is an instance of the `Gaussian` class.
-"""
-print("Model Instance: \n")
-print(instance)
-
-"""
-It has the parameters of the `Gaussian` with the values input above.
-"""
-print("Instance Parameters \n")
-print("x = ", instance.centre)
-print("normalization = ", instance.normalization)
-print("sigma = ", instance.sigma)
-
-"""
-We can use all class functions, such as the `model_data_from` function, to generate an instance of the
-1D `Gaussian` and visualize it through plotting.
-
-The code below generates the 1D Gaussian model data, which requires an input list of x values where the Gaussian is
-evaluated. The output is a NumPy array of the Gaussian's y values at the input x coordinates.
-
-Although simple, the code below is essentially the process of forward modeling, where we use the model to generate
-the data we would observe in an experiment for a given set of parameters.
-"""
-xvalues = np.arange(0.0, 100.0, 1.0)
-
-model_data = instance.model_data_from(xvalues=xvalues)
-
-plt.plot(xvalues, model_data, color="r")
-plt.title("1D Gaussian Model Data.")
-plt.xlabel("x values of profile")
-plt.ylabel("Gaussian Value")
-plt.show()
-plt.clf()
-
-"""
-__Complex Models__
-
-The code above may seem like a lot of steps just to create an instance of the `Gaussian` class. Couldn't we have
-simply done this instead?
-
-```python
-instance = Gaussian(centre=30.0, normalization=2.0, sigma=3.0)
-```
-
-Yes, we could have.
-
-However, the model composition API used above is designed to simplify the process of composing complex models that
-consist of multiple components with many free parameters. It provides a scalable approach for defining and
-manipulating models.
-
-To demonstrate this capability, let's conclude the tutorial by composing a model composed of a Gaussian
-component and another 1D profile, an `Exponential`, defined by the equation:
-
-\begin{equation*}
-g(x, I, \lambda) = N \lambda \exp{- \lambda x }
-\end{equation*}
-
-where:
-
-- `x`: Represents the x-axis coordinate where the Exponential profile is evaluated.
-
-- `N`: Describes the overall normalization of the Exponential profile.
-
-- $\lambda$: Represents the rate of decay of the exponential.
-
-We'll start by defining the `Exponential` profile using a format similar to the Gaussian definition above.
-"""
-
-
-class Exponential:
- def __init__(
- self,
- centre: float = 30.0, # <- **PyAutoFit** recognises these constructor arguments
- normalization: float = 1.0, # <- are the Exponential`s model parameters.
- rate: float = 0.01,
- ):
- """
- Represents a 1D Exponential profile.
-
- This is a model-component of example models in the **HowToFit** lectures and is used to fit example datasets
- via a non-linear search.
-
- Parameters
- ----------
- centre
- The x coordinate of the profile centre.
- normalization
- Overall normalization of the profile.
- ratw
- The decay rate controlling has fast the Exponential declines.
- """
- self.centre = centre
- self.normalization = normalization
- self.rate = rate
-
- def model_data_from(self, xvalues: np.ndarray):
- """
- Returns a 1D Gaussian on an input list of Cartesian x coordinates.
-
- The input xvalues are translated to a coordinate system centred on the `Exponential`, via its `centre`.
-
- The output is referred to as the `model_data` to signify that it is a representation of the data from the
- model.
-
- Parameters
- ----------
- xvalues
- The x coordinates in the original reference frame of the data.
- """
- transformed_xvalues = np.subtract(xvalues, self.centre)
- return self.normalization * np.multiply(
- self.rate, np.exp(-1.0 * self.rate * abs(transformed_xvalues))
- )
-
-
-"""
-We can construct a model comprising one `Gaussian` object and one `Exponential` object using the `af.Collection` object:
-"""
-model = af.Collection(gaussian=af.Model(Gaussian), exponential=af.Model(Exponential))
-
-"""
-You can retrieve all the information about the model created via the `af.Collection` by printing its `info` attribute
-in one go:
-"""
-print(model.info)
-
-"""
-When `Gaussian` and `Exponential` are added to a `Collection`, they are automatically assigned as `Model` objects.
-
-Therefore, there's no need to use the `af.Model` method when passing classes to a `Collection`, which makes the Python
-code more concise and readable.
-"""
-model = af.Collection(gaussian=Gaussian, exponential=Exponential)
-
-"""
-The `model.info` is identical to the previous example.
-"""
-print(model.info)
-
-"""
-A `Collection` functions analogously to a `Model`, but it includes multiple model components.
-
-This can be observed by examining its `paths` attribute, which displays paths to all 6 free parameters across both model components.
-
-The paths contain entries such as `.gaussian.` and `.exponential.`, corresponding to the names we provided when
-defining the `af.Collection` earlier. Modifying the names of the model components supplied to the `Collection`
-would adjust the paths accordingly.
-"""
-print(model.paths)
-
-"""
-A model instance can again be created by mapping an input `vector`, which now has 6 entries.
-"""
-instance = model.instance_from_vector(vector=[0.1, 0.2, 0.3, 0.4, 0.5, 0.01])
-
-"""
-This `instance` contains each of the model components we defined above.
-
-The argument names input into the `Collection` define the attribute names of the `instance`:
-"""
-print("Instance Parameters \n")
-print("x (Gaussian) = ", instance.gaussian.centre)
-print("normalization (Gaussian) = ", instance.gaussian.normalization)
-print("sigma (Gaussian) = ", instance.gaussian.sigma)
-print("x (Exponential) = ", instance.exponential.centre)
-print("normalization (Exponential) = ", instance.exponential.normalization)
-print("sigma (Exponential) = ", instance.exponential.rate)
-
-"""
-In the context of the model's equations, it is simply the sum of the equations defining the `Gaussian`
-and `Exponential` components.
-
-To generate the `model_data`, we sum the `model_data` of each individual model component, as demonstrated and
-visualized below.
-"""
-xvalues = np.arange(0.0, 100.0, 1.0)
-
-model_data_0 = instance.gaussian.model_data_from(xvalues=xvalues)
-model_data_1 = instance.exponential.model_data_from(xvalues=xvalues)
-
-model_data = model_data_0 + model_data_1
-
-plt.plot(xvalues, model_data, color="r")
-plt.plot(xvalues, model_data_0, "b", "--")
-plt.plot(xvalues, model_data_1, "k", "--")
-plt.title("1D Gaussian + Exponential Model Data.")
-plt.xlabel("x values of profile")
-plt.ylabel("Value")
-plt.show()
-plt.clf()
-
-"""
-__Tuple Parameters__
-
-The `Gaussian` and `Exponential` model components above only has parameters that are single-valued floats.
-
-Parameters can also be tuples, which is useful for defining model components where certain parameters are naturally
-grouped together.
-
-For example, we can define a 2D Gaussian with a center that has two coordinates and therefore free parameters, (x, y),
-using a tuple.
-"""
-from typing import Tuple
-
-
-class Gaussian2D:
- def __init__(
- self,
- centre: Tuple[float, float] = (0.0, 0.0),
- normalization: float = 0.1,
- sigma: float = 1.0,
- ):
- self.centre = centre
- self.normalization = normalization
- self.sigma = sigma
-
-
-"""
-The model's `total_free_parameters` attribute now includes 4 free parameters, as the tuple `centre` parameter accounts
-for 2 free parameters.
-"""
-model = af.Model(Gaussian2D)
-
-print("Total Free Parameters:", model.total_free_parameters)
-
-"""
-This information is again displayed in the `info` attribute:
-"""
-print("\nInfo:")
-print(model.info)
-
-"""
-The `paths` attribute provides information on the order of parameters in the model, illustrating how the
-`centre` tuple is split into two parameters.
-"""
-print("\nPaths:")
-print(model.paths)
-
-"""
-This ordering is used to create an instance of the `Gaussian2D` model component:
-"""
-instance = model.instance_from_vector(vector=[40.0, 60.0, 2.0, 3.0])
-
-print("\nInstance Parameters:")
-print("centre (x) = ", instance.centre[0])
-print("centre (y) = ", instance.centre[1])
-print("normalization = ", instance.normalization)
-print("sigma = ", instance.sigma)
-
-"""
-__Extensibility__
-
-It should now be clear why we use `Model` and `Collection` objects to construct our model.
-
-These objects facilitate the straightforward extension of our models to include multiple components and parameters.
-For instance, we can add more `Gaussian` and `Exponential` components to the `Collection`, or define new Python
-classes to represent entirely new model components with additional parameters.
-
-These objects serve numerous other essential purposes that we will explore in subsequent tutorials.
-
-**PyAutoFit** offers a comprehensive API for building models, which includes models constructed using NumPy arrays,
-hierarchies of Python classes, and graphical models where parameters are interconnected. These advanced modeling
-techniques are gradually introduced throughout the HowToFit lectures.
-
-For a detailed understanding of PyAutoFit's model composition API and a quick reference guide on how to construct
-models, you may want to take a quick look at the model cookbook in the PyAutoFit documentation. It provides an
-extensive overview and can serve as a helpful resource as you progress:
-
-[PyAutoFit Model Cookbook](https://pyautofit.readthedocs.io/en/latest/cookbooks/model.html)
-
-Don't worry if it seems a bit overwhelming at this stage; the concepts will become clearer as you continue exploring
-and working with PyAutoFit.
-
-__Wrap Up__
-
-In this tutorial, we've learned how to define and compose a model that can generate model data.
-
-Now, think about your specific field of study and the problem you want to address through model-fitting. Consider
-the following questions:
-
-- What type of model would best describe your data?
-
-- Which Python class, following the format introduced here, would you need to compose this model?
-
-- What are the free parameters of your model that need to be determined through fitting?
-
-If you decide to incorporate a new model component into your autofit_workspace tailored to your specific model-fitting
-task, refer to the following script:
-
-autofit_workspace/*/overview/new_model_component/new_model_component.ipynb
-
-This script provides guidance on setting up the PyAutoFit configuration files associated with your custom model.
-"""
+"""
+Tutorial 1: Models
+==================
+
+At the heart of model-fitting is the model: a set of equations, numerical processes, and assumptions describing a
+physical system of interest. The goal of model-fitting is to better understand this physical system and develop
+predictive models that describe it more accurately.
+
+In astronomy, a model might describe the distribution of stars within a galaxy. In biology, it might represent the
+interaction of proteins within a cell. In finance, it could describe the evolution of stock prices in a market.
+Regardless of the field, the model acts as a mathematical description of the physical system, aiming to enhance
+understanding and enable new predictions.
+
+Whatever your model, its equations are defined by "free parameters." Changing these parameters alters the
+behavior and predictions of the model.
+
+Once the model is defined and parameter values are chosen, the model creates "model data"—a realization of how the
+physical system appears given those parameters. This process, often referred to as "forward modeling," describes the
+physical system from its starting point and predicts the data we observe.
+
+By varying the model parameters, we can generate numerous model datasets. The ultimate goal of model-fitting, which
+you will learn by the end of this chapter, is to determine the model parameters and corresponding dataset that best
+fit the observed data.
+
+__Astronomy Example__
+
+For instance, in astronomy, we might model the distribution of stars, including:
+
+- A parameter describing the brightness of the stars.
+
+- Multiple parameters defining their distribution.
+
+- Several parameters describing their colors.
+
+If our model pertains to the distribution of stars within a galaxy, the forward model will produce an image of what
+that galaxy looks like when observed with a telescope. This forward model might account for physical effects such as
+the blurring of light due to diffraction in the telescope optics.
+
+By altering the parameters describing the stars, we can generate many different model images via this forward model.
+
+At the end of this chapter, we will use a real-world astronomy example to illustrate everything you have learned,
+including fitting a real galaxy observed with the Hubble Space Telescope.
+
+__Overview__
+
+In tutorial 1, we will cover the basics of defining a model, specifically:
+
+- Defining a simple model described by a few simple equations.
+
+- Showing that this model is characterized by three or more free parameters.
+
+- Using the model, with different sets of parameters, to generate model data.
+
+__Contents__
+
+This tutorial is split into the following sections:
+
+- **Paths**: Setting up the working directory path so the tutorial runs correctly on your computer.
+- **PyProjRoot**: A brief introduction to the PyProjRoot package, which sets the working directory for Jupiter notebooks.
+- **Model Parameterization**: An example of how a model is parameterized and is made up of free parameters.
+- **Model Composition**: Composing a model using PyAutoFit's model composition API.
+- **Model Creation**: Creating an instance of the model using PyAutoFit's `Model` python object.
+- **Model Mapping**: Mapping an input vector of parameters to the model to create an instance of the model.
+- **Complex Models**: Composing a more complex model with multiple model components and more free parameters.
+- **Tuple Parameters**: Defining a model component with tuple parameters.
+- **Extensibility**: Discussing how PyAutoFit's model composition API is scalable and extensible.
+- **Wrap Up**: Concluding the tutorial and considering how to apply the concepts to your own scientific problem.
+
+This tutorial introduces the PyAutoFit API for model composition, which forms the foundation of all model-fitting
+performed by PyAutoFit.
+"""
+
+import numpy as np
+import matplotlib.pyplot as plt
+
+import autofit as af
+
+"""
+__Paths__
+
+PyAutoFit assumes the current working directory is /path/to/autofit_workspace/ on your hard-disk (or in Binder).
+This setup allows PyAutoFit to:
+
+- Load configuration settings from config files in the autofit_workspace/config folder.
+
+- Load example data from the autofit_workspace/dataset folder.
+
+- Output the results of model fits to your hard disk in the autofit/output folder.
+
+If you don't have an autofit_workspace, you can download it here:
+
+ https://github.com/Jammy2211/autofit_workspace
+
+__Workspace Setup__
+
+At the top of every tutorial notebook, you will see the following cell. This cell finds the workspace root
+directory on your computer and sets it as the working directory of the notebook.
+"""
+# from autoconf import setup_notebook; setup_notebook()
+
+"""
+__Model Parameterization__
+
+A model is a set of equations, numerical processes, and assumptions that describe a physical system and dataset.
+
+In this example, our model is one or more 1-dimensional Gaussians, defined by the following equation:
+
+\begin{equation*}
+g(x, I, \sigma) = \frac{N}{\sigma\sqrt{2\pi}} \exp{(-0.5 (x / \sigma)^2)}
+\end{equation*}
+
+Where:
+
+- `x`: The x-axis coordinate where the Gaussian is evaluated.
+
+- `N`: The overall normalization of the Gaussian.
+
+
+- `\sigma`: The size of the Gaussian (Full Width Half Maximum, $\mathrm{FWHM}$, is $2{\sqrt{2\ln 2}}\;\sigma$).
+
+While a 1D Gaussian might seem like a rudimentary model, it has many real-world applications in signal processing.
+For example, 1D Gaussians are fitted to datasets to measure the size of an observed signal. Thus, this model has
+practical real-world applications.
+
+We now have a model, expressed as a simple 1D Gaussian. The model has three parameters, $(x, N, \sigma)$. Using
+different combinations of these parameters creates different realizations of the model, which we illustrate below.
+
+__Model Composition__
+
+We now define the 1D Gaussian as a "model component" in PyAutoFit. We use the term "model component" because the model
+can be extended to include multiple components, each related to different equations and numerical processes.
+
+We first illustrate a model composed of a single model component, the 1D Gaussian. We then show a model made of
+multiple model components.
+
+To define a "model component" in PyAutoFit, we simply write it as a Python class using the format shown below:
+"""
+
+
+class Gaussian:
+ def __init__(
+ self,
+ centre: float = 30.0, # <- **PyAutoFit** recognises these constructor arguments
+ normalization: float = 1.0, # <- are the Gaussian`s model parameters.
+ sigma: float = 5.0,
+ ):
+ """
+ Represents a 1D Gaussian profile.
+
+ This is a model-component of example models in the **HowToFit** lectures and is used to perform model-fitting
+ of example datasets.
+
+ Parameters
+ ----------
+ centre
+ The x coordinate of the profile centre.
+ normalization
+ Overall normalization of the profile.
+ sigma
+ The sigma value controlling the size of the Gaussian.
+ """
+ self.centre = centre
+ self.normalization = normalization
+ self.sigma = sigma
+
+ def model_data_from(self, xvalues: np.ndarray) -> np.ndarray:
+ """
+ Returns a 1D Gaussian on an input list of Cartesian x coordinates.
+
+ The input xvalues are translated to a coordinate system centred on the Gaussian, via its `centre`.
+
+ The output is referred to as the `model_data` to signify that it is a representation of the data from the
+ model.
+
+ Parameters
+ ----------
+ xvalues
+ The x coordinates in the original reference frame of the data.
+
+ Returns
+ -------
+ np.array
+ The Gaussian values at the input x coordinates.
+ """
+ transformed_xvalues = np.subtract(xvalues, self.centre)
+ return np.multiply(
+ np.divide(self.normalization, self.sigma * np.sqrt(2.0 * np.pi)),
+ np.exp(-0.5 * np.square(np.divide(transformed_xvalues, self.sigma))),
+ )
+
+
+"""
+The format of this Python class defines how PyAutoFit composes the Gaussian as a model component, where:
+
+- The name of the class is the name of the model component, in this case, "Gaussian".
+
+- The input arguments of the constructor (the `__init__` method) are the parameters of the model, in the example
+above `centre`, `normalization`, and `sigma`.
+
+- The default values and typing of the input arguments define whether a parameter is a single-valued float or a
+multi-valued tuple. For the `Gaussian` class above, no input parameters are tuples, but later examples use tuples.
+
+- It includes functions associated with that model component, specifically the model_data function. When we create
+instances of a `Gaussian` below, this function is used to generate a 1D representation of it as a NumPy array.
+
+__Model Creation__
+
+The `Gaussian` class above is a standard Python class. It does not yet act as a model component that can be used
+for model fitting with PyAutoFit.
+
+To transform the Gaussian class into a model component that can be used for model fitting with PyAutoFit, we use
+the `af.Model` object. This tells PyAutoFit to treat the input Python class as a model component.
+"""
+model = af.Model(Gaussian)
+print("Model `Gaussian` object: \n")
+print(model)
+
+"""
+In PyAutoFit, a Model object encapsulates a model component that can be used for model fitting. It provides several
+attributes that describe the model component, such as the `total_free_parameters` attribute, which indicates the
+number of free parameters in the model:
+"""
+print(model.total_free_parameters)
+
+"""
+In PyAutoFit, you can retrieve comprehensive information about a model by accessing its `info` attribute.
+
+When you print the model info, it displays detailed information about each parameter in the model, including its name,
+type, and associated prior distribution. Priors define the expected range or distribution of values for each
+parameter during the model fitting process. If you're unfamiliar with priors, they are covered in tutorial 3 of
+this chapter, which explains their role in model fitting.
+
+[The `info` below may not display optimally on your computer screen, for example the whitespace between parameter
+names on the left and parameter priors on the right may lead them to appear across multiple lines. This is a
+common issue in Jupyter notebooks.
+
+The`info_whitespace_length` parameter in the file `config/general.yaml` in the "output" section can be changed to
+increase or decrease the amount of whitespace (The Jupyter notebook kernel will need to be reset for this change to
+appear in a notebook).]
+"""
+print(model.info)
+
+"""
+__Model Mapping__
+
+In PyAutoFit, instances of model components created via the af.Model object can be instantiated by mapping an input
+vector of parameters to the Python class that the model object represents. The order of parameters in the model is
+crucial for correctly defining the input vector.
+
+To determine the order of parameters in the model, PyAutoFit provides the paths attribute of the model object.
+This attribute contains information about the parameter paths within the model.
+
+Here's how you can access the paths attribute to understand the order of parameters in the model:
+"""
+print(model.paths)
+
+"""
+To create an instance of the Gaussian model component using PyAutoFit, following the order of parameters defined by
+the paths attribute (`centre`, `normalization`, and `sigma`), you can initialize the instance as follows:
+"""
+instance = model.instance_from_vector(vector=[30.0, 2.0, 3.0])
+
+"""
+This is an instance of the `Gaussian` class.
+"""
+print("Model Instance: \n")
+print(instance)
+
+"""
+It has the parameters of the `Gaussian` with the values input above.
+"""
+print("Instance Parameters \n")
+print("x = ", instance.centre)
+print("normalization = ", instance.normalization)
+print("sigma = ", instance.sigma)
+
+"""
+We can use all class functions, such as the `model_data_from` function, to generate an instance of the
+1D `Gaussian` and visualize it through plotting.
+
+The code below generates the 1D Gaussian model data, which requires an input list of x values where the Gaussian is
+evaluated. The output is a NumPy array of the Gaussian's y values at the input x coordinates.
+
+Although simple, the code below is essentially the process of forward modeling, where we use the model to generate
+the data we would observe in an experiment for a given set of parameters.
+"""
+xvalues = np.arange(0.0, 100.0, 1.0)
+
+model_data = instance.model_data_from(xvalues=xvalues)
+
+plt.plot(xvalues, model_data, color="r")
+plt.title("1D Gaussian Model Data.")
+plt.xlabel("x values of profile")
+plt.ylabel("Gaussian Value")
+plt.show()
+plt.clf()
+
+"""
+__Complex Models__
+
+The code above may seem like a lot of steps just to create an instance of the `Gaussian` class. Couldn't we have
+simply done this instead?
+
+```python
+instance = Gaussian(centre=30.0, normalization=2.0, sigma=3.0)
+```
+
+Yes, we could have.
+
+However, the model composition API used above is designed to simplify the process of composing complex models that
+consist of multiple components with many free parameters. It provides a scalable approach for defining and
+manipulating models.
+
+To demonstrate this capability, let's conclude the tutorial by composing a model composed of a Gaussian
+component and another 1D profile, an `Exponential`, defined by the equation:
+
+\begin{equation*}
+g(x, I, \lambda) = N \lambda \exp{- \lambda x }
+\end{equation*}
+
+where:
+
+- `x`: Represents the x-axis coordinate where the Exponential profile is evaluated.
+
+- `N`: Describes the overall normalization of the Exponential profile.
+
+- $\lambda$: Represents the rate of decay of the exponential.
+
+We'll start by defining the `Exponential` profile using a format similar to the Gaussian definition above.
+"""
+
+
+class Exponential:
+ def __init__(
+ self,
+ centre: float = 30.0, # <- **PyAutoFit** recognises these constructor arguments
+ normalization: float = 1.0, # <- are the Exponential`s model parameters.
+ rate: float = 0.01,
+ ):
+ """
+ Represents a 1D Exponential profile.
+
+ This is a model-component of example models in the **HowToFit** lectures and is used to fit example datasets
+ via a non-linear search.
+
+ Parameters
+ ----------
+ centre
+ The x coordinate of the profile centre.
+ normalization
+ Overall normalization of the profile.
+ ratw
+ The decay rate controlling has fast the Exponential declines.
+ """
+ self.centre = centre
+ self.normalization = normalization
+ self.rate = rate
+
+ def model_data_from(self, xvalues: np.ndarray):
+ """
+ Returns a 1D Gaussian on an input list of Cartesian x coordinates.
+
+ The input xvalues are translated to a coordinate system centred on the `Exponential`, via its `centre`.
+
+ The output is referred to as the `model_data` to signify that it is a representation of the data from the
+ model.
+
+ Parameters
+ ----------
+ xvalues
+ The x coordinates in the original reference frame of the data.
+ """
+ transformed_xvalues = np.subtract(xvalues, self.centre)
+ return self.normalization * np.multiply(
+ self.rate, np.exp(-1.0 * self.rate * abs(transformed_xvalues))
+ )
+
+
+"""
+We can construct a model comprising one `Gaussian` object and one `Exponential` object using the `af.Collection` object:
+"""
+model = af.Collection(gaussian=af.Model(Gaussian), exponential=af.Model(Exponential))
+
+"""
+You can retrieve all the information about the model created via the `af.Collection` by printing its `info` attribute
+in one go:
+"""
+print(model.info)
+
+"""
+When `Gaussian` and `Exponential` are added to a `Collection`, they are automatically assigned as `Model` objects.
+
+Therefore, there's no need to use the `af.Model` method when passing classes to a `Collection`, which makes the Python
+code more concise and readable.
+"""
+model = af.Collection(gaussian=Gaussian, exponential=Exponential)
+
+"""
+The `model.info` is identical to the previous example.
+"""
+print(model.info)
+
+"""
+A `Collection` functions analogously to a `Model`, but it includes multiple model components.
+
+This can be observed by examining its `paths` attribute, which displays paths to all 6 free parameters across both model components.
+
+The paths contain entries such as `.gaussian.` and `.exponential.`, corresponding to the names we provided when
+defining the `af.Collection` earlier. Modifying the names of the model components supplied to the `Collection`
+would adjust the paths accordingly.
+"""
+print(model.paths)
+
+"""
+A model instance can again be created by mapping an input `vector`, which now has 6 entries.
+"""
+instance = model.instance_from_vector(vector=[0.1, 0.2, 0.3, 0.4, 0.5, 0.01])
+
+"""
+This `instance` contains each of the model components we defined above.
+
+The argument names input into the `Collection` define the attribute names of the `instance`:
+"""
+print("Instance Parameters \n")
+print("x (Gaussian) = ", instance.gaussian.centre)
+print("normalization (Gaussian) = ", instance.gaussian.normalization)
+print("sigma (Gaussian) = ", instance.gaussian.sigma)
+print("x (Exponential) = ", instance.exponential.centre)
+print("normalization (Exponential) = ", instance.exponential.normalization)
+print("sigma (Exponential) = ", instance.exponential.rate)
+
+"""
+In the context of the model's equations, it is simply the sum of the equations defining the `Gaussian`
+and `Exponential` components.
+
+To generate the `model_data`, we sum the `model_data` of each individual model component, as demonstrated and
+visualized below.
+"""
+xvalues = np.arange(0.0, 100.0, 1.0)
+
+model_data_0 = instance.gaussian.model_data_from(xvalues=xvalues)
+model_data_1 = instance.exponential.model_data_from(xvalues=xvalues)
+
+model_data = model_data_0 + model_data_1
+
+plt.plot(xvalues, model_data, color="r")
+plt.plot(xvalues, model_data_0, "b", "--")
+plt.plot(xvalues, model_data_1, "k", "--")
+plt.title("1D Gaussian + Exponential Model Data.")
+plt.xlabel("x values of profile")
+plt.ylabel("Value")
+plt.show()
+plt.clf()
+
+"""
+__Tuple Parameters__
+
+The `Gaussian` and `Exponential` model components above only has parameters that are single-valued floats.
+
+Parameters can also be tuples, which is useful for defining model components where certain parameters are naturally
+grouped together.
+
+For example, we can define a 2D Gaussian with a center that has two coordinates and therefore free parameters, (x, y),
+using a tuple.
+"""
+from typing import Tuple
+
+
+class Gaussian2D:
+ def __init__(
+ self,
+ centre: Tuple[float, float] = (0.0, 0.0),
+ normalization: float = 0.1,
+ sigma: float = 1.0,
+ ):
+ self.centre = centre
+ self.normalization = normalization
+ self.sigma = sigma
+
+
+"""
+The model's `total_free_parameters` attribute now includes 4 free parameters, as the tuple `centre` parameter accounts
+for 2 free parameters.
+"""
+model = af.Model(Gaussian2D)
+
+print("Total Free Parameters:", model.total_free_parameters)
+
+"""
+This information is again displayed in the `info` attribute:
+"""
+print("\nInfo:")
+print(model.info)
+
+"""
+The `paths` attribute provides information on the order of parameters in the model, illustrating how the
+`centre` tuple is split into two parameters.
+"""
+print("\nPaths:")
+print(model.paths)
+
+"""
+This ordering is used to create an instance of the `Gaussian2D` model component:
+"""
+instance = model.instance_from_vector(vector=[40.0, 60.0, 2.0, 3.0])
+
+print("\nInstance Parameters:")
+print("centre (x) = ", instance.centre[0])
+print("centre (y) = ", instance.centre[1])
+print("normalization = ", instance.normalization)
+print("sigma = ", instance.sigma)
+
+"""
+__Extensibility__
+
+It should now be clear why we use `Model` and `Collection` objects to construct our model.
+
+These objects facilitate the straightforward extension of our models to include multiple components and parameters.
+For instance, we can add more `Gaussian` and `Exponential` components to the `Collection`, or define new Python
+classes to represent entirely new model components with additional parameters.
+
+These objects serve numerous other essential purposes that we will explore in subsequent tutorials.
+
+**PyAutoFit** offers a comprehensive API for building models, which includes models constructed using NumPy arrays,
+hierarchies of Python classes, and graphical models where parameters are interconnected. These advanced modeling
+techniques are gradually introduced throughout the HowToFit lectures.
+
+For a detailed understanding of PyAutoFit's model composition API and a quick reference guide on how to construct
+models, you may want to take a quick look at the model cookbook in the PyAutoFit documentation. It provides an
+extensive overview and can serve as a helpful resource as you progress:
+
+[PyAutoFit Model Cookbook](https://pyautofit.readthedocs.io/en/latest/cookbooks/model.html)
+
+Don't worry if it seems a bit overwhelming at this stage; the concepts will become clearer as you continue exploring
+and working with PyAutoFit.
+
+__Wrap Up__
+
+In this tutorial, we've learned how to define and compose a model that can generate model data.
+
+Now, think about your specific field of study and the problem you want to address through model-fitting. Consider
+the following questions:
+
+- What type of model would best describe your data?
+
+- Which Python class, following the format introduced here, would you need to compose this model?
+
+- What are the free parameters of your model that need to be determined through fitting?
+
+If you decide to incorporate a new model component into your autofit_workspace tailored to your specific model-fitting
+task, refer to the following script:
+
+autofit_workspace/*/overview/new_model_component/new_model_component.ipynb
+
+This script provides guidance on setting up the PyAutoFit configuration files associated with your custom model.
+"""
diff --git a/scripts/howtofit/chapter_1_introduction/tutorial_2_fitting_data.py b/scripts/howtofit/chapter_1_introduction/tutorial_2_fitting_data.py
index a9755afd..585fc4c7 100644
--- a/scripts/howtofit/chapter_1_introduction/tutorial_2_fitting_data.py
+++ b/scripts/howtofit/chapter_1_introduction/tutorial_2_fitting_data.py
@@ -1,657 +1,653 @@
-"""
-Tutorial 2: Fitting Data
-========================
-
-We've learned that a model consists of equations, numerical processes, and assumptions that describe a physical system.
-Using **PyAutoFit**, we defined simple 1D models like the Gaussian, composed them into models using `Model` and
-`Collection` objects, and generated model data by varying their parameters.
-
-To apply our model to real-world situations, we must fit it to data. Fitting involves assessing how well the model
-matches observed data. A good fit indicates that the model's parameter values accurately describe the physical system.
-Conversely, a poor fit suggests that adjustments are needed to better reflect reality.
-
-Model-fitting is a cyclical process: define the model, fit it to data, and refine the model based on insights gained.
-Iteratively improving the model's complexity enhances its ability to accurately represent the system under study.
-This iterative process lies at the core of model-fitting in scientific analysis.
-
-__Astronomy Example__
-
-In Astronomy, this process has been crucial for understanding the distributions of stars within galaxies. By
-fitting high-quality images of galaxies with increasingly sophisticated models, astronomers have determined that
-stars within galaxies are organized into structures such as disks, bars, and bulges. This approach has also revealed
-that stars appear differently in red and blue images due to variations in their age and composition.
-
-__Overview__
-
-In this tutorial, we will explore how to fit the `model_data` generated by a model to actual data. Specifically, we will:
-
-- Load data representing a 1D Gaussian signal, which serves as our target dataset for fitting.
-
-- Compute quantities such as residuals by subtracting the model data from the observed data.
-
-- Quantitatively assess the goodness-of-fit using a critical measure in model-fitting known as the `log_likelihood`.
-
-All these steps will utilize the **PyAutoFit** API for model composition, introduced in the previous tutorial.
-
-__Contents__
-
-This tutorial is split into the following sections:
-
-- **Data**: Load and plot the 1D Gaussian dataset we will fit.
-- **Model Data**: Generate model data of the `Gaussian` model using a forward model.
-- **Residuals**: Compute and visualize residuals between the model data and observed data.
-- **Normalized Residuals**: Compute and visualize normalized residuals, which account for the noise properties of the data.
-- **Chi Squared**: Compute and visualize the chi-squared map, a measure of the overall goodness-of-fit.
-- **Noise Normalization**: Compute the noise normalization term which describes the noise properties of the data.
-- **Likelihood**: Compute the log likelihood, a key measure of the goodness-of-fit of the model to the data.
-- **Recap**: Summarize the standard metrics for quantifying model fit quality.
-- **Fitting Models**: Fit the `Gaussian` model to the 1D data and compute the log likelihood, by guessing parameters.
-- **Guess 1**: A first parameter guess with an explanation of the resulting log likelihood.
-- **Guess 2**: An improved parameter guess with a better log likelihood.
-- **Guess 3**: The optimal parameter guess providing the best fit to the data.
-- **Extensibility**: Use the `Collection` object for fitting models with multiple components.
-- **Wrap Up**: Summarize the key concepts of this tutorial.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-from os import path
-import matplotlib.pyplot as plt
-import numpy as np
-
-import autofit as af
-
-"""
-__Data__
-
-Our dataset consists of noisy 1D data containing a signal, where the underlying signal can be modeled using
-equations such as a 1D Gaussian, a 1D Exponential, or a combination of multiple 1D profiles.
-
-We load this dataset from .json files, where:
-
-- `data` is a 1D NumPy array containing values representing the observed signal.
-
-- `noise_map` is a 1D NumPy array containing values representing the estimated root mean squared (RMS) noise level at
- each data point.
-
-These datasets are generated using scripts located in `autofit_workspace/howtofit/simulators`. Feel free to explore
-these scripts for more details!
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-"""
-Next, we visualize the 1D signal using `matplotlib`.
-
-The signal is observed over uniformly spaced `xvalues`, computed using the `arange` function and `data.shape[0]` method.
-
-We will reuse these `xvalues` shortly when generating model data from the model.
-"""
-xvalues = np.arange(data.shape[0])
-plt.plot(xvalues, data, color="k")
-plt.title("1D Dataset Containing a Gaussian.")
-plt.xlabel("x values of profile")
-plt.ylabel("Signal Value")
-plt.show()
-
-"""
-The earlier plot depicted only the signal without indicating the estimated noise at each data point.
-
-To visualize both the signal and its `noise_map`, we can use `matplotlib`'s `errorbar` function.
-"""
-plt.errorbar(
- xvalues,
- data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.title("1D Gaussian dataset with errors from the noise-map.")
-plt.xlabel("x values of profile")
-plt.ylabel("Signal Value")
-plt.show()
-
-
-"""
-__Model Data__
-
-To fit our `Gaussian` model to this data, we start by generating `model_data` from the 1D `Gaussian` model,
-following the same steps as outlined in the previous tutorial.
-
-We begin by again defining the `Gaussian` class, following the **PyAutoFit** format for model components.
-"""
-
-
-class Gaussian:
- def __init__(
- self,
- centre: float = 30.0, # <- **PyAutoFit** recognises these constructor arguments
- normalization: float = 1.0, # <- are the Gaussian`s model parameters.
- sigma: float = 5.0,
- ):
- """
- Represents a 1D Gaussian profile.
-
- This is a model-component of example models in the **HowToFit** lectures and is used to perform model-fitting
- of example datasets.
-
- Parameters
- ----------
- centre
- The x coordinate of the profile centre.
- normalization
- Overall normalization of the profile.
- sigma
- The sigma value controlling the size of the Gaussian.
- """
- self.centre = centre
- self.normalization = normalization
- self.sigma = sigma
-
- def model_data_from(self, xvalues: np.ndarray) -> np.ndarray:
- """
- Returns a 1D Gaussian on an input list of Cartesian x coordinates.
-
- The input xvalues are translated to a coordinate system centred on the Gaussian, via its `centre`.
-
- The output is referred to as the `model_data` to signify that it is a representation of the data from the
- model.
-
- Parameters
- ----------
- xvalues
- The x coordinates in the original reference frame of the data.
-
- Returns
- -------
- np.array
- The Gaussian values at the input x coordinates.
- """
- transformed_xvalues = np.subtract(xvalues, self.centre)
- return np.multiply(
- np.divide(self.normalization, self.sigma * np.sqrt(2.0 * np.pi)),
- np.exp(-0.5 * np.square(np.divide(transformed_xvalues, self.sigma))),
- )
-
-
-"""
-To create `model_data` for the `Gaussian`, we use the model by providing it with `xvalues` corresponding to the
-observed data, as demonstrated in the previous tutorial.
-
-The following code essentially utilizes a forward model to generate the model data based on a specified set of
-parameters.
-"""
-model = af.Model(Gaussian)
-
-gaussian = model.instance_from_vector(vector=[60.0, 20.0, 15.0])
-
-model_data = gaussian.model_data_from(xvalues=xvalues)
-
-plt.plot(xvalues, model_data, color="r")
-plt.title("1D Gaussian model.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile Normalization")
-plt.show()
-plt.clf()
-
-"""
-For comparison purposes, it is more informative to plot both the `data` and `model_data` on the same plot.
-"""
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(xvalues, model_data, color="r")
-plt.title("Model-data fit to 1D Gaussian data.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-Changing the values of `centre`, `normalization`, and `sigma` alters the appearance of the `Gaussian`.
-
-You can modify the parameters passed into `instance_from_vector()` above. After recomputing the `model_data`, plot
-it again to observe how these changes affect the Gaussian's appearance.
-
-__Residuals__
-
-While it's informative to compare the `data` and `model_data` above, gaining insights from the residuals can be even
-more useful.
-
-Residuals are calculated as `data - model_data` in 1D:
-"""
-residual_map = data - model_data
-plt.plot(xvalues, residual_map, color="k")
-plt.title("Residuals of model-data fit to 1D Gaussian data.")
-plt.xlabel("x values of profile")
-plt.ylabel("Residuals")
-plt.show()
-plt.clf()
-
-"""
-Are these residuals indicative of a good fit to the data? Without considering the noise in the data, it's difficult
-to ascertain.
-
-We can plot the residuals with error bars based on the noise map. The plot below reveals that the model is a poor fit,
-as many residuals deviate significantly from zero even after accounting for the noise in each data point.
-
-A blue line through zero is included on the plot, to make it clear where residuals are not constent with zero
-above the noise level.
-"""
-residual_map = data - model_data
-plt.plot(range(data.shape[0]), np.zeros(data.shape[0]), "--", color="b")
-plt.errorbar(
- x=xvalues,
- y=residual_map,
- yerr=noise_map,
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
- linestyle="",
-)
-plt.title("Residuals of model-data fit to 1D Gaussian data.")
-plt.xlabel("x values of profile")
-plt.ylabel("Residuals")
-plt.show()
-plt.clf()
-
-"""
-__Normalized Residuals__
-
-Another method to quantify and visualize the quality of the fit is using the normalized residual map, also known as
-standardized residuals.
-
-The normalized residual map is computed as the residual map divided by the noise map:
-
-\[ \text{normalized\_residual} = \frac{\text{residual\_map}}{\text{noise\_map}} = \frac{\text{data} - \text{model\_data}}{\text{noise\_map}} \]
-
-If you're familiar with the concept of standard deviations (sigma) in statistics, the normalized residual map represents
-how many standard deviations the residual is from zero. For instance, a normalized residual of 2.0 (corresponding
-to a 95% confidence interval) means that the probability of the model underestimating the data by that amount is only 5%.
-
-Both the residual map with error bars and the normalized residual map convey the same information. However,
-the normalized residual map is particularly useful for visualization in multidimensional problems, as plotting
-error bars in 2D or higher dimensions is not straightforward.
-"""
-normalized_residual_map = residual_map / noise_map
-plt.plot(xvalues, normalized_residual_map, color="k")
-plt.title("Normalized residuals of model-data fit to 1D Gaussian data.")
-plt.xlabel("x values of profile")
-plt.ylabel("Normalized Residuals")
-plt.show()
-plt.clf()
-
-"""
-__Chi Squared__
-
-Next, we define the `chi_squared_map`, which is obtained by squaring the `normalized_residual_map` and serves as a
-measure of goodness of fit.
-
-The chi-squared map is calculated as:
-
-\[ \chi^2 = \left(\frac{\text{data} - \text{model\_data}}{\text{noise\_map}}\right)^2 \]
-
-The purpose of squaring the normalized residual map is to ensure all values are positive. For instance, both a
-normalized residual of -0.2 and 0.2 would square to 0.04, indicating the same level of fit in terms of `chi_squared`.
-
-As seen from the normalized residual map, it's evident that the model does not provide a good fit to the data.
-"""
-chi_squared_map = (normalized_residual_map) ** 2
-plt.plot(xvalues, chi_squared_map, color="k")
-plt.title("Chi-Squared Map of model-data fit to 1D Gaussian data.")
-plt.xlabel("x values of profile")
-plt.ylabel("Chi-Squareds")
-plt.show()
-plt.clf()
-
-"""
-Now, we consolidate all the information in our `chi_squared_map` into a single measure of goodness-of-fit
-called `chi_squared`.
-
-It is defined as the sum of all values in the `chi_squared_map` and is computed as:
-
-\[ \chi^2 = \sum \left(\frac{\text{data} - \text{model\_data}}{\text{noise\_map}}\right)^2 \]
-
-This summing process highlights why ensuring all values in the chi-squared map are positive is crucial. If we
-didn't square the values (making them positive), positive and negative residuals would cancel each other out,
-leading to an inaccurate assessment of the model's fit to the data.
-"""
-chi_squared = np.sum(chi_squared_map)
-print("Chi-squared = ", chi_squared)
-
-"""
-
-The lower the `chi_squared`, the fewer residuals exist between the model's fit and the data, indicating a better
-overall fit!
-
-__Noise Normalization__
-
-Next, we introduce another quantity that contributes to our final assessment of the goodness-of-fit:
-the `noise_normalization`.
-
-The `noise_normalization` is computed as the logarithm of the sum of squared noise values in our data:
-
-\[
-\text{{noise\_normalization}} = \sum \log(2 \pi \text{{noise\_map}}^2)
-\]
-
-This quantity is fixed because the noise-map remains constant throughout the fitting process. Despite this,
-including the `noise_normalization` is considered good practice due to its statistical significance.
-
-Understanding the exact meaning of `noise_normalization` isn't critical for our primary goal of successfully
-fitting a model to a dataset. Essentially, it provides a measure of how well the noise properties of our data align
-with a Gaussian distribution.
-"""
-noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
-
-"""
-__Likelihood__
-
-From the `chi_squared` and `noise_normalization`, we can define a final goodness-of-fit measure known as
-the `log_likelihood`.
-
-This measure is calculated by taking the sum of the `chi_squared` and `noise_normalization`, and then multiplying the
-result by -0.5:
-
-\[ \text{log\_likelihood} = -0.5 \times \left( \chi^2 + \text{noise\_normalization} \right) \]
-
-Why multiply by -0.5? The exact rationale behind this factor isn't critical for our current understanding.
-"""
-log_likelihood = -0.5 * (chi_squared + noise_normalization)
-print("Log Likelihood = ", log_likelihood)
-
-"""
-Above, we mentioned that a lower `chi_squared` indicates a better fit of the model to the data.
-
-When calculating the `log_likelihood`, we multiply the `chi_squared` by -0.5. Therefore, a higher log likelihood
-corresponds to a better model fit. This is what we aim for when fitting models to data, we want to maximize the
-log likelihood!
-
-__Recap__
-
-If you're familiar with model-fitting, you've likely encountered terms like 'residuals', 'chi-squared',
-and 'log_likelihood' before.
-
-These metrics are standard ways to quantify the quality of a model fit. They are applicable not only to 1D data but
-also to more complex data structures like 2D images, 3D data cubes, or any other multidimensional datasets.
-
-If these terms are new to you, it's important to understand their meanings as they form the basis of all
-model-fitting operations in **PyAutoFit** (and in statistical inference more broadly).
-
-Let's recap what we've learned so far:
-
-- We can define models, such as a 1D `Gaussian`, using Python classes that follow a specific format.
-
-- Models can be organized using `Collection` and `Model` objects, with parameters mapped to instances of their
- respective model classes (e.g., `Gaussian`).
-
-- Using these model instances, we can generate model data, compare it to observed data, and quantify the
- goodness-of-fit using the log likelihood.
-
-__Fitting Models__
-
-Now, armed with this knowledge, we are ready to fit our model to our data!
-
-But how do we find the best-fit model, which maximizes the log likelihood?
-
-The simplest approach is to guess parameters. Starting with initial parameter values that yield a good
-fit (i.e., a higher log likelihood), we iteratively adjust these values to refine our model until we achieve an
-optimal fit.
-
-For a 1D `Gaussian`, this iterative process works effectively. Below, we fit three different `Gaussian` models and
-identify the best-fit model—the one that matches the original dataset most closely.
-
-To streamline this process, I've developed functions that compute the `log_likelihood` of a model fit and visualize
-the data alongside the model predictions, complete with error bars.
-"""
-
-
-def log_likelihood_from(
- data: np.ndarray, noise_map: np.ndarray, model_data: np.ndarray
-) -> float:
- """
- Compute the log likelihood of a model fit to data given the noise map.
-
- Parameters
- ----------
- data
- The observed data.
- noise_map
- The root mean square noise (or uncertainty) associated with each data point.
- model_data
- The model's predicted data for the given data x points.
-
- Returns
- -------
- float
- The log likelihood of the model fit to the data.
- """
- # Calculate residuals and normalized residuals
- residual_map = data - model_data
- normalized_residual_map = residual_map / noise_map
-
- # Compute chi-squared and noise normalization
- chi_squared_map = normalized_residual_map**2
- chi_squared = np.sum(chi_squared_map)
- noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
-
- # Compute log likelihood
- log_likelihood = -0.5 * (chi_squared + noise_normalization)
-
- return log_likelihood
-
-
-def plot_model_fit(
- xvalues: np.ndarray,
- data: np.ndarray,
- noise_map: np.ndarray,
- model_data: np.ndarray,
- color: str = "k",
-):
- """
- Plot the observed data, model predictions, and error bars.
-
- Parameters
- ----------
- xvalues
- The x-axis values where the data is observed and model is predicted.
- data
- The observed data points.
- noise_map
- The root mean squared noise (or uncertainty) associated with each data point.
- model_data
- The model's predicted data for the given data x points.
- color
- The color for plotting (default is "k" for black).
- """
- plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- linestyle="",
- color=color,
- ecolor="k",
- elinewidth=1,
- capsize=2,
- )
- plt.plot(xvalues, model_data, color="r")
- plt.title("Fit of model-data to data")
- plt.xlabel("x values of profile")
- plt.ylabel("Profile Value")
- plt.show()
- plt.clf() # Clear figure to prevent overlapping plots
-
-
-"""
-__Guess 1__
-
-The first guess correctly pinpoints that the Gaussian's peak is at 50.0, but the width and normalization are off.
-
-The `log_likelihood` is computed and printed, however because we don't have a value to compare it to yet, its hard
-to assess if it is a large or small value.
-"""
-
-gaussian = model.instance_from_vector(vector=[50.0, 10.0, 5.0])
-model_data = gaussian.model_data_from(xvalues=xvalues)
-plot_model_fit(
- xvalues=xvalues,
- data=data,
- noise_map=noise_map,
- model_data=model_data,
- color="r",
-)
-
-log_likelihood = log_likelihood_from(
- data=data, noise_map=noise_map, model_data=model_data
-)
-print(f"Log Likelihood: {log_likelihood}")
-
-"""
-__Guess 2__
-
-The second guess refines the width and normalization, but the size of the Gaussian is still off.
-
-The `log_likelihood` is computed and printed, and increases a lot compared to the previous guess, indicating that
-the fit is better.
-"""
-
-gaussian = model.instance_from_vector(vector=[50.0, 25.0, 5.0])
-model_data = gaussian.model_data_from(xvalues=xvalues)
-plot_model_fit(
- xvalues=xvalues,
- data=data,
- noise_map=noise_map,
- model_data=model_data,
- color="r",
-)
-
-log_likelihood = log_likelihood_from(
- data=data, noise_map=noise_map, model_data=model_data
-)
-print(f"Log Likelihood: {log_likelihood}")
-
-"""
-__Guess 3__
-
-The third guess provides a good fit to the data, with the Gaussian's peak, width, and normalization all accurately
-representing the observed signal.
-
-The `log_likelihood` is computed and printed, and is the highest value yet, indicating that this model provides the
-best fit to the data.
-"""
-
-gaussian = model.instance_from_vector(vector=[50.0, 25.0, 10.0])
-model_data = gaussian.model_data_from(xvalues=xvalues)
-plot_model_fit(
- xvalues=xvalues,
- data=data,
- noise_map=noise_map,
- model_data=model_data,
- color="r",
-)
-
-log_likelihood = log_likelihood_from(
- data=data, noise_map=noise_map, model_data=model_data
-)
-print(f"Log Likelihood: {log_likelihood}")
-
-"""
-__Extensibility__
-
-Fitting models composed of multiple components is straightforward with PyAutoFit. Using the `Collection` object,
-we can define complex models consisting of several components. Once defined, we generate `model_data`
-from this collection and fit it to the observed data to compute the log likelihood.
-"""
-model = af.Collection(gaussian_0=Gaussian, gaussian_1=Gaussian)
-
-instance = model.instance_from_vector(vector=[40.0, 0.2, 0.3, 60.0, 0.5, 1.0])
-
-model_data_0 = instance.gaussian_0.model_data_from(xvalues=xvalues)
-model_data_1 = instance.gaussian_1.model_data_from(xvalues=xvalues)
-
-model_data = model_data_0 + model_data_1
-
-"""
-We plot the data and model data below, showing that we get a bad fit (a low log likelihood) for this model.
-
-We could attempt to improve the model-fit and find a higher log likelihood solution by varying the parameters of
-the two Gaussians. However, with 6 parameters, this would be a challenging and cumbersome task to perform by eye.
-"""
-plot_model_fit(
- xvalues=xvalues,
- data=data,
- noise_map=noise_map,
- model_data=model_data,
- color="r",
-)
-
-log_likelihood = log_likelihood_from(
- data=data, noise_map=noise_map, model_data=model_data
-)
-print(f"Log Likelihood: {log_likelihood}")
-
-
-"""
-When our model consisted of only 3 parameters, it was manageable to visually guess their values and achieve a good
-fit to the data. However, as we expanded our model to include six parameters, this approach quickly became
-inefficient. Attempting to manually optimize models with even more parameters would effectively become impossible,
-and a more systematic approach is required.
-
-In the next tutorial, we will introduce an automated approach for fitting models to data. This method will enable
-us to systematically determine the optimal values of model parameters that best describe the observed data, without
-relying on manual guesswork.
-
-__Wrap Up__
-
-To conclude, take a moment to reflect on the model you ultimately aim to fit using **PyAutoFit**. What does your
-data look like? Is it one-dimensional, like a spectrum or a time series? Or is it two-dimensional, such as an image
-or a map? Visualize the nature of your data and consider whether you can define a mathematical model that
-accurately generates similar data.
-
-Can you imagine what a residual map would look like if you were to compare your model's predictions against this
-data? A residual map shows the differences between observed data and the model's predictions, often revealing
-patterns or areas where the model fits well or poorly.
-
-Furthermore, can you foresee how you would calculate a log likelihood from this residual map? The log likelihood q
-uantifies how well your model fits the data, incorporating both the residual values and the noise characteristics of
-your observations.
-
-If you find it challenging to visualize these aspects right now, that's perfectly fine. The first step is to
-grasp the fundamentals of fitting a model to data using **PyAutoFit**, which will provide you with the tools
-and understanding needed to address these questions effectively in the future.
-"""
+"""
+Tutorial 2: Fitting Data
+========================
+
+We've learned that a model consists of equations, numerical processes, and assumptions that describe a physical system.
+Using **PyAutoFit**, we defined simple 1D models like the Gaussian, composed them into models using `Model` and
+`Collection` objects, and generated model data by varying their parameters.
+
+To apply our model to real-world situations, we must fit it to data. Fitting involves assessing how well the model
+matches observed data. A good fit indicates that the model's parameter values accurately describe the physical system.
+Conversely, a poor fit suggests that adjustments are needed to better reflect reality.
+
+Model-fitting is a cyclical process: define the model, fit it to data, and refine the model based on insights gained.
+Iteratively improving the model's complexity enhances its ability to accurately represent the system under study.
+This iterative process lies at the core of model-fitting in scientific analysis.
+
+__Astronomy Example__
+
+In Astronomy, this process has been crucial for understanding the distributions of stars within galaxies. By
+fitting high-quality images of galaxies with increasingly sophisticated models, astronomers have determined that
+stars within galaxies are organized into structures such as disks, bars, and bulges. This approach has also revealed
+that stars appear differently in red and blue images due to variations in their age and composition.
+
+__Overview__
+
+In this tutorial, we will explore how to fit the `model_data` generated by a model to actual data. Specifically, we will:
+
+- Load data representing a 1D Gaussian signal, which serves as our target dataset for fitting.
+
+- Compute quantities such as residuals by subtracting the model data from the observed data.
+
+- Quantitatively assess the goodness-of-fit using a critical measure in model-fitting known as the `log_likelihood`.
+
+All these steps will utilize the **PyAutoFit** API for model composition, introduced in the previous tutorial.
+
+__Contents__
+
+This tutorial is split into the following sections:
+
+- **Data**: Load and plot the 1D Gaussian dataset we will fit.
+- **Model Data**: Generate model data of the `Gaussian` model using a forward model.
+- **Residuals**: Compute and visualize residuals between the model data and observed data.
+- **Normalized Residuals**: Compute and visualize normalized residuals, which account for the noise properties of the data.
+- **Chi Squared**: Compute and visualize the chi-squared map, a measure of the overall goodness-of-fit.
+- **Noise Normalization**: Compute the noise normalization term which describes the noise properties of the data.
+- **Likelihood**: Compute the log likelihood, a key measure of the goodness-of-fit of the model to the data.
+- **Recap**: Summarize the standard metrics for quantifying model fit quality.
+- **Fitting Models**: Fit the `Gaussian` model to the 1D data and compute the log likelihood, by guessing parameters.
+- **Guess 1**: A first parameter guess with an explanation of the resulting log likelihood.
+- **Guess 2**: An improved parameter guess with a better log likelihood.
+- **Guess 3**: The optimal parameter guess providing the best fit to the data.
+- **Extensibility**: Use the `Collection` object for fitting models with multiple components.
+- **Wrap Up**: Summarize the key concepts of this tutorial.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+from os import path
+import matplotlib.pyplot as plt
+import numpy as np
+
+import autofit as af
+
+"""
+__Data__
+
+Our dataset consists of noisy 1D data containing a signal, where the underlying signal can be modeled using
+equations such as a 1D Gaussian, a 1D Exponential, or a combination of multiple 1D profiles.
+
+We load this dataset from .json files, where:
+
+- `data` is a 1D NumPy array containing values representing the observed signal.
+
+- `noise_map` is a 1D NumPy array containing values representing the estimated root mean squared (RMS) noise level at
+ each data point.
+
+These datasets are generated using scripts located in `autofit_workspace/howtofit/simulators`. Feel free to explore
+these scripts for more details!
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+"""
+Next, we visualize the 1D signal using `matplotlib`.
+
+The signal is observed over uniformly spaced `xvalues`, computed using the `arange` function and `data.shape[0]` method.
+
+We will reuse these `xvalues` shortly when generating model data from the model.
+"""
+xvalues = np.arange(data.shape[0])
+plt.plot(xvalues, data, color="k")
+plt.title("1D Dataset Containing a Gaussian.")
+plt.xlabel("x values of profile")
+plt.ylabel("Signal Value")
+plt.show()
+
+"""
+The earlier plot depicted only the signal without indicating the estimated noise at each data point.
+
+To visualize both the signal and its `noise_map`, we can use `matplotlib`'s `errorbar` function.
+"""
+plt.errorbar(
+ xvalues,
+ data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.title("1D Gaussian dataset with errors from the noise-map.")
+plt.xlabel("x values of profile")
+plt.ylabel("Signal Value")
+plt.show()
+
+
+"""
+__Model Data__
+
+To fit our `Gaussian` model to this data, we start by generating `model_data` from the 1D `Gaussian` model,
+following the same steps as outlined in the previous tutorial.
+
+We begin by again defining the `Gaussian` class, following the **PyAutoFit** format for model components.
+"""
+
+
+class Gaussian:
+ def __init__(
+ self,
+ centre: float = 30.0, # <- **PyAutoFit** recognises these constructor arguments
+ normalization: float = 1.0, # <- are the Gaussian`s model parameters.
+ sigma: float = 5.0,
+ ):
+ """
+ Represents a 1D Gaussian profile.
+
+ This is a model-component of example models in the **HowToFit** lectures and is used to perform model-fitting
+ of example datasets.
+
+ Parameters
+ ----------
+ centre
+ The x coordinate of the profile centre.
+ normalization
+ Overall normalization of the profile.
+ sigma
+ The sigma value controlling the size of the Gaussian.
+ """
+ self.centre = centre
+ self.normalization = normalization
+ self.sigma = sigma
+
+ def model_data_from(self, xvalues: np.ndarray) -> np.ndarray:
+ """
+ Returns a 1D Gaussian on an input list of Cartesian x coordinates.
+
+ The input xvalues are translated to a coordinate system centred on the Gaussian, via its `centre`.
+
+ The output is referred to as the `model_data` to signify that it is a representation of the data from the
+ model.
+
+ Parameters
+ ----------
+ xvalues
+ The x coordinates in the original reference frame of the data.
+
+ Returns
+ -------
+ np.array
+ The Gaussian values at the input x coordinates.
+ """
+ transformed_xvalues = np.subtract(xvalues, self.centre)
+ return np.multiply(
+ np.divide(self.normalization, self.sigma * np.sqrt(2.0 * np.pi)),
+ np.exp(-0.5 * np.square(np.divide(transformed_xvalues, self.sigma))),
+ )
+
+
+"""
+To create `model_data` for the `Gaussian`, we use the model by providing it with `xvalues` corresponding to the
+observed data, as demonstrated in the previous tutorial.
+
+The following code essentially utilizes a forward model to generate the model data based on a specified set of
+parameters.
+"""
+model = af.Model(Gaussian)
+
+gaussian = model.instance_from_vector(vector=[60.0, 20.0, 15.0])
+
+model_data = gaussian.model_data_from(xvalues=xvalues)
+
+plt.plot(xvalues, model_data, color="r")
+plt.title("1D Gaussian model.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile Normalization")
+plt.show()
+plt.clf()
+
+"""
+For comparison purposes, it is more informative to plot both the `data` and `model_data` on the same plot.
+"""
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(xvalues, model_data, color="r")
+plt.title("Model-data fit to 1D Gaussian data.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+Changing the values of `centre`, `normalization`, and `sigma` alters the appearance of the `Gaussian`.
+
+You can modify the parameters passed into `instance_from_vector()` above. After recomputing the `model_data`, plot
+it again to observe how these changes affect the Gaussian's appearance.
+
+__Residuals__
+
+While it's informative to compare the `data` and `model_data` above, gaining insights from the residuals can be even
+more useful.
+
+Residuals are calculated as `data - model_data` in 1D:
+"""
+residual_map = data - model_data
+plt.plot(xvalues, residual_map, color="k")
+plt.title("Residuals of model-data fit to 1D Gaussian data.")
+plt.xlabel("x values of profile")
+plt.ylabel("Residuals")
+plt.show()
+plt.clf()
+
+"""
+Are these residuals indicative of a good fit to the data? Without considering the noise in the data, it's difficult
+to ascertain.
+
+We can plot the residuals with error bars based on the noise map. The plot below reveals that the model is a poor fit,
+as many residuals deviate significantly from zero even after accounting for the noise in each data point.
+
+A blue line through zero is included on the plot, to make it clear where residuals are not constent with zero
+above the noise level.
+"""
+residual_map = data - model_data
+plt.plot(range(data.shape[0]), np.zeros(data.shape[0]), "--", color="b")
+plt.errorbar(
+ x=xvalues,
+ y=residual_map,
+ yerr=noise_map,
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+ linestyle="",
+)
+plt.title("Residuals of model-data fit to 1D Gaussian data.")
+plt.xlabel("x values of profile")
+plt.ylabel("Residuals")
+plt.show()
+plt.clf()
+
+"""
+__Normalized Residuals__
+
+Another method to quantify and visualize the quality of the fit is using the normalized residual map, also known as
+standardized residuals.
+
+The normalized residual map is computed as the residual map divided by the noise map:
+
+\[ \text{normalized\_residual} = \frac{\text{residual\_map}}{\text{noise\_map}} = \frac{\text{data} - \text{model\_data}}{\text{noise\_map}} \]
+
+If you're familiar with the concept of standard deviations (sigma) in statistics, the normalized residual map represents
+how many standard deviations the residual is from zero. For instance, a normalized residual of 2.0 (corresponding
+to a 95% confidence interval) means that the probability of the model underestimating the data by that amount is only 5%.
+
+Both the residual map with error bars and the normalized residual map convey the same information. However,
+the normalized residual map is particularly useful for visualization in multidimensional problems, as plotting
+error bars in 2D or higher dimensions is not straightforward.
+"""
+normalized_residual_map = residual_map / noise_map
+plt.plot(xvalues, normalized_residual_map, color="k")
+plt.title("Normalized residuals of model-data fit to 1D Gaussian data.")
+plt.xlabel("x values of profile")
+plt.ylabel("Normalized Residuals")
+plt.show()
+plt.clf()
+
+"""
+__Chi Squared__
+
+Next, we define the `chi_squared_map`, which is obtained by squaring the `normalized_residual_map` and serves as a
+measure of goodness of fit.
+
+The chi-squared map is calculated as:
+
+\[ \chi^2 = \left(\frac{\text{data} - \text{model\_data}}{\text{noise\_map}}\right)^2 \]
+
+The purpose of squaring the normalized residual map is to ensure all values are positive. For instance, both a
+normalized residual of -0.2 and 0.2 would square to 0.04, indicating the same level of fit in terms of `chi_squared`.
+
+As seen from the normalized residual map, it's evident that the model does not provide a good fit to the data.
+"""
+chi_squared_map = (normalized_residual_map) ** 2
+plt.plot(xvalues, chi_squared_map, color="k")
+plt.title("Chi-Squared Map of model-data fit to 1D Gaussian data.")
+plt.xlabel("x values of profile")
+plt.ylabel("Chi-Squareds")
+plt.show()
+plt.clf()
+
+"""
+Now, we consolidate all the information in our `chi_squared_map` into a single measure of goodness-of-fit
+called `chi_squared`.
+
+It is defined as the sum of all values in the `chi_squared_map` and is computed as:
+
+\[ \chi^2 = \sum \left(\frac{\text{data} - \text{model\_data}}{\text{noise\_map}}\right)^2 \]
+
+This summing process highlights why ensuring all values in the chi-squared map are positive is crucial. If we
+didn't square the values (making them positive), positive and negative residuals would cancel each other out,
+leading to an inaccurate assessment of the model's fit to the data.
+"""
+chi_squared = np.sum(chi_squared_map)
+print("Chi-squared = ", chi_squared)
+
+"""
+
+The lower the `chi_squared`, the fewer residuals exist between the model's fit and the data, indicating a better
+overall fit!
+
+__Noise Normalization__
+
+Next, we introduce another quantity that contributes to our final assessment of the goodness-of-fit:
+the `noise_normalization`.
+
+The `noise_normalization` is computed as the logarithm of the sum of squared noise values in our data:
+
+\[
+\text{{noise\_normalization}} = \sum \log(2 \pi \text{{noise\_map}}^2)
+\]
+
+This quantity is fixed because the noise-map remains constant throughout the fitting process. Despite this,
+including the `noise_normalization` is considered good practice due to its statistical significance.
+
+Understanding the exact meaning of `noise_normalization` isn't critical for our primary goal of successfully
+fitting a model to a dataset. Essentially, it provides a measure of how well the noise properties of our data align
+with a Gaussian distribution.
+"""
+noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
+
+"""
+__Likelihood__
+
+From the `chi_squared` and `noise_normalization`, we can define a final goodness-of-fit measure known as
+the `log_likelihood`.
+
+This measure is calculated by taking the sum of the `chi_squared` and `noise_normalization`, and then multiplying the
+result by -0.5:
+
+\[ \text{log\_likelihood} = -0.5 \times \left( \chi^2 + \text{noise\_normalization} \right) \]
+
+Why multiply by -0.5? The exact rationale behind this factor isn't critical for our current understanding.
+"""
+log_likelihood = -0.5 * (chi_squared + noise_normalization)
+print("Log Likelihood = ", log_likelihood)
+
+"""
+Above, we mentioned that a lower `chi_squared` indicates a better fit of the model to the data.
+
+When calculating the `log_likelihood`, we multiply the `chi_squared` by -0.5. Therefore, a higher log likelihood
+corresponds to a better model fit. This is what we aim for when fitting models to data, we want to maximize the
+log likelihood!
+
+__Recap__
+
+If you're familiar with model-fitting, you've likely encountered terms like 'residuals', 'chi-squared',
+and 'log_likelihood' before.
+
+These metrics are standard ways to quantify the quality of a model fit. They are applicable not only to 1D data but
+also to more complex data structures like 2D images, 3D data cubes, or any other multidimensional datasets.
+
+If these terms are new to you, it's important to understand their meanings as they form the basis of all
+model-fitting operations in **PyAutoFit** (and in statistical inference more broadly).
+
+Let's recap what we've learned so far:
+
+- We can define models, such as a 1D `Gaussian`, using Python classes that follow a specific format.
+
+- Models can be organized using `Collection` and `Model` objects, with parameters mapped to instances of their
+ respective model classes (e.g., `Gaussian`).
+
+- Using these model instances, we can generate model data, compare it to observed data, and quantify the
+ goodness-of-fit using the log likelihood.
+
+__Fitting Models__
+
+Now, armed with this knowledge, we are ready to fit our model to our data!
+
+But how do we find the best-fit model, which maximizes the log likelihood?
+
+The simplest approach is to guess parameters. Starting with initial parameter values that yield a good
+fit (i.e., a higher log likelihood), we iteratively adjust these values to refine our model until we achieve an
+optimal fit.
+
+For a 1D `Gaussian`, this iterative process works effectively. Below, we fit three different `Gaussian` models and
+identify the best-fit model—the one that matches the original dataset most closely.
+
+To streamline this process, I've developed functions that compute the `log_likelihood` of a model fit and visualize
+the data alongside the model predictions, complete with error bars.
+"""
+
+
+def log_likelihood_from(
+ data: np.ndarray, noise_map: np.ndarray, model_data: np.ndarray
+) -> float:
+ """
+ Compute the log likelihood of a model fit to data given the noise map.
+
+ Parameters
+ ----------
+ data
+ The observed data.
+ noise_map
+ The root mean square noise (or uncertainty) associated with each data point.
+ model_data
+ The model's predicted data for the given data x points.
+
+ Returns
+ -------
+ float
+ The log likelihood of the model fit to the data.
+ """
+ # Calculate residuals and normalized residuals
+ residual_map = data - model_data
+ normalized_residual_map = residual_map / noise_map
+
+ # Compute chi-squared and noise normalization
+ chi_squared_map = normalized_residual_map**2
+ chi_squared = np.sum(chi_squared_map)
+ noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
+
+ # Compute log likelihood
+ log_likelihood = -0.5 * (chi_squared + noise_normalization)
+
+ return log_likelihood
+
+
+def plot_model_fit(
+ xvalues: np.ndarray,
+ data: np.ndarray,
+ noise_map: np.ndarray,
+ model_data: np.ndarray,
+ color: str = "k",
+):
+ """
+ Plot the observed data, model predictions, and error bars.
+
+ Parameters
+ ----------
+ xvalues
+ The x-axis values where the data is observed and model is predicted.
+ data
+ The observed data points.
+ noise_map
+ The root mean squared noise (or uncertainty) associated with each data point.
+ model_data
+ The model's predicted data for the given data x points.
+ color
+ The color for plotting (default is "k" for black).
+ """
+ plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color=color,
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+ )
+ plt.plot(xvalues, model_data, color="r")
+ plt.title("Fit of model-data to data")
+ plt.xlabel("x values of profile")
+ plt.ylabel("Profile Value")
+ plt.show()
+ plt.clf() # Clear figure to prevent overlapping plots
+
+
+"""
+__Guess 1__
+
+The first guess correctly pinpoints that the Gaussian's peak is at 50.0, but the width and normalization are off.
+
+The `log_likelihood` is computed and printed, however because we don't have a value to compare it to yet, its hard
+to assess if it is a large or small value.
+"""
+
+gaussian = model.instance_from_vector(vector=[50.0, 10.0, 5.0])
+model_data = gaussian.model_data_from(xvalues=xvalues)
+plot_model_fit(
+ xvalues=xvalues,
+ data=data,
+ noise_map=noise_map,
+ model_data=model_data,
+ color="r",
+)
+
+log_likelihood = log_likelihood_from(
+ data=data, noise_map=noise_map, model_data=model_data
+)
+print(f"Log Likelihood: {log_likelihood}")
+
+"""
+__Guess 2__
+
+The second guess refines the width and normalization, but the size of the Gaussian is still off.
+
+The `log_likelihood` is computed and printed, and increases a lot compared to the previous guess, indicating that
+the fit is better.
+"""
+
+gaussian = model.instance_from_vector(vector=[50.0, 25.0, 5.0])
+model_data = gaussian.model_data_from(xvalues=xvalues)
+plot_model_fit(
+ xvalues=xvalues,
+ data=data,
+ noise_map=noise_map,
+ model_data=model_data,
+ color="r",
+)
+
+log_likelihood = log_likelihood_from(
+ data=data, noise_map=noise_map, model_data=model_data
+)
+print(f"Log Likelihood: {log_likelihood}")
+
+"""
+__Guess 3__
+
+The third guess provides a good fit to the data, with the Gaussian's peak, width, and normalization all accurately
+representing the observed signal.
+
+The `log_likelihood` is computed and printed, and is the highest value yet, indicating that this model provides the
+best fit to the data.
+"""
+
+gaussian = model.instance_from_vector(vector=[50.0, 25.0, 10.0])
+model_data = gaussian.model_data_from(xvalues=xvalues)
+plot_model_fit(
+ xvalues=xvalues,
+ data=data,
+ noise_map=noise_map,
+ model_data=model_data,
+ color="r",
+)
+
+log_likelihood = log_likelihood_from(
+ data=data, noise_map=noise_map, model_data=model_data
+)
+print(f"Log Likelihood: {log_likelihood}")
+
+"""
+__Extensibility__
+
+Fitting models composed of multiple components is straightforward with PyAutoFit. Using the `Collection` object,
+we can define complex models consisting of several components. Once defined, we generate `model_data`
+from this collection and fit it to the observed data to compute the log likelihood.
+"""
+model = af.Collection(gaussian_0=Gaussian, gaussian_1=Gaussian)
+
+instance = model.instance_from_vector(vector=[40.0, 0.2, 0.3, 60.0, 0.5, 1.0])
+
+model_data_0 = instance.gaussian_0.model_data_from(xvalues=xvalues)
+model_data_1 = instance.gaussian_1.model_data_from(xvalues=xvalues)
+
+model_data = model_data_0 + model_data_1
+
+"""
+We plot the data and model data below, showing that we get a bad fit (a low log likelihood) for this model.
+
+We could attempt to improve the model-fit and find a higher log likelihood solution by varying the parameters of
+the two Gaussians. However, with 6 parameters, this would be a challenging and cumbersome task to perform by eye.
+"""
+plot_model_fit(
+ xvalues=xvalues,
+ data=data,
+ noise_map=noise_map,
+ model_data=model_data,
+ color="r",
+)
+
+log_likelihood = log_likelihood_from(
+ data=data, noise_map=noise_map, model_data=model_data
+)
+print(f"Log Likelihood: {log_likelihood}")
+
+
+"""
+When our model consisted of only 3 parameters, it was manageable to visually guess their values and achieve a good
+fit to the data. However, as we expanded our model to include six parameters, this approach quickly became
+inefficient. Attempting to manually optimize models with even more parameters would effectively become impossible,
+and a more systematic approach is required.
+
+In the next tutorial, we will introduce an automated approach for fitting models to data. This method will enable
+us to systematically determine the optimal values of model parameters that best describe the observed data, without
+relying on manual guesswork.
+
+__Wrap Up__
+
+To conclude, take a moment to reflect on the model you ultimately aim to fit using **PyAutoFit**. What does your
+data look like? Is it one-dimensional, like a spectrum or a time series? Or is it two-dimensional, such as an image
+or a map? Visualize the nature of your data and consider whether you can define a mathematical model that
+accurately generates similar data.
+
+Can you imagine what a residual map would look like if you were to compare your model's predictions against this
+data? A residual map shows the differences between observed data and the model's predictions, often revealing
+patterns or areas where the model fits well or poorly.
+
+Furthermore, can you foresee how you would calculate a log likelihood from this residual map? The log likelihood q
+uantifies how well your model fits the data, incorporating both the residual values and the noise characteristics of
+your observations.
+
+If you find it challenging to visualize these aspects right now, that's perfectly fine. The first step is to
+grasp the fundamentals of fitting a model to data using **PyAutoFit**, which will provide you with the tools
+and understanding needed to address these questions effectively in the future.
+"""
diff --git a/scripts/howtofit/chapter_1_introduction/tutorial_3_non_linear_search.py b/scripts/howtofit/chapter_1_introduction/tutorial_3_non_linear_search.py
index d72a1564..0a9b8499 100644
--- a/scripts/howtofit/chapter_1_introduction/tutorial_3_non_linear_search.py
+++ b/scripts/howtofit/chapter_1_introduction/tutorial_3_non_linear_search.py
@@ -1,820 +1,816 @@
-"""
-Tutorial 3: Non Linear Search
-=============================
-
-In the previous tutorials, we laid the groundwork by defining a model and manually fitting it to data using fitting
-functions. We quantified the goodness of fit using the log likelihood and demonstrated that for models with only a few
-free parameters, we could achieve satisfactory fits by manually guessing parameter values. However, as the complexity
-of our models increased, this approach quickly became impractical.
-
-In this tutorial, we will delve into a more systematic approach for fitting models to data. This technique is designed
-to handle models with a larger number of parameters—ranging from tens to hundreds. By adopting this approach, we aim
-to achieve more efficient and reliable model fits, ensuring that our models accurately capture the underlying
-structure of the data.
-
-This approach not only improves the accuracy of our fits but also allows us to explore more complex models that better
-represent the systems we are studying.
-
-__Overview__
-
-In this tutorial, we will use a non-linear search to fit a 1D Gaussian profile to noisy data. Specifically, we will:
-
-- Introduce concept like a "parameter space", "likelihood surface" and "priors", and relate them to how a non-linear
- search works.
-
-- Introduce the `Analysis` class, which defines the `log_likelihood_function` that quantifies the goodness of fit of a
- model instance to the data.
-
-- Fit a 1D Gaussian model to 1D data with different non-linear searches, including a maximum likelihood estimator (MLE),
- Markok Chain Monte Carlo (MCMC) and nested sampling.
-
-All these steps utilize **PyAutoFit**'s API for model-fitting.
-
-__Contents__
-
-This tutorial is split into the following sections:
-
-- **Parameter Space**: Introduce the concept of a "parameter space" and how it relates to model-fitting.
-- **Non-Linear Search**: Introduce the concept of a "non-linear search" and how it fits models to data.
-- **Search Types**: Introduce the maximum likelihood estimator (MLE), Markov Chain Monte Carlo (MCMC) and nested sampling search algorithms used in this tutorial.
-- **Deeper Background**: Provide links to resources that more thoroughly describe the statistical principles that underpin non-linear searches.
-- **Data**: Load and plot the 1D Gaussian dataset we'll fit.
-- **Model**: Introduce the 1D `Gaussian` model we'll fit to the data.
-- **Priors**: Introduce priors and how they are used to define the parameter space and guide the non-linear search.
-- **Analysis**: Introduce the `Analysis` class, which contains the `log_likelihood_function` used to fit the model to the data.
-- **Searches**: An overview of the searches used in this tutorial.
-- **Maximum Likelihood Estimation (MLE)**: Perform a model-fit using the MLE search.
-- **Markov Chain Monte Carlo (MCMC)**: Perform a model-fit using the MCMC search.
-- **Nested Sampling**: Perform a model-fit using the nested sampling search.
-- **What is The Best Search To Use?**: Compare the strengths and weaknesses of each search method.
-- **Wrap Up**: A summary of the concepts introduced in this tutorial.
-
-__Parameter Space__
-
-In mathematics, a function is defined by its parameters, which relate inputs to outputs.
-
-For example, consider a simple function:
-
-\[ f(x) = x^2 \]
-
-Here, \( x \) is the parameter input into the function \( f \), and \( f(x) \) returns \( x^2 \). This
-mapping between \( x \) and \( f(x) \) defines the "parameter space" of the function, which in this case is a parabola.
-
-Functions can have multiple parameters, such as \( x \), \( y \), and \( z \):
-
-\[ f(x, y, z) = x + y^2 - z^3 \]
-
-Here, the mapping between \( x \), \( y \), \( z \), and \( f(x, y, z) \) defines a parameter space with three
-dimensions.
-
-This concept of a parameter space relates closely to how we define and use instances of models in model-fitting.
-For instance, in our previous tutorial, we used instances of a 1D Gaussian profile with
-parameters \( (x, I, \sigma) \) to fit data and compute a log likelihood.
-
-This process can be thought of as complete analogous to a function \( f(x, y, z) \), where the output value is the
-log likelihood. This key function, which maps parameter values to a log likelihood, is called the "likelihood function"
-in statistical inference, albeit we will refer to it hereafter as the `log_likelihood_function` to be explicit
-that it is the log of the likelihood function.
-
-By expressing the likelihood in this manner, we can consider our model as having a parameter space -— a
-multidimensional surface that spans all possible values of the model parameters \( x, I, \sigma \).
-
-This surface is often referred to as the "likelihood surface", and our objective during model-fitting is to find
-its peak.
-
-This parameter space is "non-linear", meaning the relationship between the input parameters and the log likelihood
-does not behave linearly. This non-linearity implies that we cannot predict the log likelihood from a set of model
-parameters without actually performing a fit to the data by performing the forward model calculation.
-
-__Non-Linear Search__
-
-Now that we understand our problem in terms of a non-linear parameter space with a likelihood surface, we can
-introduce the method used to fit the model to the data—the "non-linear search".
-
-Previously, our approach involved manually guessing models until finding one with a good fit and high log likelihood.
-Surprisingly, this random guessing forms the basis of how model-fitting using a non-linear search actually works!
-
-A non-linear search involves systematically guessing many models while tracking their log likelihoods. As the
-algorithm progresses, it tends to favor models with parameter combinations that have previously yielded higher
-log likelihoods. This iterative refinement helps to efficiently explore the vast parameter space.
-
-There are two key differences between guessing random models and using a non-linear search:
-
-- **Computational Efficiency**: The non-linear search can evaluate the log likelihood of a model parameter
- combinations in milliseconds and therefore many thousands of models in minutes. This computational speed enables
- it to thoroughly sample potential solutions, which would be impractical for a human.
-
-- **Effective Sampling**: The search algorithm maintains a robust memory of previously guessed models and their log
- likelihoods. This allows it to sample potential solutions more thoroughly and converge on the highest
- likelihood solutions more efficiently, which is again impractical for a human.
-
-Think of the non-linear search as systematically exploring parameter space to pinpoint regions with the highest log
-likelihood values. Its primary goal is to identify and converge on the parameter values that best describe the data.
-
-__Search Types__
-
-There are different types of non-linear searches, each of which explores parameter space in a unique way.
-In this example, we will use three types of searches, which broadly represent the various approaches to non-linear
-searches used in statistical inference.
-
-These are:
-
-- **Maximum Likelihood Estimation (MLE)**: This method aims to find the model that maximizes the likelihood function.
- It does so by testing nearby models and adjusting parameters in the direction that increases the likelihood.
-
-- **Markov Chain Monte Carlo (MCMC)**: This approach uses a group of "walkers" that explore parameter space randomly.
- The likelihood at each walker's position influences the probability of the walker moving to a new position.
-
-- **Nested Sampling**: This technique samples points from the parameter space iteratively. Lower likelihood points
- are replaced by higher likelihood ones, gradually concentrating the samples in regions of high likelihood.
-
-We will provide more details on each of these searches below.
-
-__Deeper Background__
-
-**The descriptions of how searches work in this example are simplfied and phoenomenological and do not give a full
-description of how they work at a deep statistical level. The goal is to provide you with an intuition for how to use
-them and when different searches are appropriate for different problems. Later tutorials will provide a more formal
-description of how these searches work.**
-
-If you're interested in learning more about these principles, you can explore resources such as:
-
-- [Markov Chain Monte Carlo (MCMC)](https://en.wikipedia.org/wiki/Markov_chain_Monte_Carlo)
-- [Introduction to MCMC Sampling](https://twiecki.io/blog/2015/11/10/mcmc-sampling/)
-- [Nested Sampling](https://www.imperial.ac.uk/media/imperial-college/research-centres-and-groups/astrophysics/public/icic/data-analysis-workshop/2016/NestedSampling_JRP.pdf)
-- [A Zero-Math Introduction to MCMC Methods](https://towardsdatascience.com/a-zero-math-introduction-to-markov-chain-monte-carlo-methods-dcba889e0c50)
-"""
-
-import numpy as np
-import matplotlib.pyplot as plt
-from os import path
-
-import autofit as af
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-"""
-__Data__
-
-Load and plot the dataset from the `autofit_workspace/dataset` folder.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-xvalues = np.arange(data.shape[0])
-
-plt.errorbar(
- xvalues,
- data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.title("1D Gaussian dataset.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile Normalization")
-plt.show()
-plt.clf()
-
-"""
-__Model__
-
-Create the `Gaussian` class from which we will compose model components using the standard format.
-"""
-
-
-class Gaussian:
- def __init__(
- self,
- centre: float = 30.0, # <- **PyAutoFit** recognises these constructor arguments
- normalization: float = 1.0, # <- are the Gaussian`s model parameters.
- sigma: float = 5.0,
- ):
- """
- Represents a 1D Gaussian profile.
-
- This is a model-component of example models in the **HowToFit** lectures and is used to perform model-fitting
- of example datasets.
-
- Parameters
- ----------
- centre
- The x coordinate of the profile centre.
- normalization
- Overall normalization of the profile.
- sigma
- The sigma value controlling the size of the Gaussian.
- """
- self.centre = centre
- self.normalization = normalization
- self.sigma = sigma
-
- def model_data_from(self, xvalues: np.ndarray) -> np.ndarray:
- """
- Returns a 1D Gaussian on an input list of Cartesian x coordinates.
-
- The input xvalues are translated to a coordinate system centred on the Gaussian, via its `centre`.
-
- The output is referred to as the `model_data` to signify that it is a representation of the data from the
- model.
-
- Parameters
- ----------
- xvalues
- The x coordinates in the original reference frame of the data.
-
- Returns
- -------
- np.array
- The Gaussian values at the input x coordinates.
- """
- transformed_xvalues = np.subtract(xvalues, self.centre)
- return np.multiply(
- np.divide(self.normalization, self.sigma * np.sqrt(2.0 * np.pi)),
- np.exp(-0.5 * np.square(np.divide(transformed_xvalues, self.sigma))),
- )
-
-
-"""
-We now compose our model, a single 1D Gaussian, which we will fit to the data via the non-linear search.
-"""
-model = af.Model(Gaussian)
-
-print(model.info)
-
-"""
-__Priors__
-
-When we examine the `.info` of our model, we notice that each parameter (like `centre`, `normalization`,
-and `sigma` in our Gaussian model) is associated with priors, such as `UniformPrior`. These priors define the
-range of permissible values that each parameter can assume during the model fitting process.
-
-The priors displayed above use default values defined in the `config/priors` directory. These default values have
-been chosen to be broad, and contain all plausible solutions contained in the simulated 1D Gaussian datasets.
-
-For instance, consider the `centre` parameter of our Gaussian. In theory, it could take on any value from
-negative to positive infinity. However, upon inspecting our dataset, we observe that valid values for `centre`
-fall strictly between 0.0 and 100.0. By using a `UniformPrior` with `lower_limit=0.0` and `upper_limit=100.0`,
-we restrict our parameter space to include only physically plausible values.
-
-Priors serve two primary purposes:
-
-**Defining Valid Parameter Space:** Priors specify the range of parameter values that constitute valid solutions.
-This ensures that our model explores only those solutions that are consistent with our observed data and physical
-constraints.
-
-**Incorporating Prior Knowledge:** Priors also encapsulate our prior beliefs or expectations about the model
-parameters. For instance, if we have previously fitted a similar model to another dataset and obtained certain
-parameter values, we can incorporate this knowledge into our priors for a new dataset. This approach guides the
-model fitting process towards parameter values that are more probable based on our prior understanding.
-
-While we are using `UniformPriors` in this tutorial due to their simplicity, **PyAutoFit** offers various other
-priors like `TruncatedGaussianPrior` and `LogUniformPrior`. These priors are useful for encoding different forms of prior
-information, such as normally distributed values around a mean (`TruncatedGaussianPrior`) or parameters spanning multiple
-orders of magnitude (`LogUniformPrior`).
-"""
-model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.normalization = af.UniformPrior(lower_limit=0.0, upper_limit=10.0)
-model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=10.0)
-
-"""
-__Analysis__
-
-In **PyAutoFit**, the `Analysis` class plays a crucial role in interfacing between the data being fitted and the
-model under consideration. Its primary responsibilities include:
-
-**Receiving Data:** The `Analysis` class is initialized with the data (`data`) and noise map (`noise_map`) that
- the model aims to fit.
-
-**Defining the Log Likelihood Function:** The `Analysis` class defines the `log_likelihood_function`, which
- computes the log likelihood of a model instance given the data. It evaluates how well the model, for a given set of
- parameters, fits the observed data.
-
-**Interface with Non-linear Search:** The `log_likelihood_function` is repeatedly called by the non-linear search
- algorithm to assess the goodness of fit of different parameter combinations. The search algorithm call this function
- many times and maps out regions of parameter space that yield high likelihood solutions.
-
-Below is a suitable `Analysis` class for fitting a 1D gaussian to the data loaded above.
-"""
-
-
-class Analysis(af.Analysis):
- def __init__(self, data: np.ndarray, noise_map: np.ndarray):
- """
- The `Analysis` class acts as an interface between the data and model in **PyAutoFit**.
-
- Its `log_likelihood_function` defines how the model is fitted to the data and it is called many times by
- the non-linear search fitting algorithm.
-
- In this example the `Analysis` `__init__` constructor only contains the `data` and `noise-map`, but it can be
- easily extended to include other quantities.
-
- Parameters
- ----------
- data
- A 1D numpy array containing the data (e.g. a noisy 1D signal) fitted in the workspace examples.
- noise_map
- A 1D numpy array containing the noise values of the data, used for computing the goodness of fit
- metric, the log likelihood.
- """
- super().__init__()
-
- self.data = data
- self.noise_map = noise_map
-
- def log_likelihood_function(self, instance) -> float:
- """
- Returns the log likelihood of a fit of a 1D Gaussian to the dataset.
-
- The `instance` that comes into this method is an instance of the `Gaussian` model above. The parameter values
- are chosen by the non-linear search, based on where it thinks the high likelihood regions of parameter
- space are.
-
- The lines of Python code are commented out below to prevent excessive print statements when we run the
- non-linear search, but feel free to uncomment them and run the search to see the parameters of every instance
- that it fits.
-
- print("Gaussian Instance:")
- print("Centre = ", instance.centre)
- print("Normalization = ", instance.normalization)
- print("Sigma = ", instance.sigma)
-
- The data is fitted using an `instance` of the `Gaussian` class where its `model_data_from`
- is called in order to create a model data representation of the Gaussian that is fitted to the data.
- """
- xvalues = np.arange(self.data.shape[0])
-
- model_data = instance.model_data_from(xvalues=xvalues)
- residual_map = self.data - model_data
- chi_squared_map = (residual_map / self.noise_map) ** 2.0
- chi_squared = sum(chi_squared_map)
- noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
- log_likelihood = -0.5 * (chi_squared + noise_normalization)
-
- return log_likelihood
-
-
-"""
-We create an instance of the `Analysis` class by simply passing it the `data` and `noise_map`:
-"""
-analysis = Analysis(data=data, noise_map=noise_map)
-
-"""
-__Searches__
-
-To perform a non-linear search, we create an instance of a `NonLinearSearch` object. **PyAutoFit** offers many options
-for this. A detailed description of each search method and guidance on when to use them can be found in
-the [search cookbook](https://pyautofit.readthedocs.io/en/latest/cookbooks/search.html).
-
-In this tutorial, we’ll focus on three searches that represent different approaches to model fitting:
-
-1. **Maximum Likelihood Estimation (MLE)** using the `LBFGS` non-linear search.
-2. **Markov Chain Monte Carlo (MCMC)** using the `Emcee` non-linear search.
-3. **Nested Sampling** using the `Dynesty` non-linear search.
-
-In this example, non-linear search results are stored in memory rather and not written to hard disk because the fits
-are fast and can therefore be easily regenerated. The next tutorial will perform fits which write results to the
-hard-disk.
-
-__Maximum Likelihood Estimation (MLE)__
-
-Maximum likelihood estimation (MLE) is the most straightforward type of non-linear search. Here’s a simplified
-overview of how it works:
-
-1. Starts at a point in parameter space with a set of initial values for the model parameters.
-2. Calculates the likelihood of the model at this starting point.
-3. Evaluates the likelihood at nearby points to estimate the gradient, determining the direction in which to move "up" in parameter space.
-4. Moves to a new point where, based on the gradient, the likelihood is higher.
-
-This process repeats until the search finds a point where the likelihood can no longer be improved, indicating that
-the maximum likelihood has been reached.
-
-The `LBFGS` search is an example of an MLE algorithm that follows this iterative procedure. Let’s see how it
-performs on our 1D Gaussian model.
-
-In the example below, we don’t specify a starting point for the MLE, so it begins at the center of the prior
-range for each parameter.
-"""
-search = af.LBFGS()
-
-"""
-To begin the model-fit via the non-linear search, we pass it our model and analysis and begin the fit.
-
-The fit will take a minute or so to run.
-"""
-print(
- """
- The non-linear search has begun running.
- This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
- """
-)
-
-model = af.Model(Gaussian)
-
-result = search.fit(model=model, analysis=analysis)
-
-print("The search has finished run - you may now continue the notebook.")
-
-"""
-Upon completion the non-linear search returns a `Result` object, which contains information about the model-fit.
-
-The `info` attribute shows the result in a readable format.
-
-[Above, we discussed that the `info_whitespace_length` parameter in the config files could b changed to make
-the `model.info` attribute display optimally on your computer. This attribute also controls the whitespace of the
-`result.info` attribute.]
-"""
-print(result.info)
-
-"""
-The result has a "maximum log likelihood instance", which refers to the specific set of model parameters (e.g.,
-for a `Gaussian`) that yielded the highest log likelihood among all models tested by the non-linear search.
-"""
-print("Maximum Likelihood Model:\n")
-max_log_likelihood_instance = result.samples.max_log_likelihood()
-print("Centre = ", max_log_likelihood_instance.centre)
-print("Normalization = ", max_log_likelihood_instance.normalization)
-print("Sigma = ", max_log_likelihood_instance.sigma)
-
-"""
-We can use this to plot the maximum log likelihood fit over the data and determine the quality of fit was inferred:
-"""
-model_data = result.max_log_likelihood_instance.model_data_from(
- xvalues=np.arange(data.shape[0])
-)
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(xvalues, model_data, color="r")
-plt.title("Dynesty model fit to 1D Gaussian dataset.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-The fit quality was poor, and the MLE failed to identify the correct model.
-
-This happened because the starting point of the search was a poor match to the data, placing it far from the true
-solution in parameter space. As a result, after moving "up" the likelihood gradient several times, the search
-settled into a "local maximum," where it couldn't find a better solution.
-
-To achieve a better fit with MLE, the search needs to begin in a region of parameter space where the log likelihood
-is higher. This process is known as "initialization," and it involves providing the search with an
-appropriate "starting point" in parameter space.
-"""
-initializer = af.InitializerParamStartPoints(
- {
- model.centre: 55.0,
- model.normalization: 20.0,
- model.sigma: 8.0,
- }
-)
-
-search = af.LBFGS(initializer=initializer)
-
-print(
- """
- The non-linear search has begun running.
- This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
- """
-)
-
-model = af.Model(Gaussian)
-
-result = search.fit(model=model, analysis=analysis)
-
-print("The search has finished run - you may now continue the notebook.")
-
-"""
-By printing `result.info` and looking at the maximum log likelihood model, we can confirm the search provided a
-good model fit with a much higher likelihood than the incorrect model above.
-"""
-print(result.info)
-
-model_data = result.max_log_likelihood_instance.model_data_from(
- xvalues=np.arange(data.shape[0])
-)
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(xvalues, model_data, color="r")
-plt.title("Dynesty model fit to 1D Gaussian dataset.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-MLE is a great starting point for model-fitting because it’s fast, conceptually simple, and often yields
-accurate results. It is especially effective if you can provide a good initialization, allowing it to find the
-best-fit solution quickly.
-
-However, MLE has its limitations. As seen above, it can get "stuck" in a local maximum, particularly if the
-starting point is poorly chosen. In complex model-fitting problems, providing a suitable starting point can be
-challenging. While MLE performed well in the example with just three parameters, it struggles with models that have
-many parameters, as the complexity of the likelihood surface makes simply moving "up" the gradient less effective.
-
-The MLE also does not provide any information on the errors on the parameters, which is a significant limitation.
-The next two types of searches "map out" the likelihood surface, such that they not only infer the maximum likelihood
-solution but also quantify the errors on the parameters.
-
-__Markov Chain Monte Carlo (MCMC)__
-
-Markov Chain Monte Carlo (MCMC) is a more powerful method for model-fitting, though it is also more computationally
-intensive and conceptually complex. Here’s a simplified overview:
-
-1. Place a set of "walkers" in parameter space, each with random parameter values.
-2. Calculate the likelihood of each walker's position.
-3. Move the walkers to new positions, guided by the likelihood of their current positions. Walkers in high-likelihood
-regions encourage those in lower regions to move closer to them.
-
-This process repeats, with the walkers converging on the highest-likelihood regions of parameter space.
-
-Unlike MLE, MCMC thoroughly explores parameter space. While MLE moves a single point up the likelihood gradient,
-MCMC uses many walkers to explore high-likelihood regions, making it more effective at finding the global maximum,
-though slower.
-
-In the example below, we use the `Emcee` MCMC search to fit the 1D Gaussian model. The search starts with walkers
-initialized in a "ball" around the center of the model’s priors, similar to the MLE search that failed earlier.
-"""
-search = af.Emcee(
- nwalkers=10, # The number of walkers we'll use to sample parameter space.
- nsteps=200, # The number of steps each walker takes, after which 10 * 200 = 2000 steps the non-linear search ends.
-)
-
-print(
- """
- The non-linear search has begun running.
- This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
- """
-)
-
-model = af.Model(Gaussian)
-
-result = search.fit(model=model, analysis=analysis)
-
-print("The search has finished run - you may now continue the notebook.")
-
-print(result.info)
-
-model_data = result.max_log_likelihood_instance.model_data_from(
- xvalues=np.arange(data.shape[0])
-)
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(xvalues, model_data, color="r")
-plt.title("Dynesty model fit to 1D Gaussian dataset.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-The MCMC search succeeded, finding the same high-likelihood model that the MLE search with a good starting point
-identified, even without a good initialization. Its use of multiple walkers exploring parameter space allowed it to
-avoid the local maxima that had trapped the MLE search.
-
-A major advantage of MCMC is that it provides estimates of parameter uncertainties by "mapping out" the likelihood
-surface, unlike MLE, which only finds the maximum likelihood solution. These error estimates are accessible in
-the `result.info` string and through the `result.samples` object, with further details in tutorial 5.
-
-While a good starting point wasn't necessary for this simple model, it becomes essential for efficiently mapping the
-likelihood surface in more complex models with many parameters. The code below shows an MCMC fit using a good starting
-point, with two key differences from the MLE initialization:
-
-1. Instead of single starting values, we provide bounds for each parameter. MCMC initializes each walker in a
-small "ball" in parameter space, requiring a defined range for each parameter from which values are randomly drawn.
-
-2. We do not specify a starting point for the sigma parameter, allowing its initial values to be drawn from its
-priors. This illustrates that with MCMC, it’s not necessary to know a good starting point for every parameter.
-"""
-initializer = af.InitializerParamBounds(
- {
- model.centre: (54.0, 56.0),
- model.normalization: (19.0, 21.0),
- }
-)
-
-search = af.Emcee(
- nwalkers=10, # The number of walkers we'll use to sample parameter space.
- nsteps=200, # The number of steps each walker takes, after which 10 * 200 = 2000 steps the non-linear search ends.
- initializer=initializer,
-)
-
-print(
- """
- The non-linear search has begun running.
- This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
- """
-)
-
-model = af.Model(Gaussian)
-
-result = search.fit(model=model, analysis=analysis)
-
-print("The search has finished run - you may now continue the notebook.")
-
-print(result.info)
-
-"""
-MCMC is a powerful tool for model-fitting, providing accurate parameter estimates and uncertainties. For simple models
-without a starting point, MCMC can still find the correct solution, and if a good starting point is provided, it can
-efficiently scale to more complex models with more parameters.
-
-The main limitation of MCMC is that one has to supply the number of steps the walkers take (`nsteps`). If this value
-is too low, the walkers may not explore the likelihood surface sufficiently. It can be challenging to know the right
-number of steps, especially if models of different complexity are being fitted or if datasets of varying quality are
-used. One often ends up having to perform "trial and error" to verify a sufficient number of steps are used.
-
-MCMC can perform badly in parameter spaces with certain types of complexity, for example when there are
-are local maxima "peaks" the walkers can become stuck walking around them.
-
-__Nested Sampling__
-
-**Nested Sampling** is an advanced method for model-fitting that excels in handling complex models with intricate
-parameter spaces. Here’s a simplified overview of its process:
-
-1. Start with a set of "live points" in parameter space, each initialized with random parameter values drawn from their respective priors.
-
-2. Compute the log likelihood for each live point.
-
-3. Draw a new point based on the likelihood of the current live points, favoring regions of higher likelihood.
-
-4. If the new point has a higher likelihood than any existing live point, it becomes a live point, and the lowest likelihood live point is discarded.
-
-This iterative process continues, gradually focusing the live points around higher likelihood regions of parameter
-space until they converge on the highest likelihood solution.
-
-Like MCMC, Nested Sampling effectively maps out parameter space, providing accurate estimates of parameters and
-their uncertainties.
-"""
-search = af.DynestyStatic(
- sample="rwalk", # This makes dynesty run faster, dont worry about what it means for now!
-)
-
-"""
-To begin the model-fit via the non-linear search, we pass it our model and analysis and begin the fit.
-
-The fit will take a minute or so to run.
-"""
-print(
- """
- The non-linear search has begun running.
- This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
- """
-)
-
-model = af.Model(Gaussian)
-
-result = search.fit(model=model, analysis=analysis)
-
-print("The search has finished run - you may now continue the notebook.")
-
-print(result.info)
-
-model_data = result.max_log_likelihood_instance.model_data_from(
- xvalues=np.arange(data.shape[0])
-)
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(xvalues, model_data, color="r")
-plt.title("Dynesty model fit to 1D Gaussian dataset.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-The **Nested Sampling** search was successful, identifying the same high-likelihood model as the MLE and MCMC searches.
-One of the main benefits of Nested Sampling is its ability to provide accurate parameter estimates and uncertainties,
-similar to MCMC. Additionally, it features a built-in stopping criterion, which eliminates the need for users to
-specify the number of steps the search should take.
-
-This method also excels in handling complex parameter spaces, particularly those with multiple peaks. This is because
-the live points will identify each peak and converge around them, but then begin to be discard from a peak if higher
-likelihood points are found elsewhere in parameter space. In MCMC, the walkers can get stuck indefinitely around a
-peak, causing the method to stall.
-
-Another significant advantage is that Nested Sampling estimates an important statistical quantity
-known as "evidence." This value quantifies how well the model fits the data while considering the model's complexity,
-making it essential for Bayesian model comparison, which will be covered in later tutorials.
-
-Nested sampling cannot use a starting point, as it always samples parameter space from scratch by drawing live points
-from the priors. This is both good and bad, depending on if you have access to a good starting point or not. If you do
-not, your MCMC / MLE fit will likely struggle with initialization compared to Nested Sampling. Conversely, if you do
-possess a robust starting point, it can significantly enhance the performance of MCMC, allowing it to begin closer to
-the highest likelihood regions of parameter space. This proximity can lead to faster convergence and more reliable results.
-
-However, Nested Sampling does have limitations; it often scales poorly with increased model complexity. For example,
-once a model has around 50 or more parameters, Nested Sampling can become very slow, whereas MCMC remains efficient
-even in such complex parameter spaces.
-
-__What is The Best Search To Use?__
-
-The choice of the best search method depends on several factors specific to the problem at hand. Here are key
-considerations that influence which search may be optimal:
-
-Firstly, consider the speed of the fit regardless of the search method. If the fitting process runs efficiently,
-nested sampling could be advantageous for low-dimensional parameter spaces due to its ability to handle complex
-parameter spaces and its built-in stopping criterion. However, in high-dimensional scenarios, MCMC may be more
-suitable, as it scales better with the number of parameters.
-
-Secondly, evaluate whether you have access to a robust starting point for your model fit. A strong initialization can
-make MCMC more appealing, allowing the algorithm to bypass the initial sampling stage and leading to quicker convergence.
-
-Additionally, think about the importance of error estimation in your analysis. If error estimation is not a priority,
-MLE might suffice, but this approach heavily relies on having a solid starting point and may struggle with more complex models.
-
-Ultimately, every model-fitting problem is unique, making it impossible to provide a one-size-fits-all answer regarding
-the best search method. This variability is why **PyAutoFit** offers a diverse array of search options, all
-standardized with a consistent interface. This standardization allows users to experiment with different searches on the
-same model-fitting problem and determine which yields the best results.
-
-Finally, it’s important to note that MLE, MCMC, and nested sampling represent only three categories of non-linear
-searches, each containing various algorithms. Each algorithm has its strengths and weaknesses, so experimenting with
-them can reveal the most effective approach for your specific model-fitting challenge. For further guidance, a detailed
-description of each search method can be found in the [search cookbook](https://pyautofit.readthedocs.io/en/latest/cookbooks/search.html).
-
-__Wrap Up__
-
-This tutorial has laid the foundation with several fundamental concepts in model fitting and statistical inference:
-
-1. **Parameter Space**: This refers to the range of possible values that each parameter in a model can take. It
-defines the dimensions over which the likelihood of different parameter values is evaluated.
-
-2. **Likelihood Surface**: This surface represents how the likelihood of the model varies across the parameter space.
-It helps in identifying the best-fit parameters that maximize the likelihood of the model given the data.
-
-3. **Non-linear Search**: This is an optimization technique used to explore the parameter space and find the
-combination of parameter values that best describe the data. It iteratively adjusts the parameters to maximize the
-likelihood. Many different search algorithms exist, each with their own strengths and weaknesses, and this tutorial
-used the MLE, MCMC, and nested sampling searches.
-
-4. **Priors**: Priors are probabilities assigned to different values of parameters before considering the data.
-They encapsulate our prior knowledge or assumptions about the parameter values. Priors can constrain the parameter
-space, making the search more efficient and realistic.
-
-5. **Model Fitting**: The process of adjusting model parameters to minimize the difference between model predictions
-and observed data, quantified by the likelihood function.
-
-Understanding these concepts is crucial as they form the backbone of model fitting and parameter estimation in
-scientific research and data analysis. In the next tutorials, these concepts will be further expanded upon to
-deepen your understanding and provide more advanced techniques for model fitting and analysis.
-"""
+"""
+Tutorial 3: Non Linear Search
+=============================
+
+In the previous tutorials, we laid the groundwork by defining a model and manually fitting it to data using fitting
+functions. We quantified the goodness of fit using the log likelihood and demonstrated that for models with only a few
+free parameters, we could achieve satisfactory fits by manually guessing parameter values. However, as the complexity
+of our models increased, this approach quickly became impractical.
+
+In this tutorial, we will delve into a more systematic approach for fitting models to data. This technique is designed
+to handle models with a larger number of parameters—ranging from tens to hundreds. By adopting this approach, we aim
+to achieve more efficient and reliable model fits, ensuring that our models accurately capture the underlying
+structure of the data.
+
+This approach not only improves the accuracy of our fits but also allows us to explore more complex models that better
+represent the systems we are studying.
+
+__Overview__
+
+In this tutorial, we will use a non-linear search to fit a 1D Gaussian profile to noisy data. Specifically, we will:
+
+- Introduce concept like a "parameter space", "likelihood surface" and "priors", and relate them to how a non-linear
+ search works.
+
+- Introduce the `Analysis` class, which defines the `log_likelihood_function` that quantifies the goodness of fit of a
+ model instance to the data.
+
+- Fit a 1D Gaussian model to 1D data with different non-linear searches, including a maximum likelihood estimator (MLE),
+ Markok Chain Monte Carlo (MCMC) and nested sampling.
+
+All these steps utilize **PyAutoFit**'s API for model-fitting.
+
+__Contents__
+
+This tutorial is split into the following sections:
+
+- **Parameter Space**: Introduce the concept of a "parameter space" and how it relates to model-fitting.
+- **Non-Linear Search**: Introduce the concept of a "non-linear search" and how it fits models to data.
+- **Search Types**: Introduce the maximum likelihood estimator (MLE), Markov Chain Monte Carlo (MCMC) and nested sampling search algorithms used in this tutorial.
+- **Deeper Background**: Provide links to resources that more thoroughly describe the statistical principles that underpin non-linear searches.
+- **Data**: Load and plot the 1D Gaussian dataset we'll fit.
+- **Model**: Introduce the 1D `Gaussian` model we'll fit to the data.
+- **Priors**: Introduce priors and how they are used to define the parameter space and guide the non-linear search.
+- **Analysis**: Introduce the `Analysis` class, which contains the `log_likelihood_function` used to fit the model to the data.
+- **Searches**: An overview of the searches used in this tutorial.
+- **Maximum Likelihood Estimation (MLE)**: Perform a model-fit using the MLE search.
+- **Markov Chain Monte Carlo (MCMC)**: Perform a model-fit using the MCMC search.
+- **Nested Sampling**: Perform a model-fit using the nested sampling search.
+- **What is The Best Search To Use?**: Compare the strengths and weaknesses of each search method.
+- **Wrap Up**: A summary of the concepts introduced in this tutorial.
+
+__Parameter Space__
+
+In mathematics, a function is defined by its parameters, which relate inputs to outputs.
+
+For example, consider a simple function:
+
+\[ f(x) = x^2 \]
+
+Here, \( x \) is the parameter input into the function \( f \), and \( f(x) \) returns \( x^2 \). This
+mapping between \( x \) and \( f(x) \) defines the "parameter space" of the function, which in this case is a parabola.
+
+Functions can have multiple parameters, such as \( x \), \( y \), and \( z \):
+
+\[ f(x, y, z) = x + y^2 - z^3 \]
+
+Here, the mapping between \( x \), \( y \), \( z \), and \( f(x, y, z) \) defines a parameter space with three
+dimensions.
+
+This concept of a parameter space relates closely to how we define and use instances of models in model-fitting.
+For instance, in our previous tutorial, we used instances of a 1D Gaussian profile with
+parameters \( (x, I, \sigma) \) to fit data and compute a log likelihood.
+
+This process can be thought of as complete analogous to a function \( f(x, y, z) \), where the output value is the
+log likelihood. This key function, which maps parameter values to a log likelihood, is called the "likelihood function"
+in statistical inference, albeit we will refer to it hereafter as the `log_likelihood_function` to be explicit
+that it is the log of the likelihood function.
+
+By expressing the likelihood in this manner, we can consider our model as having a parameter space -— a
+multidimensional surface that spans all possible values of the model parameters \( x, I, \sigma \).
+
+This surface is often referred to as the "likelihood surface", and our objective during model-fitting is to find
+its peak.
+
+This parameter space is "non-linear", meaning the relationship between the input parameters and the log likelihood
+does not behave linearly. This non-linearity implies that we cannot predict the log likelihood from a set of model
+parameters without actually performing a fit to the data by performing the forward model calculation.
+
+__Non-Linear Search__
+
+Now that we understand our problem in terms of a non-linear parameter space with a likelihood surface, we can
+introduce the method used to fit the model to the data—the "non-linear search".
+
+Previously, our approach involved manually guessing models until finding one with a good fit and high log likelihood.
+Surprisingly, this random guessing forms the basis of how model-fitting using a non-linear search actually works!
+
+A non-linear search involves systematically guessing many models while tracking their log likelihoods. As the
+algorithm progresses, it tends to favor models with parameter combinations that have previously yielded higher
+log likelihoods. This iterative refinement helps to efficiently explore the vast parameter space.
+
+There are two key differences between guessing random models and using a non-linear search:
+
+- **Computational Efficiency**: The non-linear search can evaluate the log likelihood of a model parameter
+ combinations in milliseconds and therefore many thousands of models in minutes. This computational speed enables
+ it to thoroughly sample potential solutions, which would be impractical for a human.
+
+- **Effective Sampling**: The search algorithm maintains a robust memory of previously guessed models and their log
+ likelihoods. This allows it to sample potential solutions more thoroughly and converge on the highest
+ likelihood solutions more efficiently, which is again impractical for a human.
+
+Think of the non-linear search as systematically exploring parameter space to pinpoint regions with the highest log
+likelihood values. Its primary goal is to identify and converge on the parameter values that best describe the data.
+
+__Search Types__
+
+There are different types of non-linear searches, each of which explores parameter space in a unique way.
+In this example, we will use three types of searches, which broadly represent the various approaches to non-linear
+searches used in statistical inference.
+
+These are:
+
+- **Maximum Likelihood Estimation (MLE)**: This method aims to find the model that maximizes the likelihood function.
+ It does so by testing nearby models and adjusting parameters in the direction that increases the likelihood.
+
+- **Markov Chain Monte Carlo (MCMC)**: This approach uses a group of "walkers" that explore parameter space randomly.
+ The likelihood at each walker's position influences the probability of the walker moving to a new position.
+
+- **Nested Sampling**: This technique samples points from the parameter space iteratively. Lower likelihood points
+ are replaced by higher likelihood ones, gradually concentrating the samples in regions of high likelihood.
+
+We will provide more details on each of these searches below.
+
+__Deeper Background__
+
+**The descriptions of how searches work in this example are simplfied and phoenomenological and do not give a full
+description of how they work at a deep statistical level. The goal is to provide you with an intuition for how to use
+them and when different searches are appropriate for different problems. Later tutorials will provide a more formal
+description of how these searches work.**
+
+If you're interested in learning more about these principles, you can explore resources such as:
+
+- [Markov Chain Monte Carlo (MCMC)](https://en.wikipedia.org/wiki/Markov_chain_Monte_Carlo)
+- [Introduction to MCMC Sampling](https://twiecki.io/blog/2015/11/10/mcmc-sampling/)
+- [Nested Sampling](https://www.imperial.ac.uk/media/imperial-college/research-centres-and-groups/astrophysics/public/icic/data-analysis-workshop/2016/NestedSampling_JRP.pdf)
+- [A Zero-Math Introduction to MCMC Methods](https://towardsdatascience.com/a-zero-math-introduction-to-markov-chain-monte-carlo-methods-dcba889e0c50)
+"""
+
+import numpy as np
+import matplotlib.pyplot as plt
+from os import path
+
+import autofit as af
+
+# from autoconf import setup_notebook; setup_notebook()
+
+"""
+__Data__
+
+Load and plot the dataset from the `autofit_workspace/dataset` folder.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+xvalues = np.arange(data.shape[0])
+
+plt.errorbar(
+ xvalues,
+ data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.title("1D Gaussian dataset.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile Normalization")
+plt.show()
+plt.clf()
+
+"""
+__Model__
+
+Create the `Gaussian` class from which we will compose model components using the standard format.
+"""
+
+
+class Gaussian:
+ def __init__(
+ self,
+ centre: float = 30.0, # <- **PyAutoFit** recognises these constructor arguments
+ normalization: float = 1.0, # <- are the Gaussian`s model parameters.
+ sigma: float = 5.0,
+ ):
+ """
+ Represents a 1D Gaussian profile.
+
+ This is a model-component of example models in the **HowToFit** lectures and is used to perform model-fitting
+ of example datasets.
+
+ Parameters
+ ----------
+ centre
+ The x coordinate of the profile centre.
+ normalization
+ Overall normalization of the profile.
+ sigma
+ The sigma value controlling the size of the Gaussian.
+ """
+ self.centre = centre
+ self.normalization = normalization
+ self.sigma = sigma
+
+ def model_data_from(self, xvalues: np.ndarray) -> np.ndarray:
+ """
+ Returns a 1D Gaussian on an input list of Cartesian x coordinates.
+
+ The input xvalues are translated to a coordinate system centred on the Gaussian, via its `centre`.
+
+ The output is referred to as the `model_data` to signify that it is a representation of the data from the
+ model.
+
+ Parameters
+ ----------
+ xvalues
+ The x coordinates in the original reference frame of the data.
+
+ Returns
+ -------
+ np.array
+ The Gaussian values at the input x coordinates.
+ """
+ transformed_xvalues = np.subtract(xvalues, self.centre)
+ return np.multiply(
+ np.divide(self.normalization, self.sigma * np.sqrt(2.0 * np.pi)),
+ np.exp(-0.5 * np.square(np.divide(transformed_xvalues, self.sigma))),
+ )
+
+
+"""
+We now compose our model, a single 1D Gaussian, which we will fit to the data via the non-linear search.
+"""
+model = af.Model(Gaussian)
+
+print(model.info)
+
+"""
+__Priors__
+
+When we examine the `.info` of our model, we notice that each parameter (like `centre`, `normalization`,
+and `sigma` in our Gaussian model) is associated with priors, such as `UniformPrior`. These priors define the
+range of permissible values that each parameter can assume during the model fitting process.
+
+The priors displayed above use default values defined in the `config/priors` directory. These default values have
+been chosen to be broad, and contain all plausible solutions contained in the simulated 1D Gaussian datasets.
+
+For instance, consider the `centre` parameter of our Gaussian. In theory, it could take on any value from
+negative to positive infinity. However, upon inspecting our dataset, we observe that valid values for `centre`
+fall strictly between 0.0 and 100.0. By using a `UniformPrior` with `lower_limit=0.0` and `upper_limit=100.0`,
+we restrict our parameter space to include only physically plausible values.
+
+Priors serve two primary purposes:
+
+**Defining Valid Parameter Space:** Priors specify the range of parameter values that constitute valid solutions.
+This ensures that our model explores only those solutions that are consistent with our observed data and physical
+constraints.
+
+**Incorporating Prior Knowledge:** Priors also encapsulate our prior beliefs or expectations about the model
+parameters. For instance, if we have previously fitted a similar model to another dataset and obtained certain
+parameter values, we can incorporate this knowledge into our priors for a new dataset. This approach guides the
+model fitting process towards parameter values that are more probable based on our prior understanding.
+
+While we are using `UniformPriors` in this tutorial due to their simplicity, **PyAutoFit** offers various other
+priors like `TruncatedGaussianPrior` and `LogUniformPrior`. These priors are useful for encoding different forms of prior
+information, such as normally distributed values around a mean (`TruncatedGaussianPrior`) or parameters spanning multiple
+orders of magnitude (`LogUniformPrior`).
+"""
+model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.normalization = af.UniformPrior(lower_limit=0.0, upper_limit=10.0)
+model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=10.0)
+
+"""
+__Analysis__
+
+In **PyAutoFit**, the `Analysis` class plays a crucial role in interfacing between the data being fitted and the
+model under consideration. Its primary responsibilities include:
+
+**Receiving Data:** The `Analysis` class is initialized with the data (`data`) and noise map (`noise_map`) that
+ the model aims to fit.
+
+**Defining the Log Likelihood Function:** The `Analysis` class defines the `log_likelihood_function`, which
+ computes the log likelihood of a model instance given the data. It evaluates how well the model, for a given set of
+ parameters, fits the observed data.
+
+**Interface with Non-linear Search:** The `log_likelihood_function` is repeatedly called by the non-linear search
+ algorithm to assess the goodness of fit of different parameter combinations. The search algorithm call this function
+ many times and maps out regions of parameter space that yield high likelihood solutions.
+
+Below is a suitable `Analysis` class for fitting a 1D gaussian to the data loaded above.
+"""
+
+
+class Analysis(af.Analysis):
+ def __init__(self, data: np.ndarray, noise_map: np.ndarray):
+ """
+ The `Analysis` class acts as an interface between the data and model in **PyAutoFit**.
+
+ Its `log_likelihood_function` defines how the model is fitted to the data and it is called many times by
+ the non-linear search fitting algorithm.
+
+ In this example the `Analysis` `__init__` constructor only contains the `data` and `noise-map`, but it can be
+ easily extended to include other quantities.
+
+ Parameters
+ ----------
+ data
+ A 1D numpy array containing the data (e.g. a noisy 1D signal) fitted in the workspace examples.
+ noise_map
+ A 1D numpy array containing the noise values of the data, used for computing the goodness of fit
+ metric, the log likelihood.
+ """
+ super().__init__()
+
+ self.data = data
+ self.noise_map = noise_map
+
+ def log_likelihood_function(self, instance) -> float:
+ """
+ Returns the log likelihood of a fit of a 1D Gaussian to the dataset.
+
+ The `instance` that comes into this method is an instance of the `Gaussian` model above. The parameter values
+ are chosen by the non-linear search, based on where it thinks the high likelihood regions of parameter
+ space are.
+
+ The lines of Python code are commented out below to prevent excessive print statements when we run the
+ non-linear search, but feel free to uncomment them and run the search to see the parameters of every instance
+ that it fits.
+
+ print("Gaussian Instance:")
+ print("Centre = ", instance.centre)
+ print("Normalization = ", instance.normalization)
+ print("Sigma = ", instance.sigma)
+
+ The data is fitted using an `instance` of the `Gaussian` class where its `model_data_from`
+ is called in order to create a model data representation of the Gaussian that is fitted to the data.
+ """
+ xvalues = np.arange(self.data.shape[0])
+
+ model_data = instance.model_data_from(xvalues=xvalues)
+ residual_map = self.data - model_data
+ chi_squared_map = (residual_map / self.noise_map) ** 2.0
+ chi_squared = sum(chi_squared_map)
+ noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
+ log_likelihood = -0.5 * (chi_squared + noise_normalization)
+
+ return log_likelihood
+
+
+"""
+We create an instance of the `Analysis` class by simply passing it the `data` and `noise_map`:
+"""
+analysis = Analysis(data=data, noise_map=noise_map)
+
+"""
+__Searches__
+
+To perform a non-linear search, we create an instance of a `NonLinearSearch` object. **PyAutoFit** offers many options
+for this. A detailed description of each search method and guidance on when to use them can be found in
+the [search cookbook](https://pyautofit.readthedocs.io/en/latest/cookbooks/search.html).
+
+In this tutorial, we’ll focus on three searches that represent different approaches to model fitting:
+
+1. **Maximum Likelihood Estimation (MLE)** using the `LBFGS` non-linear search.
+2. **Markov Chain Monte Carlo (MCMC)** using the `Emcee` non-linear search.
+3. **Nested Sampling** using the `Dynesty` non-linear search.
+
+In this example, non-linear search results are stored in memory rather and not written to hard disk because the fits
+are fast and can therefore be easily regenerated. The next tutorial will perform fits which write results to the
+hard-disk.
+
+__Maximum Likelihood Estimation (MLE)__
+
+Maximum likelihood estimation (MLE) is the most straightforward type of non-linear search. Here’s a simplified
+overview of how it works:
+
+1. Starts at a point in parameter space with a set of initial values for the model parameters.
+2. Calculates the likelihood of the model at this starting point.
+3. Evaluates the likelihood at nearby points to estimate the gradient, determining the direction in which to move "up" in parameter space.
+4. Moves to a new point where, based on the gradient, the likelihood is higher.
+
+This process repeats until the search finds a point where the likelihood can no longer be improved, indicating that
+the maximum likelihood has been reached.
+
+The `LBFGS` search is an example of an MLE algorithm that follows this iterative procedure. Let’s see how it
+performs on our 1D Gaussian model.
+
+In the example below, we don’t specify a starting point for the MLE, so it begins at the center of the prior
+range for each parameter.
+"""
+search = af.LBFGS()
+
+"""
+To begin the model-fit via the non-linear search, we pass it our model and analysis and begin the fit.
+
+The fit will take a minute or so to run.
+"""
+print(
+ """
+ The non-linear search has begun running.
+ This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
+ """
+)
+
+model = af.Model(Gaussian)
+
+result = search.fit(model=model, analysis=analysis)
+
+print("The search has finished run - you may now continue the notebook.")
+
+"""
+Upon completion the non-linear search returns a `Result` object, which contains information about the model-fit.
+
+The `info` attribute shows the result in a readable format.
+
+[Above, we discussed that the `info_whitespace_length` parameter in the config files could b changed to make
+the `model.info` attribute display optimally on your computer. This attribute also controls the whitespace of the
+`result.info` attribute.]
+"""
+print(result.info)
+
+"""
+The result has a "maximum log likelihood instance", which refers to the specific set of model parameters (e.g.,
+for a `Gaussian`) that yielded the highest log likelihood among all models tested by the non-linear search.
+"""
+print("Maximum Likelihood Model:\n")
+max_log_likelihood_instance = result.samples.max_log_likelihood()
+print("Centre = ", max_log_likelihood_instance.centre)
+print("Normalization = ", max_log_likelihood_instance.normalization)
+print("Sigma = ", max_log_likelihood_instance.sigma)
+
+"""
+We can use this to plot the maximum log likelihood fit over the data and determine the quality of fit was inferred:
+"""
+model_data = result.max_log_likelihood_instance.model_data_from(
+ xvalues=np.arange(data.shape[0])
+)
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(xvalues, model_data, color="r")
+plt.title("Dynesty model fit to 1D Gaussian dataset.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+The fit quality was poor, and the MLE failed to identify the correct model.
+
+This happened because the starting point of the search was a poor match to the data, placing it far from the true
+solution in parameter space. As a result, after moving "up" the likelihood gradient several times, the search
+settled into a "local maximum," where it couldn't find a better solution.
+
+To achieve a better fit with MLE, the search needs to begin in a region of parameter space where the log likelihood
+is higher. This process is known as "initialization," and it involves providing the search with an
+appropriate "starting point" in parameter space.
+"""
+initializer = af.InitializerParamStartPoints(
+ {
+ model.centre: 55.0,
+ model.normalization: 20.0,
+ model.sigma: 8.0,
+ }
+)
+
+search = af.LBFGS(initializer=initializer)
+
+print(
+ """
+ The non-linear search has begun running.
+ This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
+ """
+)
+
+model = af.Model(Gaussian)
+
+result = search.fit(model=model, analysis=analysis)
+
+print("The search has finished run - you may now continue the notebook.")
+
+"""
+By printing `result.info` and looking at the maximum log likelihood model, we can confirm the search provided a
+good model fit with a much higher likelihood than the incorrect model above.
+"""
+print(result.info)
+
+model_data = result.max_log_likelihood_instance.model_data_from(
+ xvalues=np.arange(data.shape[0])
+)
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(xvalues, model_data, color="r")
+plt.title("Dynesty model fit to 1D Gaussian dataset.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+MLE is a great starting point for model-fitting because it’s fast, conceptually simple, and often yields
+accurate results. It is especially effective if you can provide a good initialization, allowing it to find the
+best-fit solution quickly.
+
+However, MLE has its limitations. As seen above, it can get "stuck" in a local maximum, particularly if the
+starting point is poorly chosen. In complex model-fitting problems, providing a suitable starting point can be
+challenging. While MLE performed well in the example with just three parameters, it struggles with models that have
+many parameters, as the complexity of the likelihood surface makes simply moving "up" the gradient less effective.
+
+The MLE also does not provide any information on the errors on the parameters, which is a significant limitation.
+The next two types of searches "map out" the likelihood surface, such that they not only infer the maximum likelihood
+solution but also quantify the errors on the parameters.
+
+__Markov Chain Monte Carlo (MCMC)__
+
+Markov Chain Monte Carlo (MCMC) is a more powerful method for model-fitting, though it is also more computationally
+intensive and conceptually complex. Here’s a simplified overview:
+
+1. Place a set of "walkers" in parameter space, each with random parameter values.
+2. Calculate the likelihood of each walker's position.
+3. Move the walkers to new positions, guided by the likelihood of their current positions. Walkers in high-likelihood
+regions encourage those in lower regions to move closer to them.
+
+This process repeats, with the walkers converging on the highest-likelihood regions of parameter space.
+
+Unlike MLE, MCMC thoroughly explores parameter space. While MLE moves a single point up the likelihood gradient,
+MCMC uses many walkers to explore high-likelihood regions, making it more effective at finding the global maximum,
+though slower.
+
+In the example below, we use the `Emcee` MCMC search to fit the 1D Gaussian model. The search starts with walkers
+initialized in a "ball" around the center of the model’s priors, similar to the MLE search that failed earlier.
+"""
+search = af.Emcee(
+ nwalkers=10, # The number of walkers we'll use to sample parameter space.
+ nsteps=200, # The number of steps each walker takes, after which 10 * 200 = 2000 steps the non-linear search ends.
+)
+
+print(
+ """
+ The non-linear search has begun running.
+ This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
+ """
+)
+
+model = af.Model(Gaussian)
+
+result = search.fit(model=model, analysis=analysis)
+
+print("The search has finished run - you may now continue the notebook.")
+
+print(result.info)
+
+model_data = result.max_log_likelihood_instance.model_data_from(
+ xvalues=np.arange(data.shape[0])
+)
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(xvalues, model_data, color="r")
+plt.title("Dynesty model fit to 1D Gaussian dataset.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+The MCMC search succeeded, finding the same high-likelihood model that the MLE search with a good starting point
+identified, even without a good initialization. Its use of multiple walkers exploring parameter space allowed it to
+avoid the local maxima that had trapped the MLE search.
+
+A major advantage of MCMC is that it provides estimates of parameter uncertainties by "mapping out" the likelihood
+surface, unlike MLE, which only finds the maximum likelihood solution. These error estimates are accessible in
+the `result.info` string and through the `result.samples` object, with further details in tutorial 5.
+
+While a good starting point wasn't necessary for this simple model, it becomes essential for efficiently mapping the
+likelihood surface in more complex models with many parameters. The code below shows an MCMC fit using a good starting
+point, with two key differences from the MLE initialization:
+
+1. Instead of single starting values, we provide bounds for each parameter. MCMC initializes each walker in a
+small "ball" in parameter space, requiring a defined range for each parameter from which values are randomly drawn.
+
+2. We do not specify a starting point for the sigma parameter, allowing its initial values to be drawn from its
+priors. This illustrates that with MCMC, it’s not necessary to know a good starting point for every parameter.
+"""
+initializer = af.InitializerParamBounds(
+ {
+ model.centre: (54.0, 56.0),
+ model.normalization: (19.0, 21.0),
+ }
+)
+
+search = af.Emcee(
+ nwalkers=10, # The number of walkers we'll use to sample parameter space.
+ nsteps=200, # The number of steps each walker takes, after which 10 * 200 = 2000 steps the non-linear search ends.
+ initializer=initializer,
+)
+
+print(
+ """
+ The non-linear search has begun running.
+ This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
+ """
+)
+
+model = af.Model(Gaussian)
+
+result = search.fit(model=model, analysis=analysis)
+
+print("The search has finished run - you may now continue the notebook.")
+
+print(result.info)
+
+"""
+MCMC is a powerful tool for model-fitting, providing accurate parameter estimates and uncertainties. For simple models
+without a starting point, MCMC can still find the correct solution, and if a good starting point is provided, it can
+efficiently scale to more complex models with more parameters.
+
+The main limitation of MCMC is that one has to supply the number of steps the walkers take (`nsteps`). If this value
+is too low, the walkers may not explore the likelihood surface sufficiently. It can be challenging to know the right
+number of steps, especially if models of different complexity are being fitted or if datasets of varying quality are
+used. One often ends up having to perform "trial and error" to verify a sufficient number of steps are used.
+
+MCMC can perform badly in parameter spaces with certain types of complexity, for example when there are
+are local maxima "peaks" the walkers can become stuck walking around them.
+
+__Nested Sampling__
+
+**Nested Sampling** is an advanced method for model-fitting that excels in handling complex models with intricate
+parameter spaces. Here’s a simplified overview of its process:
+
+1. Start with a set of "live points" in parameter space, each initialized with random parameter values drawn from their respective priors.
+
+2. Compute the log likelihood for each live point.
+
+3. Draw a new point based on the likelihood of the current live points, favoring regions of higher likelihood.
+
+4. If the new point has a higher likelihood than any existing live point, it becomes a live point, and the lowest likelihood live point is discarded.
+
+This iterative process continues, gradually focusing the live points around higher likelihood regions of parameter
+space until they converge on the highest likelihood solution.
+
+Like MCMC, Nested Sampling effectively maps out parameter space, providing accurate estimates of parameters and
+their uncertainties.
+"""
+search = af.DynestyStatic(
+ sample="rwalk", # This makes dynesty run faster, dont worry about what it means for now!
+)
+
+"""
+To begin the model-fit via the non-linear search, we pass it our model and analysis and begin the fit.
+
+The fit will take a minute or so to run.
+"""
+print(
+ """
+ The non-linear search has begun running.
+ This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
+ """
+)
+
+model = af.Model(Gaussian)
+
+result = search.fit(model=model, analysis=analysis)
+
+print("The search has finished run - you may now continue the notebook.")
+
+print(result.info)
+
+model_data = result.max_log_likelihood_instance.model_data_from(
+ xvalues=np.arange(data.shape[0])
+)
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(xvalues, model_data, color="r")
+plt.title("Dynesty model fit to 1D Gaussian dataset.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+The **Nested Sampling** search was successful, identifying the same high-likelihood model as the MLE and MCMC searches.
+One of the main benefits of Nested Sampling is its ability to provide accurate parameter estimates and uncertainties,
+similar to MCMC. Additionally, it features a built-in stopping criterion, which eliminates the need for users to
+specify the number of steps the search should take.
+
+This method also excels in handling complex parameter spaces, particularly those with multiple peaks. This is because
+the live points will identify each peak and converge around them, but then begin to be discard from a peak if higher
+likelihood points are found elsewhere in parameter space. In MCMC, the walkers can get stuck indefinitely around a
+peak, causing the method to stall.
+
+Another significant advantage is that Nested Sampling estimates an important statistical quantity
+known as "evidence." This value quantifies how well the model fits the data while considering the model's complexity,
+making it essential for Bayesian model comparison, which will be covered in later tutorials.
+
+Nested sampling cannot use a starting point, as it always samples parameter space from scratch by drawing live points
+from the priors. This is both good and bad, depending on if you have access to a good starting point or not. If you do
+not, your MCMC / MLE fit will likely struggle with initialization compared to Nested Sampling. Conversely, if you do
+possess a robust starting point, it can significantly enhance the performance of MCMC, allowing it to begin closer to
+the highest likelihood regions of parameter space. This proximity can lead to faster convergence and more reliable results.
+
+However, Nested Sampling does have limitations; it often scales poorly with increased model complexity. For example,
+once a model has around 50 or more parameters, Nested Sampling can become very slow, whereas MCMC remains efficient
+even in such complex parameter spaces.
+
+__What is The Best Search To Use?__
+
+The choice of the best search method depends on several factors specific to the problem at hand. Here are key
+considerations that influence which search may be optimal:
+
+Firstly, consider the speed of the fit regardless of the search method. If the fitting process runs efficiently,
+nested sampling could be advantageous for low-dimensional parameter spaces due to its ability to handle complex
+parameter spaces and its built-in stopping criterion. However, in high-dimensional scenarios, MCMC may be more
+suitable, as it scales better with the number of parameters.
+
+Secondly, evaluate whether you have access to a robust starting point for your model fit. A strong initialization can
+make MCMC more appealing, allowing the algorithm to bypass the initial sampling stage and leading to quicker convergence.
+
+Additionally, think about the importance of error estimation in your analysis. If error estimation is not a priority,
+MLE might suffice, but this approach heavily relies on having a solid starting point and may struggle with more complex models.
+
+Ultimately, every model-fitting problem is unique, making it impossible to provide a one-size-fits-all answer regarding
+the best search method. This variability is why **PyAutoFit** offers a diverse array of search options, all
+standardized with a consistent interface. This standardization allows users to experiment with different searches on the
+same model-fitting problem and determine which yields the best results.
+
+Finally, it’s important to note that MLE, MCMC, and nested sampling represent only three categories of non-linear
+searches, each containing various algorithms. Each algorithm has its strengths and weaknesses, so experimenting with
+them can reveal the most effective approach for your specific model-fitting challenge. For further guidance, a detailed
+description of each search method can be found in the [search cookbook](https://pyautofit.readthedocs.io/en/latest/cookbooks/search.html).
+
+__Wrap Up__
+
+This tutorial has laid the foundation with several fundamental concepts in model fitting and statistical inference:
+
+1. **Parameter Space**: This refers to the range of possible values that each parameter in a model can take. It
+defines the dimensions over which the likelihood of different parameter values is evaluated.
+
+2. **Likelihood Surface**: This surface represents how the likelihood of the model varies across the parameter space.
+It helps in identifying the best-fit parameters that maximize the likelihood of the model given the data.
+
+3. **Non-linear Search**: This is an optimization technique used to explore the parameter space and find the
+combination of parameter values that best describe the data. It iteratively adjusts the parameters to maximize the
+likelihood. Many different search algorithms exist, each with their own strengths and weaknesses, and this tutorial
+used the MLE, MCMC, and nested sampling searches.
+
+4. **Priors**: Priors are probabilities assigned to different values of parameters before considering the data.
+They encapsulate our prior knowledge or assumptions about the parameter values. Priors can constrain the parameter
+space, making the search more efficient and realistic.
+
+5. **Model Fitting**: The process of adjusting model parameters to minimize the difference between model predictions
+and observed data, quantified by the likelihood function.
+
+Understanding these concepts is crucial as they form the backbone of model fitting and parameter estimation in
+scientific research and data analysis. In the next tutorials, these concepts will be further expanded upon to
+deepen your understanding and provide more advanced techniques for model fitting and analysis.
+"""
diff --git a/scripts/howtofit/chapter_1_introduction/tutorial_4_why_modeling_is_hard.py b/scripts/howtofit/chapter_1_introduction/tutorial_4_why_modeling_is_hard.py
index 86ce9a2a..fec7a69b 100644
--- a/scripts/howtofit/chapter_1_introduction/tutorial_4_why_modeling_is_hard.py
+++ b/scripts/howtofit/chapter_1_introduction/tutorial_4_why_modeling_is_hard.py
@@ -1,950 +1,946 @@
-"""
-Tutorial 4: Why Modeling Is Hard
-================================
-
-We have successfully fitted a simple 1D Gaussian profile to a dataset using a non-linear search. While achieving an
-accurate model fit has been straightforward, the reality is that model fitting is a challenging problem where many things can go wrong.
-
-This tutorial will illustrate why modeling is challenging, highlight common problems that occur when fitting complex
-models, and show how a good scientific approach can help us overcome these challenges.
-
-We will build on concepts introduced in previous tutorials, such as the non-linear parameter space, likelihood surface,
-and the role of priors.
-
-__Overview__
-
-In this tutorial, we will fit complex models with up to 15 free parameters and consider the following:
-
-- Why more complex models are more difficult to fit and may lead the non-linear search to infer an incorrect solution.
-
-- Strategies for ensuring the non-linear search estimates the correct solution.
-
-- What drives the run-times of a model fit and how to carefully balance run-times with model complexity.
-
-__Contents__
-
-- **Data**: Load and plot the 1D Gaussian dataset we'll fit, which is more complex than the previous tutorial.
-- **Model**: The `Gaussian` model component that we will fit to the data.
-- **Analysis**: The log likelihood function used to fit the model to the data.
-- **Alternative Syntax**: An alternative loop-based approach for creating a summed profile from multiple model components.
-- **Collection**: The `Collection` model used to compose the model-fit.
-- **Search**: Set up the nested sampling search (Dynesty) for the model-fit.
-- **Model Fit**: Perform the model-fit and examine the results.
-- **Result**: Determine if the model-fit was successful and what can be done to ensure a good model-fit.
-- **Why Modeling is Hard**: Introduce the concept of randomness and local maxima and why they make model-fitting challenging.
-- **Prior Tuning**: Adjust the priors of the model to help the non-linear search find the global maxima solution.
-- **Reducing Complexity**: Simplify the model to reduce the dimensionality of the parameter space.
-- **Search More Thoroughly**: Adjust the non-linear search settings to search parameter space more thoroughly.
-- **Summary**: Summarize the three strategies for ensuring successful model-fitting.
-- **Run Times**: Discuss how the likelihood function and complexity of a model impacts the run-time of a model-fit.
-- **Model Mismatch**: Introduce the concept of model mismatches and how it makes inferring the correct model challenging.
-- **Astronomy Example**: How the concepts of this tutorial are applied to real astronomical problems.
-- **Wrap Up**: A summary of the key takeaways of this tutorial.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-from os import path
-import numpy as np
-import matplotlib.pyplot as plt
-
-import autofit as af
-
-"""
-__Data__
-
-Load the dataset we fit.
-
-This is a new `dataset` where the underlying signal is a sum of five `Gaussian` profiles.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x5")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-"""
-Plotting the data reveals that the signal is more complex than a simple 1D Gaussian, as the wings to the left and
-right are more extended than what a single Gaussian profile can account for.
-"""
-xvalues = np.arange(data.shape[0])
-plt.errorbar(
- xvalues,
- data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.title("1D Gaussian dataset with errors from the noise-map.")
-plt.xlabel("x values of profile")
-plt.ylabel("Signal Value")
-plt.show()
-plt.clf()
-plt.close()
-
-"""
-__Model__
-
-Create the `Gaussian` class from which we will compose model components using the standard format.
-"""
-
-
-class Gaussian:
- def __init__(
- self,
- centre: float = 30.0, # <- **PyAutoFit** recognises these constructor arguments
- normalization: float = 1.0, # <- are the Gaussian`s model parameters.
- sigma: float = 5.0,
- ):
- """
- Represents a 1D Gaussian profile.
-
- This is a model-component of example models in the **HowToFit** lectures and is used to perform model-fitting
- of example datasets.
-
- Parameters
- ----------
- centre
- The x coordinate of the profile centre.
- normalization
- Overall normalization of the profile.
- sigma
- The sigma value controlling the size of the Gaussian.
- """
- self.centre = centre
- self.normalization = normalization
- self.sigma = sigma
-
- def model_data_from(self, xvalues: np.ndarray) -> np.ndarray:
- """
- Returns a 1D Gaussian on an input list of Cartesian x coordinates.
-
- The input xvalues are translated to a coordinate system centred on the Gaussian, via its `centre`.
-
- The output is referred to as the `model_data` to signify that it is a representation of the data from the
- model.
-
- Parameters
- ----------
- xvalues
- The x coordinates in the original reference frame of the data.
-
- Returns
- -------
- np.array
- The Gaussian values at the input x coordinates.
- """
- transformed_xvalues = np.subtract(xvalues, self.centre)
- return np.multiply(
- np.divide(self.normalization, self.sigma * np.sqrt(2.0 * np.pi)),
- np.exp(-0.5 * np.square(np.divide(transformed_xvalues, self.sigma))),
- )
-
-
-"""
-__Analysis__
-
-To define the Analysis class for this model-fit, we need to ensure that the `log_likelihood_function` can handle an
-instance containing multiple 1D profiles. Below is an expanded explanation and the corresponding class definition:
-
-The log_likelihood_function will now assume that the instance it receives consists of multiple Gaussian profiles.
-For each Gaussian in the instance, it will compute the model_data and then sum these to create the overall `model_data`
-that is compared to the observed data.
-"""
-
-
-class Analysis(af.Analysis):
- def __init__(self, data: np.ndarray, noise_map: np.ndarray):
- """
- The `Analysis` class acts as an interface between the data and model in **PyAutoFit**.
-
- Its `log_likelihood_function` defines how the model is fitted to the data and it is called many times by
- the non-linear search fitting algorithm.
-
- In this example, the `log_likelihood_function` receives an instance containing multiple instances of
- the `Gaussian` class and sums the `model_data` of each to create the overall model fit to the data.
-
- In this example the `Analysis` `__init__` constructor only contains the `data` and `noise-map`, but it can be
- easily extended to include other quantities.
-
- Parameters
- ----------
- data
- A 1D numpy array containing the data (e.g. a noisy 1D signal) fitted in the workspace examples.
- noise_map
- A 1D numpy array containing the noise values of the data, used for computing the goodness of fit
- metric, the log likelihood.
- """
- super().__init__()
-
- self.data = data
- self.noise_map = noise_map
-
- def log_likelihood_function(self, instance) -> float:
- """
- Returns the log likelihood of a fit of a 1D Gaussian to the dataset.
-
- In the previous tutorial, the instance was a single `Gaussian` profile, however this function now assumes
- the instance contains multiple `Gaussian` profiles.
-
- The `model_data` is therefore the summed `model_data` of all individual Gaussians in the model.
-
- The docstring below describes this in more detail.
-
- Parameters
- ----------
- instance
- A list of 1D profiles with parameters set via the non-linear search.
-
- Returns
- -------
- float
- The log likelihood value indicating how well this model fit the `MaskedDataset`.
- """
-
- """
- In the previous tutorial the instance was a single `Gaussian` profile, meaning we could create the model data
- using the line:
-
- model_data = instance.gaussian.model_data_from(xvalues=self.data.xvalues)
-
- In this tutorial our instance is comprised of three 1D Gaussians, because we will use a `Collection` to
- compose the model:
-
- model = Collection(gaussian_0=Gaussian, gaussian_1=Gaussian, gaussian_2=Gaussian).
-
- By using a Collection, this means the instance parameter input into the fit function is a
- dictionary where individual profiles (and their parameters) can be accessed as followed:
-
- print(instance.gaussian_0)
- print(instance.gaussian_1)
- print(instance.gaussian_2)
-
- print(instance.gaussian_0.centre)
- print(instance.gaussian_1.centre)
- print(instance.gaussian_2.centre)
-
- The `model_data` is therefore the summed `model_data` of all individual Gaussians in the model.
-
- The function `model_data_from_instance` performs this summation.
- """
- model_data = self.model_data_from_instance(instance=instance)
-
- residual_map = self.data - model_data
- chi_squared_map = (residual_map / self.noise_map) ** 2.0
- chi_squared = sum(chi_squared_map)
- noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
- log_likelihood = -0.5 * (chi_squared + noise_normalization)
-
- return log_likelihood
-
- def model_data_from_instance(self, instance):
- """
- To create the summed profile of all individual profiles, we use a list comprehension to iterate over
- all profiles in the instance.
-
- The `instance` has the properties of a Python `iterator` and therefore can be looped over using the standard
- Python for syntax (e.g. `for profile in instance`).
-
- __Alternative Syntax__
-
- For those not familiar with Python list comprehensions, the code below shows how to use the instance to
- create the summed profile using a for loop and numpy array:
-
- model_data = np.zeros(shape=self.data.xvalues.shape[0])
-
- for profile in instance:
- model_data += profile.model_data_from(xvalues=self.data.xvalues)
-
- return model_data
- """
- xvalues = np.arange(self.data.shape[0])
-
- return sum([profile.model_data_from(xvalues=xvalues) for profile in instance])
-
-
-"""
-__Collection__
-
-In the previous tutorial, we fitted a single `Gaussian` profile to the dataset by turning it into a model
-component using the `Model` class.
-
-In this tutorial, we will fit a model composed of five `Gaussian` profiles. To do this, we need to combine
-five `Gaussian` model components into a single model.
-
-This can be achieved using a `Collection` object, which was introduced in tutorial 1. The `Collection` object allows
-us to group together multiple model components—in this case, five `Gaussian` profiles—into one model that can be
-passed to the non-linear search.
-"""
-model = af.Collection(
- gaussian_0=Gaussian,
- gaussian_1=Gaussian,
- gaussian_2=Gaussian,
- gaussian_3=Gaussian,
- gaussian_4=Gaussian,
-)
-
-"""
-The `model.info` confirms the model is composed of 5 `Gaussian` profiles.
-"""
-print(model.info)
-
-"""
-__Search__
-
-We again use the nested sampling algorithm Dynesty to fit the model to the data.
-"""
-search = af.DynestyStatic(
- sample="rwalk", # This makes dynesty run faster, don't worry about what it means for now!
-)
-
-"""
-__Model Fit__
-
-Perform the fit using our five `Gaussian` model, which has 15 free parameters.
-
-This means the non-linear parameter space has a dimensionality of N=15, making it significantly more complex
-than the simpler model we fitted in the previous tutorial.
-
-Consequently, the non-linear search takes slightly longer to run but still completes in under a minute.
-"""
-analysis = Analysis(data=data, noise_map=noise_map)
-
-print(
- """
- The non-linear search has begun running.
- This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
- """
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-print("The search has finished run - you may now continue the notebook.")
-
-"""
-__Result__
-
-The `info` attribute shows the result in a readable format, which contains information on the full collection
-of all 5 model components.
-"""
-print(result.info)
-
-"""
-From the result info, it is hard to assess if the model fit was good or not.
-
-A good way to evaluate the fit is through a visual inspection of the model data plotted over the actual data.
-
-If the model data (red line) consistently aligns with the data points (black error bars), the fit is good.
-However, if the model misses certain features of the data, such as peaks or regions of high intensity,
-the fit was not successful.
-"""
-instance = result.max_log_likelihood_instance
-
-model_data_0 = instance.gaussian_0.model_data_from(xvalues=np.arange(data.shape[0]))
-model_data_1 = instance.gaussian_1.model_data_from(xvalues=np.arange(data.shape[0]))
-model_data_2 = instance.gaussian_2.model_data_from(xvalues=np.arange(data.shape[0]))
-model_data_3 = instance.gaussian_3.model_data_from(xvalues=np.arange(data.shape[0]))
-model_data_4 = instance.gaussian_4.model_data_from(xvalues=np.arange(data.shape[0]))
-
-model_data_list = [model_data_0, model_data_1, model_data_2, model_data_3, model_data_4]
-
-model_data = sum(model_data_list)
-
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(range(data.shape[0]), model_data, color="r")
-for model_data_1d_individual in model_data_list:
- plt.plot(range(data.shape[0]), model_data_1d_individual, "--")
-plt.title(f"Fit (log likelihood = {result.log_likelihood})")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-It's challenging to determine from the plot whether the data and model data perfectly overlap across the entire dataset.
-
-To clarify this, the residual map introduced in tutorial 2 is useful. It provides a clear representation of where
-the differences between the model and data exceed the noise level.
-
-Regions where the black error bars do not align with the zero line in the residual map indicate areas where the model
-did not fit the data well and is inconsistent with the data above the noise level. Furthermore, regions where
-larger values of residuals are next to one another indicate that the model failed to accurate fit that
-region of the data.
-"""
-residual_map = data - model_data
-plt.plot(range(data.shape[0]), np.zeros(data.shape[0]), "--", color="b")
-plt.errorbar(
- x=xvalues,
- y=residual_map,
- yerr=noise_map,
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
- linestyle="",
-)
-plt.title(f"Residuals (log likelihood = {result.log_likelihood})")
-plt.xlabel("x values of profile")
-plt.ylabel("Residuals")
-plt.show()
-plt.clf()
-plt.close()
-
-"""
-The normalized residual map, as discussed in tutorial 2, provides an alternative visualization of the fit quality.
-
-Normalized residuals indicate the standard deviation (σ) level at which the residuals could have been drawn from the
-noise. For instance, a normalized residual of 2.0 suggests that a residual value is 2.0σ away from the noise,
-implying there is a 5% chance such a residual would occur due to noise.
-
-Values of normalized residuals above 3.0 are particularly improbable (occurring only 0.3% of the time), which is
-generally considered a threshold where issues with the model-fit are likely the cause of the residual as opposed
-to it being a noise fluctuation.
-"""
-residual_map = data - model_data
-normalized_residual_map = residual_map / noise_map
-plt.plot(xvalues, normalized_residual_map, color="k")
-plt.title(f"Normalized Residuals (log likelihood = {result.log_likelihood})")
-plt.xlabel("x values of profile")
-plt.ylabel("Normalized Residuals ($\sigma$)")
-plt.show()
-plt.clf()
-plt.close()
-
-"""
-So, did you achieve a good fit? Maybe a bad one? Or just an okay one?
-
-The truth is, I don't know, and I can't tell you for sure. Modeling is inherently random. It's not uncommon to
-fit the same model to the same dataset using the same non-linear search and get a different result each time.
-
-When I ran the model fit above, that's exactly what happened. It produced a range of fits: some bad, some okay, and
-some good, as shown in the images below:
-
-
-
-Distinguishing between the good and okay fit is difficult, however the normalized residuals make this easier. They show
-that for the okay fit there are residuals above 3.0 sigma, indicating that the model did not perfectly fit the data.
-
-
-
-You should quickly rerun the code above a couple of times to see this variability for yourself.
-
-__Why Modeling is Hard__
-
-This variability is at the heart of why modeling is challenging. The process of model-fitting is stochastic,
-meaning it's hard as the scientist to determine if a better fit is possible or not.
-
-Why does modeling produce different results each time, and why might it sometimes infer solutions that are not good fits?
-
-In the previous tutorial, the non-linear search consistently found models that visually matched the data well,
-minimizing residuals and yielding high log likelihood values. These optimal solutions are called 'global maxima',
-they are where the model parameters correspond to the highest likelihood regions across the entire parameter space.
-This ideal scenario is illustrated in the `good_fit.png` image above.
-
-However, non-linear searches do not always find these global maxima. Instead, they might settle on 'local maxima'
-solutions, which have high log likelihood values relative to nearby models in parameter space but are significantly
-lower than the true global maxima found elsewhere.
-
-This is what happened for the okay and bad fits above. The non-linear search converged on solutions that were locally
-peaks on the likelihood surface but were not the global maximum solution. This is why the residuals were higher and
-the normalized residuals above 3.0 sigma.
-
-Why does a non-linear search end up at local maxima? As discussed, the search iterates through many models,
-focusing more on regions where previous guesses yielded higher likelihoods. It gradually converges around
-solutions with higher likelihoods compared to surrounding models. If the search isn't exhaustive enough, it might
-converge on a local maxima that appears good compared to nearby models but isn't the global maximum.
-
-Modeling is challenging because the parameter spaces of complex models are typically filled with local maxima,
-making it hard for a non-linear search to locate the global maximum.
-
-Fortunately, there are strategies to help non-linear searches find the global maxima, and we'll now explore three of
-them.
-
-__Prior Tuning__
-
-First, let's assist our non-linear search by tuning our priors. Priors provide guidance to the search on where to
-explore in the parameter space. By setting more accurate priors ('tuning' them), we can help the search find the
-global solution instead of settling for a local maximum.
-
-For instance, from the data itself, it's evident that all `Gaussian` profiles are centered around pixel 50. In our
-previous fit, the `centre` parameter of each `Gaussian` had a `UniformPrior` spanning from 0.0 to 100.0, which is
-much broader than necessary given the data's range.
-
-Additionally, the peak value of the data's `normalization` parameter was around 17.5. This indicates that
-the `normalization` values of our `Gaussians` do not exceed 500.0, allowing us to refine our prior accordingly.
-
-The following code snippet adjusts these priors for the `centre` and `normalization` parameters of
-each `Gaussian` using **PyAutoFit**'s API for model and prior customization:
-"""
-gaussian_0 = af.Model(Gaussian)
-
-gaussian_0.centre = af.UniformPrior(lower_limit=45.0, upper_limit=55.0)
-gaussian_0.normalization = af.LogUniformPrior(lower_limit=0.1, upper_limit=500.0)
-
-gaussian_1 = af.Model(Gaussian)
-
-gaussian_1.centre = af.UniformPrior(lower_limit=45.0, upper_limit=55.0)
-gaussian_1.normalization = af.LogUniformPrior(lower_limit=0.1, upper_limit=500.0)
-
-gaussian_2 = af.Model(Gaussian)
-
-gaussian_2.centre = af.UniformPrior(lower_limit=45.0, upper_limit=55.0)
-gaussian_2.normalization = af.LogUniformPrior(lower_limit=0.1, upper_limit=500.0)
-
-gaussian_3 = af.Model(Gaussian)
-
-gaussian_3.centre = af.UniformPrior(lower_limit=45.0, upper_limit=55.0)
-gaussian_3.normalization = af.LogUniformPrior(lower_limit=0.1, upper_limit=500.0)
-
-gaussian_4 = af.Model(Gaussian)
-
-gaussian_4.centre = af.UniformPrior(lower_limit=45.0, upper_limit=55.0)
-gaussian_4.normalization = af.LogUniformPrior(lower_limit=0.1, upper_limit=500.0)
-
-model = af.Collection(
- gaussian_0=gaussian_0,
- gaussian_1=gaussian_1,
- gaussian_2=gaussian_2,
- gaussian_3=gaussian_3,
- gaussian_4=gaussian_4,
-)
-
-"""
-The `info` attribute shows the model is now using the priors specified above.
-"""
-print(model.info)
-
-"""
-We now repeat the model-fit using these updated priors.
-
-First, you should note that the run time of the fit is significantly faster than the previous fit. This is because
-the prior is telling the non-linear search where to look, meaning it converges on solutions more quickly
-and spends less time searching regions of parameter space that do not contain solutions.
-
-Second, the model-fit consistently produces a good model-fit more often, because our tuned priors are centred
-on the global maxima solution ensuring the non-linear search is less likely to converge on a local maxima.
-"""
-print(
- """
- The non-linear search has begun running.
- This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
- """
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-print("The search has finished run - you may now continue the notebook.")
-
-"""
-Lets print the result info and plot the fit to the dataset to confirm the priors have provided a better model-fit.
-"""
-print(result.info)
-
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
- linestyle="",
-)
-plt.plot(range(data.shape[0]), model_data, color="r")
-for model_data_1d_individual in model_data_list:
- plt.plot(range(data.shape[0]), model_data_1d_individual, "--")
-plt.title(f"Fit (log likelihood = {result.log_likelihood})")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.clf()
-plt.close()
-
-residual_map = data - model_data
-normalized_residual_map = residual_map / noise_map
-plt.plot(xvalues, normalized_residual_map, color="k")
-plt.title(f"Normalized Residuals (log likelihood = {result.log_likelihood})")
-plt.xlabel("x values of profile")
-plt.ylabel("Normalized Residuals ($\sigma$)")
-plt.show()
-plt.clf()
-plt.close()
-
-"""
-Lets consider the advantages and disadvantages of prior tuning:
-
-**Advantages:**
-
-- Higher likelihood of finding the global maximum log likelihood solutions in parameter space.
-
-- Faster search times, as the non-linear search explores less of the parameter space.
-
-**Disadvantages:**
-
-- Incorrectly specified priors could lead the non-linear search to an incorrect solution.
-
-- It is not always clear how the priors should be tuned, especially for complex models.
-
-- Priors tuning must be applied to each dataset fitted. For large datasets, this process would be very time-consuming.
-
-__Reducing Complexity__
-
-The non-linear search may fail because the model is too complex, making its parameter space too difficult to
-sample accurately consistent. To address this, we may be able to simplify the model while ensuring it remains
-realistic enough for our scientific study. By making certain assumptions, we can reduce the number of model
-parameters, thereby lowering the dimensionality of the parameter space and improving the search's performance.
-
-For example, we may know that the `Gaussian`'s in our model are aligned at the same `centre`. We can therefore
-compose a model that assumes that the `centre` of each `Gaussian` is the same, reducing the dimensionality of the
-model from N=15 to N=11.
-
-The code below shows how we can customize the model components to ensure the `centre` of each `Gaussian` is the same:
-"""
-gaussian_0 = af.Model(Gaussian)
-gaussian_1 = af.Model(Gaussian)
-gaussian_2 = af.Model(Gaussian)
-gaussian_3 = af.Model(Gaussian)
-gaussian_4 = af.Model(Gaussian)
-
-gaussian_1.centre = gaussian_0.centre
-gaussian_2.centre = gaussian_0.centre
-gaussian_3.centre = gaussian_0.centre
-gaussian_4.centre = gaussian_0.centre
-
-model = af.Collection(
- gaussian_0=gaussian_0,
- gaussian_1=gaussian_1,
- gaussian_2=gaussian_2,
- gaussian_3=gaussian_3,
- gaussian_4=gaussian_4,
-)
-
-"""
-The `info` attribute shows the model is now using the same `centre` for all `Gaussian`'s and has 11 free parameters.
-"""
-print(model.info)
-
-"""
-We now repeat the model-fit using this updated model where the `centre` of each `Gaussian` is the same.
-
-You should again note that the run time of the fit is significantly faster than the previous fits
-and that it consistently produces a good model-fit more often. This is because the model is less complex,
-non-linear parameter space is less difficult to sample and the search is less likely to converge on a local maxima.
-"""
-print(
- """
- The non-linear search has begun running.
- This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
- """
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-print("The search has finished run - you may now continue the notebook.")
-
-"""
-Lets print the result info and plot the fit to the dataset to confirm the reduced model complexity has
-provided a better model-fit.
-"""
-print(result.info)
-
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
- linestyle="",
-)
-plt.plot(range(data.shape[0]), model_data, color="r")
-for model_data_1d_individual in model_data_list:
- plt.plot(range(data.shape[0]), model_data_1d_individual, "--")
-plt.title(f"Fit (log likelihood = {result.log_likelihood})")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.clf()
-plt.close()
-
-residual_map = data - model_data
-normalized_residual_map = residual_map / noise_map
-plt.plot(xvalues, normalized_residual_map, color="k")
-plt.title(f"Normalized Residuals (log likelihood = {result.log_likelihood})")
-plt.xlabel("x values of profile")
-plt.ylabel("Normalized Residuals ($\sigma$)")
-plt.show()
-plt.clf()
-plt.close()
-
-"""
-Let’s consider the advantages and disadvantages of simplifying the model:
-
-Advantages:
-
-- By reducing the complexity of the parameter space, we increase the chances of finding the global maximum log
-likelihood, and the search requires less time to do so.
-
-- Unlike with tuned priors, this approach is not specific to a single dataset, allowing us to use it on many datasets.
-
-Disadvantages:
-
-- Our model is less realistic, which may negatively impact the accuracy of our fit and the scientific results we
-derive from it.
-
-__Search More Thoroughly__
-
-In approaches 1 and 2, we assisted our non-linear search to find the highest log likelihood regions of parameter
-space. In approach 3, we're simply going to tell the search to look more thoroughly through parameter space.
-
-Every non-linear search has settings that control how thoroughly it explores parameter space. For Dynesty, the
-primary setting is the number of live points `nlive`. The more thoroughly the search examines the space, the more
-likely it is to find the global maximum model. However, this also means the search will take longer!
-
-Below, we configure a more thorough Dynesty search with `nlive=500`. It is currently unclear what changing
-this setting actually does and what the number of live points actually means. These will be covered in chapter 2
-of the **HowToFit** lectures, where we will also expand on how a non-linear search actually works and the different
-types of methods that can be used to search parameter space.
-"""
-model = af.Collection(
- gaussian_0=Gaussian,
- gaussian_1=Gaussian,
- gaussian_2=Gaussian,
- gaussian_3=Gaussian,
- gaussian_4=Gaussian,
-)
-
-"""
-The `model.info` confirms the model is the same model fitted initially, composed of 5 `Gaussian` profiles.
-"""
-print(model.info)
-
-"""
-__Search__
-
-We again use the nested sampling algorithm Dynesty to fit the model to the data, but now increase the number of live
-points to 300 meaning it will search parameter space more thoroughly.
-"""
-search = af.DynestyStatic(
- nlive=300,
- sample="rwalk", # This makes dynesty run faster, don't worry about what it means for now!
-)
-
-"""
-__Model Fit__
-
-Perform the fit using our five `Gaussian` model, which has 15 free parameters.
-"""
-analysis = Analysis(data=data, noise_map=noise_map)
-
-print(
- """
- The non-linear search has begun running.
- This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
- """
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-print("The search has finished run - you may now continue the notebook.")
-
-
-"""
-Lets print the result info and plot the fit to the dataset to confirm the more thorough search has provided a better
-model-fit.
-"""
-print(result.info)
-
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
- linestyle="",
-)
-plt.plot(range(data.shape[0]), model_data, color="r")
-for model_data_1d_individual in model_data_list:
- plt.plot(range(data.shape[0]), model_data_1d_individual, "--")
-plt.title(f"Fit (log likelihood = {result.log_likelihood})")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.clf()
-plt.close()
-
-residual_map = data - model_data
-normalized_residual_map = residual_map / noise_map
-plt.plot(xvalues, normalized_residual_map, color="k")
-plt.title(f"Normalized Residuals (log likelihood = {result.log_likelihood})")
-plt.xlabel("x values of profile")
-plt.ylabel("Normalized Residuals ($\sigma$)")
-plt.show()
-plt.clf()
-plt.close()
-
-"""
-If you repeat the fit multiple times, you will find that the model-fit is more likely to produce a good fit than
-previously.
-
-However, the run-time of the search is noticeably longer, taking a few minutes to complete, owining
-to the increased number of live points and fact it is searching parameter space more thoroughly.
-
-Let's list the advantages and disadvantages of simply adjusting the non-linear search:
-
-**Advantages:**
-
-- It’s easy to set up; just change the settings of the non-linear search.
-
-- It generalizes to any dataset.
-
-- We can retain a more complex model.
-
-**Disadvantage:**
-
-- It can be very expensive in terms of run time, producing run-times that are five, tens or even longer than the
- original run-time.
-
-__Summary__
-
-We have covered three strategies for ensuring the non-linear search estimates the correct solution:
-
-1) Prior Tuning: By setting our priors more accurately, we can help the search find the global solution instead of
- getting stuck at a local maxima.
-
-2) Reducing Complexity: By making certain assumptions, we can reduce the number of model parameters, thereby lowering
- the dimensionality of the parameter space and improving the search's performance.
-
-3) Search More Thoroughly: By increasing the number of live points, we can make the search explore parameter space
- more thoroughly, increasing the likelihood of finding the global maximum model.
-
-Each of these strategies has its advantages and disadvantages, and your ability to fit models successfully will
-likely rely on a combination of these strategies. Which one works best depends on the specific model, dataset and
-scientific question you are trying to answer. Therefore, when you begin your own model-fitting, it is a good idea to
-try each of these strategies and assess which works best for your model-fit.
-
-__Run Times__
-
-One challenging aspect of model-fitting which was not properly covered in this tutorial is the run-time of a model-fit.
-This example fits simple 1D datasets, which are computationally inexpensive to fit. That is, the `log_likelihood_function`
-is evaluated in a fraction of a second, meaning the non-linear search fitted the model in mere minutes.
-
-Many model-fitting tasks are not as fast. For example, when fitting a model to a 2D image, the `log_likelihood_function`
-may take of order seconds, or longer, because it comprises a number of expensive calculations (e.g. a Fourier transform,
-2D convolution, etc.). Depending on the model complexity, this means that the non-linear search may take hours, days
-or even weeks to fit the model.
-
-Run times are also dictated by the complexity of the model and the nature of the log likelihood function. For models
-with many more dimensions than the simple 1D model used in this tutorial (e.g. hundreds or thousands of free parameters),
-non-linear search may take tens or hundreds of more iterations to converge on a solution. This is because the parameter
-space is significantly more complex and difficult to sample accurately. More iterations mean longer run times,
-which in combination with a slow likelihood function can make model-fitting infeasible.
-
-Whether or not run times will pose a challenge to your model-fitting task depends on the complexity of the model and
-nature of the log likelihood function. If your problem is computationally expensive, **PyAutoFit** provides many
-tools to help, which will be the topic of tutorials in chapter 2 of the **HowToFit** lectures.
-
-**PyAutoFit** provides tools to profile the run-time of your log likelihood function, which can be used to
-assess the computational expense of your model-fit and plan accordingly. Below is an example of the simplest use
-of these tools, an estimate of the run-time of the log likelihood function using one random instance of the model.
-
-
-Feature still being developed, IGNORE FOR NOW.
-
-run_time_dict, info_dict = analysis.profile_log_likelihood_function(
- instance=model.random_instance()
-)
-print(f"Log Likelihood Evaluation Time (second) = {run_time_dict['fit_time']}")
-
-__Model Mismatch__
-
-In this example, interpreting how well the model fitted the data, and whether it found the global maxima, was
-relatively straightforward. This is because the same model was used to simulate the data and fit it, meaning the
-global maxima fit corresponded to one where the normalized residuals were minimized and consistent with the noise
-(e.g. they went to sigma values below 3.0 or so).
-
-In many scientific studies, the data that one is fitting may have come from an instrument or simulation where the
-exact physical processes that generate the data are not perfectly known. This then means that the model is
-not a perfect representation of the data, and it may not ever be possible to fit the data perfectly. In this case,
-we might infer a fit with significant residuals, but it may still correspond to the global maxima solution,
-at least for that particular model.
-
-This makes it even more difficult to be certain if the non-linear search is sampling parameter space correctly,
-and therefore requires even more care and attention to the strategies we have discussed above.
-
-Whether or not this is the case for your model-fitting task is something you will have to determine yourself.
-**PyAutoFit** provides many tools to help assess the quality of a model-fit, which will be the topic of tutorials
-in chapter 2 of the **HowToFit** lectures.
-
-__Astronomy Example__
-
-At the end of chapter 1, we will fit a complex model to a real astronomical dataset in order to quantify
-the distribution of stars in 2D images of galaxies.
-
-This example will illustrate many of the challenges discussed in this tutorial, including:
-
-- Fits using more complex models consisting of 15-20 parameters often infer local maxima, unless we assist the
- non-linear search with tuned priors, reduced complexity or a more thorough search.
-
-- Fitting 2D imaging data requires a 2D convolution, which is somewhat computationally expensive and means run times
- become something we must balance with model complexity.
-
-- The model is not a perfect representation of the data. For example, the model assumes the galaxy is elliptically
- symmetric, whereas the real galaxy may not be. In certain examples, this means that the global maxima solution
- actually leaves significant residuals, above 3.0 $\sigma$, in the data.
-
-__Wrap Up__
-
-Now is a good time to assess how straightforward or difficult you think your model-fitting task will be.
-
-Are the models you will be fitting made up of tens of parameters? or thousands? Are there ways you can simplify
-the model parameter or tune priors to make the model-fitting task more feasible? Will run times be an issue, or is
-your likelihood function computationally cheap? And how confident are you that the model you are fitting is a good
-representation of the data?
-
-These are all questions you should be asking yourself before beginning your model-fitting task, but they will
-become easier to answer as you gain experience with model-fitting and **PyAutoFit**.
-"""
+"""
+Tutorial 4: Why Modeling Is Hard
+================================
+
+We have successfully fitted a simple 1D Gaussian profile to a dataset using a non-linear search. While achieving an
+accurate model fit has been straightforward, the reality is that model fitting is a challenging problem where many things can go wrong.
+
+This tutorial will illustrate why modeling is challenging, highlight common problems that occur when fitting complex
+models, and show how a good scientific approach can help us overcome these challenges.
+
+We will build on concepts introduced in previous tutorials, such as the non-linear parameter space, likelihood surface,
+and the role of priors.
+
+__Overview__
+
+In this tutorial, we will fit complex models with up to 15 free parameters and consider the following:
+
+- Why more complex models are more difficult to fit and may lead the non-linear search to infer an incorrect solution.
+
+- Strategies for ensuring the non-linear search estimates the correct solution.
+
+- What drives the run-times of a model fit and how to carefully balance run-times with model complexity.
+
+__Contents__
+
+- **Data**: Load and plot the 1D Gaussian dataset we'll fit, which is more complex than the previous tutorial.
+- **Model**: The `Gaussian` model component that we will fit to the data.
+- **Analysis**: The log likelihood function used to fit the model to the data.
+- **Alternative Syntax**: An alternative loop-based approach for creating a summed profile from multiple model components.
+- **Collection**: The `Collection` model used to compose the model-fit.
+- **Search**: Set up the nested sampling search (Dynesty) for the model-fit.
+- **Model Fit**: Perform the model-fit and examine the results.
+- **Result**: Determine if the model-fit was successful and what can be done to ensure a good model-fit.
+- **Why Modeling is Hard**: Introduce the concept of randomness and local maxima and why they make model-fitting challenging.
+- **Prior Tuning**: Adjust the priors of the model to help the non-linear search find the global maxima solution.
+- **Reducing Complexity**: Simplify the model to reduce the dimensionality of the parameter space.
+- **Search More Thoroughly**: Adjust the non-linear search settings to search parameter space more thoroughly.
+- **Summary**: Summarize the three strategies for ensuring successful model-fitting.
+- **Run Times**: Discuss how the likelihood function and complexity of a model impacts the run-time of a model-fit.
+- **Model Mismatch**: Introduce the concept of model mismatches and how it makes inferring the correct model challenging.
+- **Astronomy Example**: How the concepts of this tutorial are applied to real astronomical problems.
+- **Wrap Up**: A summary of the key takeaways of this tutorial.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+from os import path
+import numpy as np
+import matplotlib.pyplot as plt
+
+import autofit as af
+
+"""
+__Data__
+
+Load the dataset we fit.
+
+This is a new `dataset` where the underlying signal is a sum of five `Gaussian` profiles.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x5")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+"""
+Plotting the data reveals that the signal is more complex than a simple 1D Gaussian, as the wings to the left and
+right are more extended than what a single Gaussian profile can account for.
+"""
+xvalues = np.arange(data.shape[0])
+plt.errorbar(
+ xvalues,
+ data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.title("1D Gaussian dataset with errors from the noise-map.")
+plt.xlabel("x values of profile")
+plt.ylabel("Signal Value")
+plt.show()
+plt.clf()
+plt.close()
+
+"""
+__Model__
+
+Create the `Gaussian` class from which we will compose model components using the standard format.
+"""
+
+
+class Gaussian:
+ def __init__(
+ self,
+ centre: float = 30.0, # <- **PyAutoFit** recognises these constructor arguments
+ normalization: float = 1.0, # <- are the Gaussian`s model parameters.
+ sigma: float = 5.0,
+ ):
+ """
+ Represents a 1D Gaussian profile.
+
+ This is a model-component of example models in the **HowToFit** lectures and is used to perform model-fitting
+ of example datasets.
+
+ Parameters
+ ----------
+ centre
+ The x coordinate of the profile centre.
+ normalization
+ Overall normalization of the profile.
+ sigma
+ The sigma value controlling the size of the Gaussian.
+ """
+ self.centre = centre
+ self.normalization = normalization
+ self.sigma = sigma
+
+ def model_data_from(self, xvalues: np.ndarray) -> np.ndarray:
+ """
+ Returns a 1D Gaussian on an input list of Cartesian x coordinates.
+
+ The input xvalues are translated to a coordinate system centred on the Gaussian, via its `centre`.
+
+ The output is referred to as the `model_data` to signify that it is a representation of the data from the
+ model.
+
+ Parameters
+ ----------
+ xvalues
+ The x coordinates in the original reference frame of the data.
+
+ Returns
+ -------
+ np.array
+ The Gaussian values at the input x coordinates.
+ """
+ transformed_xvalues = np.subtract(xvalues, self.centre)
+ return np.multiply(
+ np.divide(self.normalization, self.sigma * np.sqrt(2.0 * np.pi)),
+ np.exp(-0.5 * np.square(np.divide(transformed_xvalues, self.sigma))),
+ )
+
+
+"""
+__Analysis__
+
+To define the Analysis class for this model-fit, we need to ensure that the `log_likelihood_function` can handle an
+instance containing multiple 1D profiles. Below is an expanded explanation and the corresponding class definition:
+
+The log_likelihood_function will now assume that the instance it receives consists of multiple Gaussian profiles.
+For each Gaussian in the instance, it will compute the model_data and then sum these to create the overall `model_data`
+that is compared to the observed data.
+"""
+
+
+class Analysis(af.Analysis):
+ def __init__(self, data: np.ndarray, noise_map: np.ndarray):
+ """
+ The `Analysis` class acts as an interface between the data and model in **PyAutoFit**.
+
+ Its `log_likelihood_function` defines how the model is fitted to the data and it is called many times by
+ the non-linear search fitting algorithm.
+
+ In this example, the `log_likelihood_function` receives an instance containing multiple instances of
+ the `Gaussian` class and sums the `model_data` of each to create the overall model fit to the data.
+
+ In this example the `Analysis` `__init__` constructor only contains the `data` and `noise-map`, but it can be
+ easily extended to include other quantities.
+
+ Parameters
+ ----------
+ data
+ A 1D numpy array containing the data (e.g. a noisy 1D signal) fitted in the workspace examples.
+ noise_map
+ A 1D numpy array containing the noise values of the data, used for computing the goodness of fit
+ metric, the log likelihood.
+ """
+ super().__init__()
+
+ self.data = data
+ self.noise_map = noise_map
+
+ def log_likelihood_function(self, instance) -> float:
+ """
+ Returns the log likelihood of a fit of a 1D Gaussian to the dataset.
+
+ In the previous tutorial, the instance was a single `Gaussian` profile, however this function now assumes
+ the instance contains multiple `Gaussian` profiles.
+
+ The `model_data` is therefore the summed `model_data` of all individual Gaussians in the model.
+
+ The docstring below describes this in more detail.
+
+ Parameters
+ ----------
+ instance
+ A list of 1D profiles with parameters set via the non-linear search.
+
+ Returns
+ -------
+ float
+ The log likelihood value indicating how well this model fit the `MaskedDataset`.
+ """
+
+ """
+ In the previous tutorial the instance was a single `Gaussian` profile, meaning we could create the model data
+ using the line:
+
+ model_data = instance.gaussian.model_data_from(xvalues=self.data.xvalues)
+
+ In this tutorial our instance is comprised of three 1D Gaussians, because we will use a `Collection` to
+ compose the model:
+
+ model = Collection(gaussian_0=Gaussian, gaussian_1=Gaussian, gaussian_2=Gaussian).
+
+ By using a Collection, this means the instance parameter input into the fit function is a
+ dictionary where individual profiles (and their parameters) can be accessed as followed:
+
+ print(instance.gaussian_0)
+ print(instance.gaussian_1)
+ print(instance.gaussian_2)
+
+ print(instance.gaussian_0.centre)
+ print(instance.gaussian_1.centre)
+ print(instance.gaussian_2.centre)
+
+ The `model_data` is therefore the summed `model_data` of all individual Gaussians in the model.
+
+ The function `model_data_from_instance` performs this summation.
+ """
+ model_data = self.model_data_from_instance(instance=instance)
+
+ residual_map = self.data - model_data
+ chi_squared_map = (residual_map / self.noise_map) ** 2.0
+ chi_squared = sum(chi_squared_map)
+ noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
+ log_likelihood = -0.5 * (chi_squared + noise_normalization)
+
+ return log_likelihood
+
+ def model_data_from_instance(self, instance):
+ """
+ To create the summed profile of all individual profiles, we use a list comprehension to iterate over
+ all profiles in the instance.
+
+ The `instance` has the properties of a Python `iterator` and therefore can be looped over using the standard
+ Python for syntax (e.g. `for profile in instance`).
+
+ __Alternative Syntax__
+
+ For those not familiar with Python list comprehensions, the code below shows how to use the instance to
+ create the summed profile using a for loop and numpy array:
+
+ model_data = np.zeros(shape=self.data.xvalues.shape[0])
+
+ for profile in instance:
+ model_data += profile.model_data_from(xvalues=self.data.xvalues)
+
+ return model_data
+ """
+ xvalues = np.arange(self.data.shape[0])
+
+ return sum([profile.model_data_from(xvalues=xvalues) for profile in instance])
+
+
+"""
+__Collection__
+
+In the previous tutorial, we fitted a single `Gaussian` profile to the dataset by turning it into a model
+component using the `Model` class.
+
+In this tutorial, we will fit a model composed of five `Gaussian` profiles. To do this, we need to combine
+five `Gaussian` model components into a single model.
+
+This can be achieved using a `Collection` object, which was introduced in tutorial 1. The `Collection` object allows
+us to group together multiple model components—in this case, five `Gaussian` profiles—into one model that can be
+passed to the non-linear search.
+"""
+model = af.Collection(
+ gaussian_0=Gaussian,
+ gaussian_1=Gaussian,
+ gaussian_2=Gaussian,
+ gaussian_3=Gaussian,
+ gaussian_4=Gaussian,
+)
+
+"""
+The `model.info` confirms the model is composed of 5 `Gaussian` profiles.
+"""
+print(model.info)
+
+"""
+__Search__
+
+We again use the nested sampling algorithm Dynesty to fit the model to the data.
+"""
+search = af.DynestyStatic(
+ sample="rwalk", # This makes dynesty run faster, don't worry about what it means for now!
+)
+
+"""
+__Model Fit__
+
+Perform the fit using our five `Gaussian` model, which has 15 free parameters.
+
+This means the non-linear parameter space has a dimensionality of N=15, making it significantly more complex
+than the simpler model we fitted in the previous tutorial.
+
+Consequently, the non-linear search takes slightly longer to run but still completes in under a minute.
+"""
+analysis = Analysis(data=data, noise_map=noise_map)
+
+print(
+ """
+ The non-linear search has begun running.
+ This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
+ """
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+print("The search has finished run - you may now continue the notebook.")
+
+"""
+__Result__
+
+The `info` attribute shows the result in a readable format, which contains information on the full collection
+of all 5 model components.
+"""
+print(result.info)
+
+"""
+From the result info, it is hard to assess if the model fit was good or not.
+
+A good way to evaluate the fit is through a visual inspection of the model data plotted over the actual data.
+
+If the model data (red line) consistently aligns with the data points (black error bars), the fit is good.
+However, if the model misses certain features of the data, such as peaks or regions of high intensity,
+the fit was not successful.
+"""
+instance = result.max_log_likelihood_instance
+
+model_data_0 = instance.gaussian_0.model_data_from(xvalues=np.arange(data.shape[0]))
+model_data_1 = instance.gaussian_1.model_data_from(xvalues=np.arange(data.shape[0]))
+model_data_2 = instance.gaussian_2.model_data_from(xvalues=np.arange(data.shape[0]))
+model_data_3 = instance.gaussian_3.model_data_from(xvalues=np.arange(data.shape[0]))
+model_data_4 = instance.gaussian_4.model_data_from(xvalues=np.arange(data.shape[0]))
+
+model_data_list = [model_data_0, model_data_1, model_data_2, model_data_3, model_data_4]
+
+model_data = sum(model_data_list)
+
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(range(data.shape[0]), model_data, color="r")
+for model_data_1d_individual in model_data_list:
+ plt.plot(range(data.shape[0]), model_data_1d_individual, "--")
+plt.title(f"Fit (log likelihood = {result.log_likelihood})")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+It's challenging to determine from the plot whether the data and model data perfectly overlap across the entire dataset.
+
+To clarify this, the residual map introduced in tutorial 2 is useful. It provides a clear representation of where
+the differences between the model and data exceed the noise level.
+
+Regions where the black error bars do not align with the zero line in the residual map indicate areas where the model
+did not fit the data well and is inconsistent with the data above the noise level. Furthermore, regions where
+larger values of residuals are next to one another indicate that the model failed to accurate fit that
+region of the data.
+"""
+residual_map = data - model_data
+plt.plot(range(data.shape[0]), np.zeros(data.shape[0]), "--", color="b")
+plt.errorbar(
+ x=xvalues,
+ y=residual_map,
+ yerr=noise_map,
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+ linestyle="",
+)
+plt.title(f"Residuals (log likelihood = {result.log_likelihood})")
+plt.xlabel("x values of profile")
+plt.ylabel("Residuals")
+plt.show()
+plt.clf()
+plt.close()
+
+"""
+The normalized residual map, as discussed in tutorial 2, provides an alternative visualization of the fit quality.
+
+Normalized residuals indicate the standard deviation (σ) level at which the residuals could have been drawn from the
+noise. For instance, a normalized residual of 2.0 suggests that a residual value is 2.0σ away from the noise,
+implying there is a 5% chance such a residual would occur due to noise.
+
+Values of normalized residuals above 3.0 are particularly improbable (occurring only 0.3% of the time), which is
+generally considered a threshold where issues with the model-fit are likely the cause of the residual as opposed
+to it being a noise fluctuation.
+"""
+residual_map = data - model_data
+normalized_residual_map = residual_map / noise_map
+plt.plot(xvalues, normalized_residual_map, color="k")
+plt.title(f"Normalized Residuals (log likelihood = {result.log_likelihood})")
+plt.xlabel("x values of profile")
+plt.ylabel("Normalized Residuals ($\sigma$)")
+plt.show()
+plt.clf()
+plt.close()
+
+"""
+So, did you achieve a good fit? Maybe a bad one? Or just an okay one?
+
+The truth is, I don't know, and I can't tell you for sure. Modeling is inherently random. It's not uncommon to
+fit the same model to the same dataset using the same non-linear search and get a different result each time.
+
+When I ran the model fit above, that's exactly what happened. It produced a range of fits: some bad, some okay, and
+some good, as shown in the images below:
+
+
+
+Distinguishing between the good and okay fit is difficult, however the normalized residuals make this easier. They show
+that for the okay fit there are residuals above 3.0 sigma, indicating that the model did not perfectly fit the data.
+
+
+
+You should quickly rerun the code above a couple of times to see this variability for yourself.
+
+__Why Modeling is Hard__
+
+This variability is at the heart of why modeling is challenging. The process of model-fitting is stochastic,
+meaning it's hard as the scientist to determine if a better fit is possible or not.
+
+Why does modeling produce different results each time, and why might it sometimes infer solutions that are not good fits?
+
+In the previous tutorial, the non-linear search consistently found models that visually matched the data well,
+minimizing residuals and yielding high log likelihood values. These optimal solutions are called 'global maxima',
+they are where the model parameters correspond to the highest likelihood regions across the entire parameter space.
+This ideal scenario is illustrated in the `good_fit.png` image above.
+
+However, non-linear searches do not always find these global maxima. Instead, they might settle on 'local maxima'
+solutions, which have high log likelihood values relative to nearby models in parameter space but are significantly
+lower than the true global maxima found elsewhere.
+
+This is what happened for the okay and bad fits above. The non-linear search converged on solutions that were locally
+peaks on the likelihood surface but were not the global maximum solution. This is why the residuals were higher and
+the normalized residuals above 3.0 sigma.
+
+Why does a non-linear search end up at local maxima? As discussed, the search iterates through many models,
+focusing more on regions where previous guesses yielded higher likelihoods. It gradually converges around
+solutions with higher likelihoods compared to surrounding models. If the search isn't exhaustive enough, it might
+converge on a local maxima that appears good compared to nearby models but isn't the global maximum.
+
+Modeling is challenging because the parameter spaces of complex models are typically filled with local maxima,
+making it hard for a non-linear search to locate the global maximum.
+
+Fortunately, there are strategies to help non-linear searches find the global maxima, and we'll now explore three of
+them.
+
+__Prior Tuning__
+
+First, let's assist our non-linear search by tuning our priors. Priors provide guidance to the search on where to
+explore in the parameter space. By setting more accurate priors ('tuning' them), we can help the search find the
+global solution instead of settling for a local maximum.
+
+For instance, from the data itself, it's evident that all `Gaussian` profiles are centered around pixel 50. In our
+previous fit, the `centre` parameter of each `Gaussian` had a `UniformPrior` spanning from 0.0 to 100.0, which is
+much broader than necessary given the data's range.
+
+Additionally, the peak value of the data's `normalization` parameter was around 17.5. This indicates that
+the `normalization` values of our `Gaussians` do not exceed 500.0, allowing us to refine our prior accordingly.
+
+The following code snippet adjusts these priors for the `centre` and `normalization` parameters of
+each `Gaussian` using **PyAutoFit**'s API for model and prior customization:
+"""
+gaussian_0 = af.Model(Gaussian)
+
+gaussian_0.centre = af.UniformPrior(lower_limit=45.0, upper_limit=55.0)
+gaussian_0.normalization = af.LogUniformPrior(lower_limit=0.1, upper_limit=500.0)
+
+gaussian_1 = af.Model(Gaussian)
+
+gaussian_1.centre = af.UniformPrior(lower_limit=45.0, upper_limit=55.0)
+gaussian_1.normalization = af.LogUniformPrior(lower_limit=0.1, upper_limit=500.0)
+
+gaussian_2 = af.Model(Gaussian)
+
+gaussian_2.centre = af.UniformPrior(lower_limit=45.0, upper_limit=55.0)
+gaussian_2.normalization = af.LogUniformPrior(lower_limit=0.1, upper_limit=500.0)
+
+gaussian_3 = af.Model(Gaussian)
+
+gaussian_3.centre = af.UniformPrior(lower_limit=45.0, upper_limit=55.0)
+gaussian_3.normalization = af.LogUniformPrior(lower_limit=0.1, upper_limit=500.0)
+
+gaussian_4 = af.Model(Gaussian)
+
+gaussian_4.centre = af.UniformPrior(lower_limit=45.0, upper_limit=55.0)
+gaussian_4.normalization = af.LogUniformPrior(lower_limit=0.1, upper_limit=500.0)
+
+model = af.Collection(
+ gaussian_0=gaussian_0,
+ gaussian_1=gaussian_1,
+ gaussian_2=gaussian_2,
+ gaussian_3=gaussian_3,
+ gaussian_4=gaussian_4,
+)
+
+"""
+The `info` attribute shows the model is now using the priors specified above.
+"""
+print(model.info)
+
+"""
+We now repeat the model-fit using these updated priors.
+
+First, you should note that the run time of the fit is significantly faster than the previous fit. This is because
+the prior is telling the non-linear search where to look, meaning it converges on solutions more quickly
+and spends less time searching regions of parameter space that do not contain solutions.
+
+Second, the model-fit consistently produces a good model-fit more often, because our tuned priors are centred
+on the global maxima solution ensuring the non-linear search is less likely to converge on a local maxima.
+"""
+print(
+ """
+ The non-linear search has begun running.
+ This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
+ """
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+print("The search has finished run - you may now continue the notebook.")
+
+"""
+Lets print the result info and plot the fit to the dataset to confirm the priors have provided a better model-fit.
+"""
+print(result.info)
+
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+ linestyle="",
+)
+plt.plot(range(data.shape[0]), model_data, color="r")
+for model_data_1d_individual in model_data_list:
+ plt.plot(range(data.shape[0]), model_data_1d_individual, "--")
+plt.title(f"Fit (log likelihood = {result.log_likelihood})")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.clf()
+plt.close()
+
+residual_map = data - model_data
+normalized_residual_map = residual_map / noise_map
+plt.plot(xvalues, normalized_residual_map, color="k")
+plt.title(f"Normalized Residuals (log likelihood = {result.log_likelihood})")
+plt.xlabel("x values of profile")
+plt.ylabel("Normalized Residuals ($\sigma$)")
+plt.show()
+plt.clf()
+plt.close()
+
+"""
+Lets consider the advantages and disadvantages of prior tuning:
+
+**Advantages:**
+
+- Higher likelihood of finding the global maximum log likelihood solutions in parameter space.
+
+- Faster search times, as the non-linear search explores less of the parameter space.
+
+**Disadvantages:**
+
+- Incorrectly specified priors could lead the non-linear search to an incorrect solution.
+
+- It is not always clear how the priors should be tuned, especially for complex models.
+
+- Priors tuning must be applied to each dataset fitted. For large datasets, this process would be very time-consuming.
+
+__Reducing Complexity__
+
+The non-linear search may fail because the model is too complex, making its parameter space too difficult to
+sample accurately consistent. To address this, we may be able to simplify the model while ensuring it remains
+realistic enough for our scientific study. By making certain assumptions, we can reduce the number of model
+parameters, thereby lowering the dimensionality of the parameter space and improving the search's performance.
+
+For example, we may know that the `Gaussian`'s in our model are aligned at the same `centre`. We can therefore
+compose a model that assumes that the `centre` of each `Gaussian` is the same, reducing the dimensionality of the
+model from N=15 to N=11.
+
+The code below shows how we can customize the model components to ensure the `centre` of each `Gaussian` is the same:
+"""
+gaussian_0 = af.Model(Gaussian)
+gaussian_1 = af.Model(Gaussian)
+gaussian_2 = af.Model(Gaussian)
+gaussian_3 = af.Model(Gaussian)
+gaussian_4 = af.Model(Gaussian)
+
+gaussian_1.centre = gaussian_0.centre
+gaussian_2.centre = gaussian_0.centre
+gaussian_3.centre = gaussian_0.centre
+gaussian_4.centre = gaussian_0.centre
+
+model = af.Collection(
+ gaussian_0=gaussian_0,
+ gaussian_1=gaussian_1,
+ gaussian_2=gaussian_2,
+ gaussian_3=gaussian_3,
+ gaussian_4=gaussian_4,
+)
+
+"""
+The `info` attribute shows the model is now using the same `centre` for all `Gaussian`'s and has 11 free parameters.
+"""
+print(model.info)
+
+"""
+We now repeat the model-fit using this updated model where the `centre` of each `Gaussian` is the same.
+
+You should again note that the run time of the fit is significantly faster than the previous fits
+and that it consistently produces a good model-fit more often. This is because the model is less complex,
+non-linear parameter space is less difficult to sample and the search is less likely to converge on a local maxima.
+"""
+print(
+ """
+ The non-linear search has begun running.
+ This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
+ """
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+print("The search has finished run - you may now continue the notebook.")
+
+"""
+Lets print the result info and plot the fit to the dataset to confirm the reduced model complexity has
+provided a better model-fit.
+"""
+print(result.info)
+
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+ linestyle="",
+)
+plt.plot(range(data.shape[0]), model_data, color="r")
+for model_data_1d_individual in model_data_list:
+ plt.plot(range(data.shape[0]), model_data_1d_individual, "--")
+plt.title(f"Fit (log likelihood = {result.log_likelihood})")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.clf()
+plt.close()
+
+residual_map = data - model_data
+normalized_residual_map = residual_map / noise_map
+plt.plot(xvalues, normalized_residual_map, color="k")
+plt.title(f"Normalized Residuals (log likelihood = {result.log_likelihood})")
+plt.xlabel("x values of profile")
+plt.ylabel("Normalized Residuals ($\sigma$)")
+plt.show()
+plt.clf()
+plt.close()
+
+"""
+Let’s consider the advantages and disadvantages of simplifying the model:
+
+Advantages:
+
+- By reducing the complexity of the parameter space, we increase the chances of finding the global maximum log
+likelihood, and the search requires less time to do so.
+
+- Unlike with tuned priors, this approach is not specific to a single dataset, allowing us to use it on many datasets.
+
+Disadvantages:
+
+- Our model is less realistic, which may negatively impact the accuracy of our fit and the scientific results we
+derive from it.
+
+__Search More Thoroughly__
+
+In approaches 1 and 2, we assisted our non-linear search to find the highest log likelihood regions of parameter
+space. In approach 3, we're simply going to tell the search to look more thoroughly through parameter space.
+
+Every non-linear search has settings that control how thoroughly it explores parameter space. For Dynesty, the
+primary setting is the number of live points `nlive`. The more thoroughly the search examines the space, the more
+likely it is to find the global maximum model. However, this also means the search will take longer!
+
+Below, we configure a more thorough Dynesty search with `nlive=500`. It is currently unclear what changing
+this setting actually does and what the number of live points actually means. These will be covered in chapter 2
+of the **HowToFit** lectures, where we will also expand on how a non-linear search actually works and the different
+types of methods that can be used to search parameter space.
+"""
+model = af.Collection(
+ gaussian_0=Gaussian,
+ gaussian_1=Gaussian,
+ gaussian_2=Gaussian,
+ gaussian_3=Gaussian,
+ gaussian_4=Gaussian,
+)
+
+"""
+The `model.info` confirms the model is the same model fitted initially, composed of 5 `Gaussian` profiles.
+"""
+print(model.info)
+
+"""
+__Search__
+
+We again use the nested sampling algorithm Dynesty to fit the model to the data, but now increase the number of live
+points to 300 meaning it will search parameter space more thoroughly.
+"""
+search = af.DynestyStatic(
+ nlive=300,
+ sample="rwalk", # This makes dynesty run faster, don't worry about what it means for now!
+)
+
+"""
+__Model Fit__
+
+Perform the fit using our five `Gaussian` model, which has 15 free parameters.
+"""
+analysis = Analysis(data=data, noise_map=noise_map)
+
+print(
+ """
+ The non-linear search has begun running.
+ This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
+ """
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+print("The search has finished run - you may now continue the notebook.")
+
+
+"""
+Lets print the result info and plot the fit to the dataset to confirm the more thorough search has provided a better
+model-fit.
+"""
+print(result.info)
+
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+ linestyle="",
+)
+plt.plot(range(data.shape[0]), model_data, color="r")
+for model_data_1d_individual in model_data_list:
+ plt.plot(range(data.shape[0]), model_data_1d_individual, "--")
+plt.title(f"Fit (log likelihood = {result.log_likelihood})")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.clf()
+plt.close()
+
+residual_map = data - model_data
+normalized_residual_map = residual_map / noise_map
+plt.plot(xvalues, normalized_residual_map, color="k")
+plt.title(f"Normalized Residuals (log likelihood = {result.log_likelihood})")
+plt.xlabel("x values of profile")
+plt.ylabel("Normalized Residuals ($\sigma$)")
+plt.show()
+plt.clf()
+plt.close()
+
+"""
+If you repeat the fit multiple times, you will find that the model-fit is more likely to produce a good fit than
+previously.
+
+However, the run-time of the search is noticeably longer, taking a few minutes to complete, owining
+to the increased number of live points and fact it is searching parameter space more thoroughly.
+
+Let's list the advantages and disadvantages of simply adjusting the non-linear search:
+
+**Advantages:**
+
+- It’s easy to set up; just change the settings of the non-linear search.
+
+- It generalizes to any dataset.
+
+- We can retain a more complex model.
+
+**Disadvantage:**
+
+- It can be very expensive in terms of run time, producing run-times that are five, tens or even longer than the
+ original run-time.
+
+__Summary__
+
+We have covered three strategies for ensuring the non-linear search estimates the correct solution:
+
+1) Prior Tuning: By setting our priors more accurately, we can help the search find the global solution instead of
+ getting stuck at a local maxima.
+
+2) Reducing Complexity: By making certain assumptions, we can reduce the number of model parameters, thereby lowering
+ the dimensionality of the parameter space and improving the search's performance.
+
+3) Search More Thoroughly: By increasing the number of live points, we can make the search explore parameter space
+ more thoroughly, increasing the likelihood of finding the global maximum model.
+
+Each of these strategies has its advantages and disadvantages, and your ability to fit models successfully will
+likely rely on a combination of these strategies. Which one works best depends on the specific model, dataset and
+scientific question you are trying to answer. Therefore, when you begin your own model-fitting, it is a good idea to
+try each of these strategies and assess which works best for your model-fit.
+
+__Run Times__
+
+One challenging aspect of model-fitting which was not properly covered in this tutorial is the run-time of a model-fit.
+This example fits simple 1D datasets, which are computationally inexpensive to fit. That is, the `log_likelihood_function`
+is evaluated in a fraction of a second, meaning the non-linear search fitted the model in mere minutes.
+
+Many model-fitting tasks are not as fast. For example, when fitting a model to a 2D image, the `log_likelihood_function`
+may take of order seconds, or longer, because it comprises a number of expensive calculations (e.g. a Fourier transform,
+2D convolution, etc.). Depending on the model complexity, this means that the non-linear search may take hours, days
+or even weeks to fit the model.
+
+Run times are also dictated by the complexity of the model and the nature of the log likelihood function. For models
+with many more dimensions than the simple 1D model used in this tutorial (e.g. hundreds or thousands of free parameters),
+non-linear search may take tens or hundreds of more iterations to converge on a solution. This is because the parameter
+space is significantly more complex and difficult to sample accurately. More iterations mean longer run times,
+which in combination with a slow likelihood function can make model-fitting infeasible.
+
+Whether or not run times will pose a challenge to your model-fitting task depends on the complexity of the model and
+nature of the log likelihood function. If your problem is computationally expensive, **PyAutoFit** provides many
+tools to help, which will be the topic of tutorials in chapter 2 of the **HowToFit** lectures.
+
+**PyAutoFit** provides tools to profile the run-time of your log likelihood function, which can be used to
+assess the computational expense of your model-fit and plan accordingly. Below is an example of the simplest use
+of these tools, an estimate of the run-time of the log likelihood function using one random instance of the model.
+
+
+Feature still being developed, IGNORE FOR NOW.
+
+run_time_dict, info_dict = analysis.profile_log_likelihood_function(
+ instance=model.random_instance()
+)
+print(f"Log Likelihood Evaluation Time (second) = {run_time_dict['fit_time']}")
+
+__Model Mismatch__
+
+In this example, interpreting how well the model fitted the data, and whether it found the global maxima, was
+relatively straightforward. This is because the same model was used to simulate the data and fit it, meaning the
+global maxima fit corresponded to one where the normalized residuals were minimized and consistent with the noise
+(e.g. they went to sigma values below 3.0 or so).
+
+In many scientific studies, the data that one is fitting may have come from an instrument or simulation where the
+exact physical processes that generate the data are not perfectly known. This then means that the model is
+not a perfect representation of the data, and it may not ever be possible to fit the data perfectly. In this case,
+we might infer a fit with significant residuals, but it may still correspond to the global maxima solution,
+at least for that particular model.
+
+This makes it even more difficult to be certain if the non-linear search is sampling parameter space correctly,
+and therefore requires even more care and attention to the strategies we have discussed above.
+
+Whether or not this is the case for your model-fitting task is something you will have to determine yourself.
+**PyAutoFit** provides many tools to help assess the quality of a model-fit, which will be the topic of tutorials
+in chapter 2 of the **HowToFit** lectures.
+
+__Astronomy Example__
+
+At the end of chapter 1, we will fit a complex model to a real astronomical dataset in order to quantify
+the distribution of stars in 2D images of galaxies.
+
+This example will illustrate many of the challenges discussed in this tutorial, including:
+
+- Fits using more complex models consisting of 15-20 parameters often infer local maxima, unless we assist the
+ non-linear search with tuned priors, reduced complexity or a more thorough search.
+
+- Fitting 2D imaging data requires a 2D convolution, which is somewhat computationally expensive and means run times
+ become something we must balance with model complexity.
+
+- The model is not a perfect representation of the data. For example, the model assumes the galaxy is elliptically
+ symmetric, whereas the real galaxy may not be. In certain examples, this means that the global maxima solution
+ actually leaves significant residuals, above 3.0 $\sigma$, in the data.
+
+__Wrap Up__
+
+Now is a good time to assess how straightforward or difficult you think your model-fitting task will be.
+
+Are the models you will be fitting made up of tens of parameters? or thousands? Are there ways you can simplify
+the model parameter or tune priors to make the model-fitting task more feasible? Will run times be an issue, or is
+your likelihood function computationally cheap? And how confident are you that the model you are fitting is a good
+representation of the data?
+
+These are all questions you should be asking yourself before beginning your model-fitting task, but they will
+become easier to answer as you gain experience with model-fitting and **PyAutoFit**.
+"""
diff --git a/scripts/howtofit/chapter_1_introduction/tutorial_5_results_and_samples.py b/scripts/howtofit/chapter_1_introduction/tutorial_5_results_and_samples.py
index e484f81b..65c0891e 100644
--- a/scripts/howtofit/chapter_1_introduction/tutorial_5_results_and_samples.py
+++ b/scripts/howtofit/chapter_1_introduction/tutorial_5_results_and_samples.py
@@ -1,697 +1,693 @@
-"""
-Tutorial 5: Results And Samples
-===============================
-
-In this tutorial, we'll cover all of the output that comes from a non-linear search's `Result` object.
-
-We used this object at various points in the chapter. The bulk of material covered here is described in the example
-script `autofit_workspace/overview/simple/result.py`. Nevertheless, it is a good idea to refresh ourselves about how
-results in **PyAutoFit** work before covering more advanced material.
-
-__Contents__
-
-This tutorial is split into the following sections:
-
-- **Data**: Load the dataset from the autofit_workspace/dataset folder.
-- **Reused Functions**: Reuse the `plot_profile_1d` and `Analysis` classes from the previous tutorial.
-- **Model Fit**: Run a non-linear search to generate a `Result` object.
-- **Result**: Examine the `Result` object and its info attribute.
-- **Samples**: Introduce the `Samples` object containing the non-linear search samples.
-- **Parameters**: Access parameter values from the samples.
-- **Figures of Merit**: Examine log likelihood, log prior, and log posterior values.
-- **Instances**: Return results as model instances from samples.
-- **Vectors**: Return results as 1D parameter vectors.
-- **Labels**: Access the paths, names, and labels for model parameters.
-- **Posterior / PDF**: Access median PDF estimates for the model parameters.
-- **Plot**: Visualize model fit results using instances.
-- **Errors**: Compute parameter error estimates at specified sigma confidence limits.
-- **PDF**: Plot Probability Density Functions using corner.py.
-- **Other Results**: Access maximum log posterior and other sample statistics.
-- **Sample Instance**: Create instances from individual samples in the sample list.
-- **Bayesian Evidence**: Access the log evidence for nested sampling searches.
-- **Derived Errors (PDF from samples)**: Compute errors on derived quantities from sample PDFs.
-- **Samples Filtering**: Filter samples by parameter paths for specific parameter analysis.
-- **Latex**: Generate LaTeX table code for modeling results.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import autofit as af
-import autofit.plot as aplt
-import os
-from os import path
-import numpy as np
-import matplotlib.pyplot as plt
-
-"""
-__Data__
-
-Load the dataset from the `autofit_workspace/dataset` folder.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1__exponential_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-"""
-__Reused Functions__
-
-We'll reuse the `plot_profile_1d` and `Analysis` classes of the previous tutorial.
-"""
-
-
-def plot_profile_1d(
- xvalues,
- profile_1d,
- title=None,
- ylabel=None,
- errors=None,
- color="k",
- output_path=None,
- output_filename=None,
-):
- plt.errorbar(
- x=xvalues,
- y=profile_1d,
- yerr=errors,
- linestyle="",
- color=color,
- ecolor="k",
- elinewidth=1,
- capsize=2,
- )
- plt.title(title)
- plt.xlabel("x value of profile")
- plt.ylabel(ylabel)
- if not path.exists(output_path):
- os.makedirs(output_path)
- plt.savefig(path.join(output_path, f"{output_filename}.png"))
- plt.clf()
-
-
-class Analysis(af.Analysis):
- def __init__(self, data, noise_map):
- super().__init__()
-
- self.data = data
- self.noise_map = noise_map
-
- def log_likelihood_function(self, instance):
- model_data = self.model_data_from_instance(instance=instance)
-
- residual_map = self.data - model_data
- chi_squared_map = (residual_map / self.noise_map) ** 2.0
- chi_squared = sum(chi_squared_map)
- noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
- log_likelihood = -0.5 * (chi_squared + noise_normalization)
-
- return log_likelihood
-
- def model_data_from_instance(self, instance):
- """
- To create the summed profile of all individual profiles in an instance, we can use a dictionary comprehension
- to iterate over all profiles in the instance.
- """
- xvalues = np.arange(self.data.shape[0])
-
- return sum([profile.model_data_from(xvalues=xvalues) for profile in instance])
-
- def visualize(self, paths, instance, during_analysis):
- """
- This method is identical to the previous tutorial, except it now uses the `model_data_from_instance` method
- to create the profile.
- """
- xvalues = np.arange(self.data.shape[0])
-
- model_data = self.model_data_from_instance(instance=instance)
-
- residual_map = self.data - model_data
- chi_squared_map = (residual_map / self.noise_map) ** 2.0
-
- """The visualizer now outputs images of the best-fit results to hard-disk (checkout `visualizer.py`)."""
- plot_profile_1d(
- xvalues=xvalues,
- profile_1d=self.data,
- title="Data",
- ylabel="Data Values",
- color="k",
- output_path=paths.image_path,
- output_filename="data",
- )
-
- plot_profile_1d(
- xvalues=xvalues,
- profile_1d=model_data,
- title="Model Data",
- ylabel="Model Data Values",
- color="k",
- output_path=paths.image_path,
- output_filename="model_data",
- )
-
- plot_profile_1d(
- xvalues=xvalues,
- profile_1d=residual_map,
- title="Residual Map",
- ylabel="Residuals",
- color="k",
- output_path=paths.image_path,
- output_filename="residual_map",
- )
-
- plot_profile_1d(
- xvalues=xvalues,
- profile_1d=chi_squared_map,
- title="Chi-Squared Map",
- ylabel="Chi-Squareds",
- color="k",
- output_path=paths.image_path,
- output_filename="chi_squared_map",
- )
-
-
-"""
-__Model Fit__
-
-Now lets run the non-linear search to get ourselves a `Result`.
-"""
-
-
-class Gaussian:
- def __init__(
- self,
- centre=30.0, # <- **PyAutoFit** recognises these constructor arguments
- normalization=1.0, # <- are the Gaussian`s model parameters.
- sigma=5.0,
- ):
- """
- Represents a 1D Gaussian profile.
-
- This is a model-component of example models in the **HowToFit** lectures and is used to fit example datasets
- via a non-linear search.
-
- Parameters
- ----------
- centre
- The x coordinate of the profile centre.
- normalization
- Overall normalization of the profile.
- sigma
- The sigma value controlling the size of the Gaussian.
- """
- self.centre = centre
- self.normalization = normalization
- self.sigma = sigma
-
- def model_data_from(self, xvalues: np.ndarray):
- """
-
- Returns a 1D Gaussian on an input list of Cartesian x coordinates.
-
- The input xvalues are translated to a coordinate system centred on the Gaussian, via its `centre`.
-
- The output is referred to as the `model_data` to signify that it is a representation of the data from the
- model.
-
- Parameters
- ----------
- xvalues
- The x coordinates in the original reference frame of the data.
- """
- transformed_xvalues = np.subtract(xvalues, self.centre)
- return np.multiply(
- np.divide(self.normalization, self.sigma * np.sqrt(2.0 * np.pi)),
- np.exp(-0.5 * np.square(np.divide(transformed_xvalues, self.sigma))),
- )
-
-
-class Exponential:
- def __init__(
- self,
- centre=30.0, # <- **PyAutoFit** recognises these constructor arguments
- normalization=1.0, # <- are the Exponential`s model parameters.
- rate=0.01,
- ):
- """
- Represents a 1D Exponential profile.
-
- This is a model-component of example models in the **HowToFit** lectures and is used to fit example datasets
- via a non-linear search.
-
- Parameters
- ----------
- centre
- The x coordinate of the profile centre.
- normalization
- Overall normalization of the profile.
- ratw
- The decay rate controlling has fast the Exponential declines.
- """
- self.centre = centre
- self.normalization = normalization
- self.rate = rate
-
- def model_data_from(self, xvalues: np.ndarray):
- """
- Returns a 1D Gaussian on an input list of Cartesian x coordinates.
-
- The input xvalues are translated to a coordinate system centred on the Gaussian, via its `centre`.
-
- The output is referred to as the `model_data` to signify that it is a representation of the data from the
- model.
-
- Parameters
- ----------
- xvalues
- The x coordinates in the original reference frame of the data.
- """
- transformed_xvalues = np.subtract(xvalues, self.centre)
- return self.normalization * np.multiply(
- self.rate, np.exp(-1.0 * self.rate * abs(transformed_xvalues))
- )
-
-
-model = af.Collection(gaussian=af.Model(Gaussian), exponential=af.Model(Exponential))
-
-analysis = Analysis(data=data, noise_map=noise_map)
-
-search = af.Emcee(
- name="tutorial_5_results_and_samples",
- path_prefix=path.join("howtofit", "chapter_1"),
-)
-
-print(
- """
- The non-linear search has begun running.
- Checkout the autofit_workspace/output/howtofit/tutorial_6__results_and_samples
- folder for live output of the results.
- This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
- """
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-print("The search has finished run - you may now continue the notebook.")
-
-"""
-__Result__
-
-Here, we'll look in detail at what information is contained in the `Result`.
-
-It contains an `info` attribute which prints the result in readable format.
-"""
-print(result.info)
-
-"""
-__Samples__
-
-The result contains a `Samples` object, which contains all of the non-linear search samples.
-
-Each sample corresponds to a set of model parameters that were evaluated and accepted by our non linear search,
-in this example emcee.
-
-This also includes their log likelihoods, which are used for computing additional information about the model-fit,
-for example the error on every parameter.
-
-Our model-fit used the MCMC algorithm Emcee, so the `Samples` object returned is a `SamplesMCMC` object.
-"""
-samples = result.samples
-
-print("MCMC Samples: \n")
-print(samples)
-
-"""
-__Parameters__
-
-The parameters are stored as a list of lists, where:
-
- - The outer list is the size of the total number of samples.
- - The inner list is the size of the number of free parameters in the fit.
-"""
-samples = result.samples
-print("Sample 5's second parameter value (Gaussian -> normalization):")
-print(samples.parameter_lists[4][1])
-print("Sample 10`s third parameter value (Gaussian -> sigma)")
-print(samples.parameter_lists[9][2], "\n")
-
-"""
-__Figures of Merit__
-
-The Samples class also contains the log likelihood, log prior, log posterior and weight_list of every accepted sample,
-where:
-
-- The log likelihood is the value evaluated from the likelihood function (e.g. -0.5 * chi_squared + the noise
-normalized).
-
-- The log prior encodes information on how the priors on the parameters maps the log likelihood value to the log
-posterior value.
-
-- The log posterior is log_likelihood + log_prior.
-
-- The weight gives information on how samples should be combined to estimate the posterior. The weight values depend on
-the sampler used, for MCMC samples they are all 1 (e.g. all weighted equally).
-
-Lets inspect the last 10 values of each for the analysis.
-"""
-print("log(likelihood), log(prior), log(posterior) and weight of the tenth sample.")
-print(samples.log_likelihood_list[9])
-print(samples.log_prior_list[9])
-print(samples.log_posterior_list[9])
-print(samples.weight_list[9])
-
-"""
-__Instances__
-
-The `Samples` contains many results which are returned as an instance of the model, using the Python class structure
-of the model composition.
-
-For example, we can return the model parameters corresponding to the maximum log likelihood sample.
-"""
-max_lh_instance = samples.max_log_likelihood()
-
-print("Max Log Likelihood `Gaussian` Instance:")
-print("Centre = ", max_lh_instance.gaussian.centre)
-print("Normalization = ", max_lh_instance.gaussian.normalization)
-print("Sigma = ", max_lh_instance.gaussian.sigma, "\n")
-
-print("Max Log Likelihood Exponential Instance:")
-print("Centre = ", max_lh_instance.exponential.centre)
-print("Normalization = ", max_lh_instance.exponential.normalization)
-print("Sigma = ", max_lh_instance.exponential.rate, "\n")
-
-"""
-__Vectors__
-
-All results can alternatively be returned as a 1D vector of values, by passing `as_instance=False`:
-"""
-max_lh_vector = samples.max_log_likelihood(as_instance=False)
-print("Max Log Likelihood Model Parameters: \n")
-print(max_lh_vector, "\n\n")
-
-"""
-__Labels__
-
-Vectors return a lists of all model parameters, but do not tell us which values correspond to which parameters.
-
-The following quantities are available in the `Model`, where the order of their entries correspond to the parameters
-in the `ml_vector` above:
-
- - `paths`: a list of tuples which give the path of every parameter in the `Model`.
- - `parameter_names`: a list of shorthand parameter names derived from the `paths`.
- - `parameter_labels`: a list of parameter labels used when visualizing non-linear search results (see below).
-
-"""
-model = samples.model
-
-print(model.paths)
-print(model.parameter_names)
-print(model.parameter_labels)
-print(model.model_component_and_parameter_names)
-print("\n")
-
-"""
-From here on, we will returned all results information as instances, but every method below can be returned as a
-vector via the `as_instance=False` input.
-
-__Posterior / PDF__
-
-The ``Result`` object contains the full posterior information of our non-linear search, which can be used for
-parameter estimation.
-
-The median pdf vector is available from the `Samples` object, which estimates the every parameter via 1D
-marginalization of their PDFs.
-"""
-median_pdf_instance = samples.median_pdf()
-
-print("Max Log Likelihood `Gaussian` Instance:")
-print("Centre = ", median_pdf_instance.gaussian.centre)
-print("Normalization = ", median_pdf_instance.gaussian.normalization)
-print("Sigma = ", median_pdf_instance.gaussian.sigma, "\n")
-
-print("Max Log Likelihood Exponential Instance:")
-print("Centre = ", median_pdf_instance.exponential.centre)
-print("Normalization = ", median_pdf_instance.exponential.normalization)
-print("Sigma = ", median_pdf_instance.exponential.rate, "\n")
-
-"""
-__Plot__
-
-Because results are returned as instances, it is straight forward to use them and their associated functionality
-to make plots of the results:
-"""
-model_gaussian = max_lh_instance.gaussian.model_data_from(
- xvalues=np.arange(data.shape[0])
-)
-model_exponential = max_lh_instance.exponential.model_data_from(
- xvalues=np.arange(data.shape[0])
-)
-model_data = model_gaussian + model_exponential
-
-plt.plot(range(data.shape[0]), data)
-plt.plot(range(data.shape[0]), model_data)
-plt.plot(range(data.shape[0]), model_gaussian, "--")
-plt.plot(range(data.shape[0]), model_exponential, "--")
-plt.title("Illustrative model fit to 1D `Gaussian` + Exponential profile data.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-__Errors__
-
-The samples include methods for computing the error estimates of all parameters, via 1D marginalization at an
-input sigma confidence limit.
-"""
-errors_at_upper_sigma_instance = samples.errors_at_upper_sigma(sigma=3.0)
-errors_at_lower_sigma_instance = samples.errors_at_lower_sigma(sigma=3.0)
-
-print("Upper Error values of Gaussian (at 3.0 sigma confidence):")
-print("Centre = ", errors_at_upper_sigma_instance.gaussian.centre)
-print("Normalization = ", errors_at_upper_sigma_instance.gaussian.normalization)
-print("Sigma = ", errors_at_upper_sigma_instance.gaussian.sigma, "\n")
-
-print("lower Error values of Gaussian (at 3.0 sigma confidence):")
-print("Centre = ", errors_at_lower_sigma_instance.gaussian.centre)
-print("Normalization = ", errors_at_lower_sigma_instance.gaussian.normalization)
-print("Sigma = ", errors_at_lower_sigma_instance.gaussian.sigma, "\n")
-
-"""
-They can also be returned at the values of the parameters at their error values:
-"""
-values_at_upper_sigma_instance = samples.values_at_upper_sigma(sigma=3.0)
-values_at_lower_sigma_instance = samples.values_at_lower_sigma(sigma=3.0)
-
-print("Upper Parameter values w/ error of Gaussian (at 3.0 sigma confidence):")
-print("Centre = ", values_at_upper_sigma_instance.gaussian.centre)
-print("Normalization = ", values_at_upper_sigma_instance.gaussian.normalization)
-print("Sigma = ", values_at_upper_sigma_instance.gaussian.sigma, "\n")
-
-print("lower Parameter values w/ errors of Gaussian (at 3.0 sigma confidence):")
-print("Centre = ", values_at_lower_sigma_instance.gaussian.centre)
-print("Normalization = ", values_at_lower_sigma_instance.gaussian.normalization)
-print("Sigma = ", values_at_lower_sigma_instance.gaussian.sigma, "\n")
-
-"""
-__PDF__
-
-The Probability Density Functions (PDF's) of the results can be plotted using the Emcee's visualization
-tool `corner.py`, which is wrapped via the `aplt.corner_cornerpy` function.
-"""
-aplt.corner_cornerpy(samples=result.samples)
-
-"""
-__Other Results__
-
-The samples contain many useful vectors, including the samples with the highest posterior values.
-"""
-max_log_posterior_instance = samples.max_log_posterior()
-
-print("Maximum Log Posterior Vector:")
-print("Centre = ", max_log_posterior_instance.gaussian.centre)
-print("Normalization = ", max_log_posterior_instance.gaussian.normalization)
-print("Sigma = ", max_log_posterior_instance.gaussian.sigma, "\n")
-
-
-"""
-All methods above are available as a vector:
-"""
-median_pdf_instance = samples.median_pdf(as_instance=False)
-values_at_upper_sigma = samples.values_at_upper_sigma(sigma=3.0, as_instance=False)
-values_at_lower_sigma = samples.values_at_lower_sigma(sigma=3.0, as_instance=False)
-errors_at_upper_sigma = samples.errors_at_upper_sigma(sigma=3.0, as_instance=False)
-errors_at_lower_sigma = samples.errors_at_lower_sigma(sigma=3.0, as_instance=False)
-
-"""
-__Sample Instance__
-
-A non-linear search retains every model that is accepted during the model-fit.
-
-We can create an instance of any lens model -- below we create an instance of the last accepted model.
-"""
-instance = samples.from_sample_index(sample_index=-1)
-
-print("Gaussian Instance of last sample")
-print("Centre = ", instance.gaussian.centre)
-print("Normalization = ", instance.gaussian.normalization)
-print("Sigma = ", instance.gaussian.sigma, "\n")
-
-"""
-__Bayesian Evidence__
-
-If a nested sampling `NonLinearSearch` is used, the evidence of the model is also available which enables Bayesian
-model comparison to be performed (given we are using Emcee, which is not a nested sampling algorithm, the log evidence
-is None).:
-"""
-log_evidence = samples.log_evidence
-
-"""
-__Derived Errors (PDF from samples)__
-
-Computing the errors of a quantity like the `sigma` of the Gaussian is simple, because it is sampled by the non-linear
-search. Thus, to get their errors above we used the `Samples` object to simply marginalize over all over parameters
-via the 1D Probability Density Function (PDF).
-
-Computing errors on derived quantities is more tricky, because they are not sampled directly by the non-linear search.
-For example, what if we want the error on the full width half maximum (FWHM) of the Gaussian? In order to do this
-we need to create the PDF of that derived quantity, which we can then marginalize over using the same function we
-use to marginalize model parameters.
-
-Below, we compute the FWHM of every accepted model sampled by the non-linear search and use this determine the PDF
-of the FWHM. When combining the FWHM's we weight each value by its `weight`. For Emcee, an MCMC algorithm, the
-weight of every sample is 1, but weights may take different values for other non-linear searches.
-
-In order to pass these samples to the function `marginalize`, which marginalizes over the PDF of the FWHM to compute
-its error, we also pass the weight list of the samples.
-
-(Computing the error on the FWHM could be done in much simpler ways than creating its PDF from the list of every
-sample. We chose this example for simplicity, in order to show this functionality, which can easily be extended to more
-complicated derived quantities.)
-"""
-fwhm_list = []
-
-for sample in samples.sample_list:
- instance = sample.instance_for_model(model=samples.model)
-
- sigma = instance.gaussian.sigma
-
- fwhm = 2 * np.sqrt(2 * np.log(2)) * sigma
-
- fwhm_list.append(fwhm)
-
-median_fwhm, lower_fwhm, upper_fwhm = af.marginalize(
- parameter_list=fwhm_list, sigma=3.0, weight_list=samples.weight_list
-)
-
-print(f"FWHM = {median_fwhm} ({upper_fwhm} {lower_fwhm}")
-
-"""
-__Samples Filtering__
-
-Our samples object has the results for all three parameters in our model. However, we might only be interested in the
-results of a specific parameter.
-
-The basic form of filtering specifies parameters via their path, which was printed above via the model and is printed
-again below.
-"""
-samples = result.samples
-
-print("Parameter paths in the model which are used for filtering:")
-print(samples.model.paths)
-
-print("All parameters of the very first sample")
-print(samples.parameter_lists[0])
-
-samples = samples.with_paths([("gaussian", "centre")])
-
-print("All parameters of the very first sample (containing only the Gaussian centre.")
-print(samples.parameter_lists[0])
-
-print("Maximum Log Likelihood Model Instances (containing only the Gaussian centre):\n")
-print(samples.max_log_likelihood(as_instance=False))
-
-"""
-Above, we specified each path as a list of tuples of strings.
-
-This is how the source code internally stores the path to different components of the model, but it is not
-in-profile_1d with the PyAutoFIT API used to compose a model.
-
-We can alternatively use the following API:
-"""
-samples = result.samples
-
-samples = samples.with_paths(["gaussian.centre"])
-
-print("All parameters of the very first sample (containing only the Gaussian centre).")
-print(samples.parameter_lists[0])
-
-"""
-Above, we filtered the `Samples` but asking for all parameters which included the path ("gaussian", "centre").
-
-We can alternatively filter the `Samples` object by removing all parameters with a certain path. Below, we remove
-the Gaussian's `centre` to be left with 2 parameters; the `normalization` and `sigma`.
-"""
-samples = result.samples
-
-print("Parameter paths in the model which are used for filtering:")
-print(samples.model.paths)
-
-print("All parameters of the very first sample")
-print(samples.parameter_lists[0])
-
-samples = samples.without_paths(["gaussian.centre"])
-
-print(
- "All parameters of the very first sample (containing only the Gaussian normalization and sigma)."
-)
-print(samples.parameter_lists[0])
-
-"""
-__Latex__
-
-If you are writing modeling results up in a paper, you can use inbuilt latex tools to create latex table
-code which you can copy to your .tex document.
-
-By combining this with the filtering tools below, specific parameters can be included or removed from the latex.
-
-Remember that the superscripts of a parameter are loaded from the config file `notation/label.yaml`, providing high
-levels of customization for how the parameter names appear in the latex table. This is especially useful if your model
-uses the same model components with the same parameter, which therefore need to be distinguished via superscripts.
-"""
-latex = af.text.Samples.latex(
- samples=result.samples,
- median_pdf_model=True,
- sigma=3.0,
- name_to_label=True,
- include_name=True,
- include_quickmath=True,
- prefix="Example Prefix ",
- suffix=" \\[-2pt]",
-)
-
-print(latex)
-
-"""
-Finish.
-"""
+"""
+Tutorial 5: Results And Samples
+===============================
+
+In this tutorial, we'll cover all of the output that comes from a non-linear search's `Result` object.
+
+We used this object at various points in the chapter. The bulk of material covered here is described in the example
+script `autofit_workspace/overview/simple/result.py`. Nevertheless, it is a good idea to refresh ourselves about how
+results in **PyAutoFit** work before covering more advanced material.
+
+__Contents__
+
+This tutorial is split into the following sections:
+
+- **Data**: Load the dataset from the autofit_workspace/dataset folder.
+- **Reused Functions**: Reuse the `plot_profile_1d` and `Analysis` classes from the previous tutorial.
+- **Model Fit**: Run a non-linear search to generate a `Result` object.
+- **Result**: Examine the `Result` object and its info attribute.
+- **Samples**: Introduce the `Samples` object containing the non-linear search samples.
+- **Parameters**: Access parameter values from the samples.
+- **Figures of Merit**: Examine log likelihood, log prior, and log posterior values.
+- **Instances**: Return results as model instances from samples.
+- **Vectors**: Return results as 1D parameter vectors.
+- **Labels**: Access the paths, names, and labels for model parameters.
+- **Posterior / PDF**: Access median PDF estimates for the model parameters.
+- **Plot**: Visualize model fit results using instances.
+- **Errors**: Compute parameter error estimates at specified sigma confidence limits.
+- **PDF**: Plot Probability Density Functions using corner.py.
+- **Other Results**: Access maximum log posterior and other sample statistics.
+- **Sample Instance**: Create instances from individual samples in the sample list.
+- **Bayesian Evidence**: Access the log evidence for nested sampling searches.
+- **Derived Errors (PDF from samples)**: Compute errors on derived quantities from sample PDFs.
+- **Samples Filtering**: Filter samples by parameter paths for specific parameter analysis.
+- **Latex**: Generate LaTeX table code for modeling results.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import autofit as af
+import autofit.plot as aplt
+import os
+from os import path
+import numpy as np
+import matplotlib.pyplot as plt
+
+"""
+__Data__
+
+Load the dataset from the `autofit_workspace/dataset` folder.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1__exponential_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+"""
+__Reused Functions__
+
+We'll reuse the `plot_profile_1d` and `Analysis` classes of the previous tutorial.
+"""
+
+
+def plot_profile_1d(
+ xvalues,
+ profile_1d,
+ title=None,
+ ylabel=None,
+ errors=None,
+ color="k",
+ output_path=None,
+ output_filename=None,
+):
+ plt.errorbar(
+ x=xvalues,
+ y=profile_1d,
+ yerr=errors,
+ linestyle="",
+ color=color,
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+ )
+ plt.title(title)
+ plt.xlabel("x value of profile")
+ plt.ylabel(ylabel)
+ if not path.exists(output_path):
+ os.makedirs(output_path)
+ plt.savefig(path.join(output_path, f"{output_filename}.png"))
+ plt.clf()
+
+
+class Analysis(af.Analysis):
+ def __init__(self, data, noise_map):
+ super().__init__()
+
+ self.data = data
+ self.noise_map = noise_map
+
+ def log_likelihood_function(self, instance):
+ model_data = self.model_data_from_instance(instance=instance)
+
+ residual_map = self.data - model_data
+ chi_squared_map = (residual_map / self.noise_map) ** 2.0
+ chi_squared = sum(chi_squared_map)
+ noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
+ log_likelihood = -0.5 * (chi_squared + noise_normalization)
+
+ return log_likelihood
+
+ def model_data_from_instance(self, instance):
+ """
+ To create the summed profile of all individual profiles in an instance, we can use a dictionary comprehension
+ to iterate over all profiles in the instance.
+ """
+ xvalues = np.arange(self.data.shape[0])
+
+ return sum([profile.model_data_from(xvalues=xvalues) for profile in instance])
+
+ def visualize(self, paths, instance, during_analysis):
+ """
+ This method is identical to the previous tutorial, except it now uses the `model_data_from_instance` method
+ to create the profile.
+ """
+ xvalues = np.arange(self.data.shape[0])
+
+ model_data = self.model_data_from_instance(instance=instance)
+
+ residual_map = self.data - model_data
+ chi_squared_map = (residual_map / self.noise_map) ** 2.0
+
+ """The visualizer now outputs images of the best-fit results to hard-disk (checkout `visualizer.py`)."""
+ plot_profile_1d(
+ xvalues=xvalues,
+ profile_1d=self.data,
+ title="Data",
+ ylabel="Data Values",
+ color="k",
+ output_path=paths.image_path,
+ output_filename="data",
+ )
+
+ plot_profile_1d(
+ xvalues=xvalues,
+ profile_1d=model_data,
+ title="Model Data",
+ ylabel="Model Data Values",
+ color="k",
+ output_path=paths.image_path,
+ output_filename="model_data",
+ )
+
+ plot_profile_1d(
+ xvalues=xvalues,
+ profile_1d=residual_map,
+ title="Residual Map",
+ ylabel="Residuals",
+ color="k",
+ output_path=paths.image_path,
+ output_filename="residual_map",
+ )
+
+ plot_profile_1d(
+ xvalues=xvalues,
+ profile_1d=chi_squared_map,
+ title="Chi-Squared Map",
+ ylabel="Chi-Squareds",
+ color="k",
+ output_path=paths.image_path,
+ output_filename="chi_squared_map",
+ )
+
+
+"""
+__Model Fit__
+
+Now lets run the non-linear search to get ourselves a `Result`.
+"""
+
+
+class Gaussian:
+ def __init__(
+ self,
+ centre=30.0, # <- **PyAutoFit** recognises these constructor arguments
+ normalization=1.0, # <- are the Gaussian`s model parameters.
+ sigma=5.0,
+ ):
+ """
+ Represents a 1D Gaussian profile.
+
+ This is a model-component of example models in the **HowToFit** lectures and is used to fit example datasets
+ via a non-linear search.
+
+ Parameters
+ ----------
+ centre
+ The x coordinate of the profile centre.
+ normalization
+ Overall normalization of the profile.
+ sigma
+ The sigma value controlling the size of the Gaussian.
+ """
+ self.centre = centre
+ self.normalization = normalization
+ self.sigma = sigma
+
+ def model_data_from(self, xvalues: np.ndarray):
+ """
+
+ Returns a 1D Gaussian on an input list of Cartesian x coordinates.
+
+ The input xvalues are translated to a coordinate system centred on the Gaussian, via its `centre`.
+
+ The output is referred to as the `model_data` to signify that it is a representation of the data from the
+ model.
+
+ Parameters
+ ----------
+ xvalues
+ The x coordinates in the original reference frame of the data.
+ """
+ transformed_xvalues = np.subtract(xvalues, self.centre)
+ return np.multiply(
+ np.divide(self.normalization, self.sigma * np.sqrt(2.0 * np.pi)),
+ np.exp(-0.5 * np.square(np.divide(transformed_xvalues, self.sigma))),
+ )
+
+
+class Exponential:
+ def __init__(
+ self,
+ centre=30.0, # <- **PyAutoFit** recognises these constructor arguments
+ normalization=1.0, # <- are the Exponential`s model parameters.
+ rate=0.01,
+ ):
+ """
+ Represents a 1D Exponential profile.
+
+ This is a model-component of example models in the **HowToFit** lectures and is used to fit example datasets
+ via a non-linear search.
+
+ Parameters
+ ----------
+ centre
+ The x coordinate of the profile centre.
+ normalization
+ Overall normalization of the profile.
+ ratw
+ The decay rate controlling has fast the Exponential declines.
+ """
+ self.centre = centre
+ self.normalization = normalization
+ self.rate = rate
+
+ def model_data_from(self, xvalues: np.ndarray):
+ """
+ Returns a 1D Gaussian on an input list of Cartesian x coordinates.
+
+ The input xvalues are translated to a coordinate system centred on the Gaussian, via its `centre`.
+
+ The output is referred to as the `model_data` to signify that it is a representation of the data from the
+ model.
+
+ Parameters
+ ----------
+ xvalues
+ The x coordinates in the original reference frame of the data.
+ """
+ transformed_xvalues = np.subtract(xvalues, self.centre)
+ return self.normalization * np.multiply(
+ self.rate, np.exp(-1.0 * self.rate * abs(transformed_xvalues))
+ )
+
+
+model = af.Collection(gaussian=af.Model(Gaussian), exponential=af.Model(Exponential))
+
+analysis = Analysis(data=data, noise_map=noise_map)
+
+search = af.Emcee(
+ name="tutorial_5_results_and_samples",
+ path_prefix=path.join("howtofit", "chapter_1"),
+)
+
+print(
+ """
+ The non-linear search has begun running.
+ Checkout the autofit_workspace/output/howtofit/tutorial_6__results_and_samples
+ folder for live output of the results.
+ This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
+ """
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+print("The search has finished run - you may now continue the notebook.")
+
+"""
+__Result__
+
+Here, we'll look in detail at what information is contained in the `Result`.
+
+It contains an `info` attribute which prints the result in readable format.
+"""
+print(result.info)
+
+"""
+__Samples__
+
+The result contains a `Samples` object, which contains all of the non-linear search samples.
+
+Each sample corresponds to a set of model parameters that were evaluated and accepted by our non linear search,
+in this example emcee.
+
+This also includes their log likelihoods, which are used for computing additional information about the model-fit,
+for example the error on every parameter.
+
+Our model-fit used the MCMC algorithm Emcee, so the `Samples` object returned is a `SamplesMCMC` object.
+"""
+samples = result.samples
+
+print("MCMC Samples: \n")
+print(samples)
+
+"""
+__Parameters__
+
+The parameters are stored as a list of lists, where:
+
+ - The outer list is the size of the total number of samples.
+ - The inner list is the size of the number of free parameters in the fit.
+"""
+samples = result.samples
+print("Sample 5's second parameter value (Gaussian -> normalization):")
+print(samples.parameter_lists[4][1])
+print("Sample 10`s third parameter value (Gaussian -> sigma)")
+print(samples.parameter_lists[9][2], "\n")
+
+"""
+__Figures of Merit__
+
+The Samples class also contains the log likelihood, log prior, log posterior and weight_list of every accepted sample,
+where:
+
+- The log likelihood is the value evaluated from the likelihood function (e.g. -0.5 * chi_squared + the noise
+normalized).
+
+- The log prior encodes information on how the priors on the parameters maps the log likelihood value to the log
+posterior value.
+
+- The log posterior is log_likelihood + log_prior.
+
+- The weight gives information on how samples should be combined to estimate the posterior. The weight values depend on
+the sampler used, for MCMC samples they are all 1 (e.g. all weighted equally).
+
+Lets inspect the last 10 values of each for the analysis.
+"""
+print("log(likelihood), log(prior), log(posterior) and weight of the tenth sample.")
+print(samples.log_likelihood_list[9])
+print(samples.log_prior_list[9])
+print(samples.log_posterior_list[9])
+print(samples.weight_list[9])
+
+"""
+__Instances__
+
+The `Samples` contains many results which are returned as an instance of the model, using the Python class structure
+of the model composition.
+
+For example, we can return the model parameters corresponding to the maximum log likelihood sample.
+"""
+max_lh_instance = samples.max_log_likelihood()
+
+print("Max Log Likelihood `Gaussian` Instance:")
+print("Centre = ", max_lh_instance.gaussian.centre)
+print("Normalization = ", max_lh_instance.gaussian.normalization)
+print("Sigma = ", max_lh_instance.gaussian.sigma, "\n")
+
+print("Max Log Likelihood Exponential Instance:")
+print("Centre = ", max_lh_instance.exponential.centre)
+print("Normalization = ", max_lh_instance.exponential.normalization)
+print("Sigma = ", max_lh_instance.exponential.rate, "\n")
+
+"""
+__Vectors__
+
+All results can alternatively be returned as a 1D vector of values, by passing `as_instance=False`:
+"""
+max_lh_vector = samples.max_log_likelihood(as_instance=False)
+print("Max Log Likelihood Model Parameters: \n")
+print(max_lh_vector, "\n\n")
+
+"""
+__Labels__
+
+Vectors return a lists of all model parameters, but do not tell us which values correspond to which parameters.
+
+The following quantities are available in the `Model`, where the order of their entries correspond to the parameters
+in the `ml_vector` above:
+
+ - `paths`: a list of tuples which give the path of every parameter in the `Model`.
+ - `parameter_names`: a list of shorthand parameter names derived from the `paths`.
+ - `parameter_labels`: a list of parameter labels used when visualizing non-linear search results (see below).
+
+"""
+model = samples.model
+
+print(model.paths)
+print(model.parameter_names)
+print(model.parameter_labels)
+print(model.model_component_and_parameter_names)
+print("\n")
+
+"""
+From here on, we will returned all results information as instances, but every method below can be returned as a
+vector via the `as_instance=False` input.
+
+__Posterior / PDF__
+
+The ``Result`` object contains the full posterior information of our non-linear search, which can be used for
+parameter estimation.
+
+The median pdf vector is available from the `Samples` object, which estimates the every parameter via 1D
+marginalization of their PDFs.
+"""
+median_pdf_instance = samples.median_pdf()
+
+print("Max Log Likelihood `Gaussian` Instance:")
+print("Centre = ", median_pdf_instance.gaussian.centre)
+print("Normalization = ", median_pdf_instance.gaussian.normalization)
+print("Sigma = ", median_pdf_instance.gaussian.sigma, "\n")
+
+print("Max Log Likelihood Exponential Instance:")
+print("Centre = ", median_pdf_instance.exponential.centre)
+print("Normalization = ", median_pdf_instance.exponential.normalization)
+print("Sigma = ", median_pdf_instance.exponential.rate, "\n")
+
+"""
+__Plot__
+
+Because results are returned as instances, it is straight forward to use them and their associated functionality
+to make plots of the results:
+"""
+model_gaussian = max_lh_instance.gaussian.model_data_from(
+ xvalues=np.arange(data.shape[0])
+)
+model_exponential = max_lh_instance.exponential.model_data_from(
+ xvalues=np.arange(data.shape[0])
+)
+model_data = model_gaussian + model_exponential
+
+plt.plot(range(data.shape[0]), data)
+plt.plot(range(data.shape[0]), model_data)
+plt.plot(range(data.shape[0]), model_gaussian, "--")
+plt.plot(range(data.shape[0]), model_exponential, "--")
+plt.title("Illustrative model fit to 1D `Gaussian` + Exponential profile data.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+__Errors__
+
+The samples include methods for computing the error estimates of all parameters, via 1D marginalization at an
+input sigma confidence limit.
+"""
+errors_at_upper_sigma_instance = samples.errors_at_upper_sigma(sigma=3.0)
+errors_at_lower_sigma_instance = samples.errors_at_lower_sigma(sigma=3.0)
+
+print("Upper Error values of Gaussian (at 3.0 sigma confidence):")
+print("Centre = ", errors_at_upper_sigma_instance.gaussian.centre)
+print("Normalization = ", errors_at_upper_sigma_instance.gaussian.normalization)
+print("Sigma = ", errors_at_upper_sigma_instance.gaussian.sigma, "\n")
+
+print("lower Error values of Gaussian (at 3.0 sigma confidence):")
+print("Centre = ", errors_at_lower_sigma_instance.gaussian.centre)
+print("Normalization = ", errors_at_lower_sigma_instance.gaussian.normalization)
+print("Sigma = ", errors_at_lower_sigma_instance.gaussian.sigma, "\n")
+
+"""
+They can also be returned at the values of the parameters at their error values:
+"""
+values_at_upper_sigma_instance = samples.values_at_upper_sigma(sigma=3.0)
+values_at_lower_sigma_instance = samples.values_at_lower_sigma(sigma=3.0)
+
+print("Upper Parameter values w/ error of Gaussian (at 3.0 sigma confidence):")
+print("Centre = ", values_at_upper_sigma_instance.gaussian.centre)
+print("Normalization = ", values_at_upper_sigma_instance.gaussian.normalization)
+print("Sigma = ", values_at_upper_sigma_instance.gaussian.sigma, "\n")
+
+print("lower Parameter values w/ errors of Gaussian (at 3.0 sigma confidence):")
+print("Centre = ", values_at_lower_sigma_instance.gaussian.centre)
+print("Normalization = ", values_at_lower_sigma_instance.gaussian.normalization)
+print("Sigma = ", values_at_lower_sigma_instance.gaussian.sigma, "\n")
+
+"""
+__PDF__
+
+The Probability Density Functions (PDF's) of the results can be plotted using the Emcee's visualization
+tool `corner.py`, which is wrapped via the `aplt.corner_cornerpy` function.
+"""
+aplt.corner_cornerpy(samples=result.samples)
+
+"""
+__Other Results__
+
+The samples contain many useful vectors, including the samples with the highest posterior values.
+"""
+max_log_posterior_instance = samples.max_log_posterior()
+
+print("Maximum Log Posterior Vector:")
+print("Centre = ", max_log_posterior_instance.gaussian.centre)
+print("Normalization = ", max_log_posterior_instance.gaussian.normalization)
+print("Sigma = ", max_log_posterior_instance.gaussian.sigma, "\n")
+
+
+"""
+All methods above are available as a vector:
+"""
+median_pdf_instance = samples.median_pdf(as_instance=False)
+values_at_upper_sigma = samples.values_at_upper_sigma(sigma=3.0, as_instance=False)
+values_at_lower_sigma = samples.values_at_lower_sigma(sigma=3.0, as_instance=False)
+errors_at_upper_sigma = samples.errors_at_upper_sigma(sigma=3.0, as_instance=False)
+errors_at_lower_sigma = samples.errors_at_lower_sigma(sigma=3.0, as_instance=False)
+
+"""
+__Sample Instance__
+
+A non-linear search retains every model that is accepted during the model-fit.
+
+We can create an instance of any lens model -- below we create an instance of the last accepted model.
+"""
+instance = samples.from_sample_index(sample_index=-1)
+
+print("Gaussian Instance of last sample")
+print("Centre = ", instance.gaussian.centre)
+print("Normalization = ", instance.gaussian.normalization)
+print("Sigma = ", instance.gaussian.sigma, "\n")
+
+"""
+__Bayesian Evidence__
+
+If a nested sampling `NonLinearSearch` is used, the evidence of the model is also available which enables Bayesian
+model comparison to be performed (given we are using Emcee, which is not a nested sampling algorithm, the log evidence
+is None).:
+"""
+log_evidence = samples.log_evidence
+
+"""
+__Derived Errors (PDF from samples)__
+
+Computing the errors of a quantity like the `sigma` of the Gaussian is simple, because it is sampled by the non-linear
+search. Thus, to get their errors above we used the `Samples` object to simply marginalize over all over parameters
+via the 1D Probability Density Function (PDF).
+
+Computing errors on derived quantities is more tricky, because they are not sampled directly by the non-linear search.
+For example, what if we want the error on the full width half maximum (FWHM) of the Gaussian? In order to do this
+we need to create the PDF of that derived quantity, which we can then marginalize over using the same function we
+use to marginalize model parameters.
+
+Below, we compute the FWHM of every accepted model sampled by the non-linear search and use this determine the PDF
+of the FWHM. When combining the FWHM's we weight each value by its `weight`. For Emcee, an MCMC algorithm, the
+weight of every sample is 1, but weights may take different values for other non-linear searches.
+
+In order to pass these samples to the function `marginalize`, which marginalizes over the PDF of the FWHM to compute
+its error, we also pass the weight list of the samples.
+
+(Computing the error on the FWHM could be done in much simpler ways than creating its PDF from the list of every
+sample. We chose this example for simplicity, in order to show this functionality, which can easily be extended to more
+complicated derived quantities.)
+"""
+fwhm_list = []
+
+for sample in samples.sample_list:
+ instance = sample.instance_for_model(model=samples.model)
+
+ sigma = instance.gaussian.sigma
+
+ fwhm = 2 * np.sqrt(2 * np.log(2)) * sigma
+
+ fwhm_list.append(fwhm)
+
+median_fwhm, lower_fwhm, upper_fwhm = af.marginalize(
+ parameter_list=fwhm_list, sigma=3.0, weight_list=samples.weight_list
+)
+
+print(f"FWHM = {median_fwhm} ({upper_fwhm} {lower_fwhm}")
+
+"""
+__Samples Filtering__
+
+Our samples object has the results for all three parameters in our model. However, we might only be interested in the
+results of a specific parameter.
+
+The basic form of filtering specifies parameters via their path, which was printed above via the model and is printed
+again below.
+"""
+samples = result.samples
+
+print("Parameter paths in the model which are used for filtering:")
+print(samples.model.paths)
+
+print("All parameters of the very first sample")
+print(samples.parameter_lists[0])
+
+samples = samples.with_paths([("gaussian", "centre")])
+
+print("All parameters of the very first sample (containing only the Gaussian centre.")
+print(samples.parameter_lists[0])
+
+print("Maximum Log Likelihood Model Instances (containing only the Gaussian centre):\n")
+print(samples.max_log_likelihood(as_instance=False))
+
+"""
+Above, we specified each path as a list of tuples of strings.
+
+This is how the source code internally stores the path to different components of the model, but it is not
+in-profile_1d with the PyAutoFIT API used to compose a model.
+
+We can alternatively use the following API:
+"""
+samples = result.samples
+
+samples = samples.with_paths(["gaussian.centre"])
+
+print("All parameters of the very first sample (containing only the Gaussian centre).")
+print(samples.parameter_lists[0])
+
+"""
+Above, we filtered the `Samples` but asking for all parameters which included the path ("gaussian", "centre").
+
+We can alternatively filter the `Samples` object by removing all parameters with a certain path. Below, we remove
+the Gaussian's `centre` to be left with 2 parameters; the `normalization` and `sigma`.
+"""
+samples = result.samples
+
+print("Parameter paths in the model which are used for filtering:")
+print(samples.model.paths)
+
+print("All parameters of the very first sample")
+print(samples.parameter_lists[0])
+
+samples = samples.without_paths(["gaussian.centre"])
+
+print(
+ "All parameters of the very first sample (containing only the Gaussian normalization and sigma)."
+)
+print(samples.parameter_lists[0])
+
+"""
+__Latex__
+
+If you are writing modeling results up in a paper, you can use inbuilt latex tools to create latex table
+code which you can copy to your .tex document.
+
+By combining this with the filtering tools below, specific parameters can be included or removed from the latex.
+
+Remember that the superscripts of a parameter are loaded from the config file `notation/label.yaml`, providing high
+levels of customization for how the parameter names appear in the latex table. This is especially useful if your model
+uses the same model components with the same parameter, which therefore need to be distinguished via superscripts.
+"""
+latex = af.text.Samples.latex(
+ samples=result.samples,
+ median_pdf_model=True,
+ sigma=3.0,
+ name_to_label=True,
+ include_name=True,
+ include_quickmath=True,
+ prefix="Example Prefix ",
+ suffix=" \\[-2pt]",
+)
+
+print(latex)
+
+"""
+Finish.
+"""
diff --git a/scripts/howtofit/chapter_1_introduction/tutorial_8_astronomy_example.py b/scripts/howtofit/chapter_1_introduction/tutorial_8_astronomy_example.py
index 5283ebb4..c596a4b2 100644
--- a/scripts/howtofit/chapter_1_introduction/tutorial_8_astronomy_example.py
+++ b/scripts/howtofit/chapter_1_introduction/tutorial_8_astronomy_example.py
@@ -1,1067 +1,1063 @@
-"""
-Tutorial 8: Astronomy Example
-=============================
-
-In this tutorial, we'll apply the tools we've learned in this chapter to tackle a read world problem and fit
-2D Hubble Space Telescope imaging data of galaxies.
-
-One of the most well-known results in astronomy is the Hubble sequence, which classifies galaxies based on their
-visual appearance. The sequence is divided into three broad groups:
-
-- **Early-type**: Galaxies that are round in shape with a smooth blob of light, often called a "bulge".
-
-- **Late-type**: Galaxies that are elliptical in shape with a flattened light distribution, often called a "disk".
-
-- **Irregular**: Galaxies that do not have a regular shape.
-
-Here is the Hubble Sequence, showing early-type galaxies on the left and late-type galaxies on the right:
-
-
-
-This example fits a Hubble Space Telescope image of a galaxy with two models: one representing bulge-like early-type
-galaxies and one representing disk-like late-type galaxies. This will help us determine whether the galaxy is an early
-or late-type galaxy.
-
-The aim of this example is to illustrate that everything you have learnt in this chapter can be applied to real-world
-problems, and to show you how many of the tools and challenges you have learnt about are used in practice.
-
-After fitting the Hubble Space Telescope data, we will explore some common model-fitting problems astronomers face
-when classifying galaxies and measuring their properties. This will be an open exercise for you to consider how these
-problems can be addressed.
-
-__Overview__
-
-In this tutorial, we will:
-
-- Use the tools from tutorials 1, 2, and 3 to set up a model that fits 2D astronomical images of galaxies with light
- profile models.
-
-- Encounter and discuss the challenges of model fitting described in tutorial 4, along with strategies to overcome
- these challenges.
-
-- Use the result properties introduced in tutorial 5 to interpret the model fit and determine whether the galaxy is an
- early or late-type galaxy.
-
- __Contents__
-
-- **Plot**: Convensional plotting functions for 2D data and grids of 2D coordinates.
-- **Data**: Load and plot Hubble Space Telescope imaging data of a galaxy.
-- **Mask**: Apply a mask to the data to remove regions of the image that are not relevant to the model fitting.
-- **PSF**: Load and plot the Point Spread Function (PSF) of the telescope.
-- **Grid**: Create a grid of (y,x) coordinates that overlap the observed galaxy data.
-- **Light Profiles**: Define light profile classes representing the light of galaxies.
-- **Model Data**: Create the model image of a galaxy by convolving its light profile with the PSF.
-- **Model**: Define a model with a light profile to fit the galaxy data.
-- **Analysis**: Define the log likelihood function, which compares the model image to the observed image.
-- **Model Fit**: Fit the model to the data and display the results.
-- **Result**: Interpret the model fit to determine whether the galaxy is an early or late-type galaxy.
-- **Bulgey**: Repeat the fit using a bulgey light profile to determine the galaxy's type.
-- **Model Mismatch**: Analyze the challenges from model mismatches in galaxy classification.
-- **Extensions**: Illustrate examples of how this problem can be extended and the challenges that arise.
-- **Chapter Wrap Up**: Summarize the completion of Chapter 1 and its applications to real astronomy.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-from os import path
-import numpy as np
-import matplotlib.pyplot as plt
-from scipy import signal
-from typing import Tuple
-
-import autofit as af
-
-"""
-__Plot__
-
-We will plot a lot of arrays of 2D data and grids of 2D coordinates in this example, so lets make a convenience
-functions.
-"""
-
-
-def plot_array(array, title=None, norm=None, filename=None):
- plt.imshow(array, norm=norm)
- plt.colorbar()
- plt.title(title)
- if filename is not None:
- plt.savefig(filename)
- plt.show()
- plt.clf()
- plt.close()
-
-
-def plot_grid(grid, title=None):
- plt.scatter(x=grid[:, :, 0], y=grid[:, :, 1], s=1)
- plt.title(title)
- plt.show()
- plt.clf()
- plt.close()
-
-
-"""
-__Data__
-
-First, let's load and plot Hubble Space Telescope imaging data of a galaxy. This data includes:
-
-1) The image of the galaxy, which is the data we'll fit.
-2) The noise in each pixel of this image, which will be used to evaluate the log likelihood.
-
-The noise-map has a few strange off-centre features which are an artefact of the telescope. Don't worry about these
-features.
-"""
-dataset_path = path.join("dataset", "howtofit", "chapter_1", "astro", "simple")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = np.load(file=path.join(dataset_path, "data.npy"))
-plot_array(array=data, title="Image of Galaxy")
-
-noise_map = np.load(file=path.join(dataset_path, "noise_map.npy"))
-plot_array(array=noise_map, title="Noise Map of Galaxy")
-
-"""
-__Mask__
-
-When fitting 2D imaging data, it is common to apply a mask which removes regions of the image that are not relevant to
-the model fitting.
-
-For example, when fitting the galaxy, we remove the edges of the image where the galaxy's light is not visible.
-
-We load and plot the mask below to show you how it is applied to the data, and we will use it in
-the `log_likelihood_function` below to ensure these regions are not fitted.
-"""
-mask = np.load(file=path.join(dataset_path, "mask.npy"))
-plot_array(array=mask, title="Mask of Galaxy")
-
-"""
-In the image of the galaxy a bright blob of light is clearly visible, which is the galaxy we'll fit with a model.
-
-__PSF__
-
-Another important component of imaging data is the Point Spread Function (PSF), which describes how the light from
-galaxies is blurred when it enters the Hubble Space Telescope.
-
-This blurring occurs due to diffraction as light passes through the HST's optics. The PSF is represented as a
-two-dimensional array, which acts as a 2D convolution kernel.
-
-When fitting the data and in the `log_likelihood_function` below, the PSF is used to create the model data. This
-demonstrates how an `Analysis` class can be extended to include additional steps in the model fitting process.
-"""
-psf = np.load(file=path.join(dataset_path, "psf.npy"))
-plot_array(array=psf, title="Point Spread Function of Galaxy ?")
-
-"""
-__Grid__
-
-To perform certain calculations, we need a grid of (x,y) coordinates that overlap the observed galaxy data.
-
-We create a 2D grid of coordinates where the origin is (0.0, 0.0) and each pixel is 0.05 in size, matching the
-resolution of our image data.
-
-This grid includes only (y,x) coordinates within the circular mask applied to the data, as we only need to perform
-calculations within this masked region.
-"""
-grid = np.load(file=path.join(dataset_path, "grid.npy"))
-
-plot_grid(
- grid=grid,
- title="Cartesian grid of (x,y) coordinates aligned with dataset",
-)
-
-"""
-__Light Profiles__
-
-Our galaxy model must describe the light of each galaxy, which we refer to as a "light profile." Below, we define two
-light profile classes named `LightBulgey` and `LightDisky`.
-
-These Python classes serve as the model components representing each galaxy's light, similar to the `Gaussian` class
-in previous tutorials. The `__init__` constructor's input parameters (e.g., `centre`, `axis_ratio`, `angle`) are the
-model parameters that the non-linear search will fit.
-
-These classes also include functions that create an image from the light profile based on an input grid of (x,y) 2D
-coordinates, which we will use to generate an image of a light profile. There is a lot of maths going on in these
-functions which understanding in detail isn't necessary for you to use **PyAutoFit**. So, if the code below looks
-complicated, don't worry about it!
-
-The Python code below uses Python's class inheritance functionality, which novice users may not be familiar with.
-Inheritance is the part of the code which reads `class LightProfile(GeometryProfile):`. This means that
-the `LightProfile` class inherits the functions and attributes of the `GeometryProfile` class.
-
-This is a common object-oriented programming feature in Python, but is not something you need to be familiar with to
-use **PyAutoFit**. If the code below is confusing, don't worry about it for now, it won't impact your ability to use
-**PyAutoFit**.
-"""
-
-
-class GeometryProfile:
- def __init__(
- self,
- centre: Tuple[float, float] = (0.0, 0.0),
- axis_ratio: float = 1.0,
- angle: float = 0.0,
- ):
- """
- Abstract base class for the geometry of a profile representing the light or mass of a galaxy.
-
- Using the centre, axis-ratio and position angle of the profile this class describes how to convert a
- (y,x) grid of Cartesian coordinates to the elliptical geometry of the profile.
-
- Parameters
- ----------
- centre
- The (y,x) coordinates of the profile centre.
- axis_ratio
- The axis-ratio of the ellipse (minor axis / major axis).
- angle
- The rotation angle in degrees counter-clockwise from the positive x-axis.
- """
-
- self.centre = centre
- self.axis_ratio = axis_ratio
- self.angle = angle
-
- def transformed_to_reference_frame_grid_from(self, grid: np.ndarray):
- """
- Transform a grid of (y,x) coordinates to the geometric reference frame of the profile via a translation using
- its `centre` and a rotation using its `angle`.
-
- This performs the following steps:
-
- 1) Translate the input (y,x) coordinates from the (0.0, 0.0) origin to the centre of the profile by subtracting
- the profile centre.
-
- 2) Compute the radial distance of every translated coordinate from the centre.
-
- 3) Rotate the coordinates from the above step counter-clockwise from the positive x-axis by the
- profile's `angle`.
-
- Parameters
- ----------
- grid
- The (y, x) coordinate grid in its original reference frame.
- """
-
- shifted_grid = grid - self.centre
- effective_radius = (
- (shifted_grid[:, :, 0] ** 2.0 + shifted_grid[:, :, 1] ** 2.0)
- ) ** 0.5
-
- theta_coordinate_to_profile = np.arctan2(
- shifted_grid[:, :, 1], shifted_grid[:, :, 0]
- ) - np.radians(self.angle)
-
- transformed_grid = np.zeros(grid.shape)
-
- transformed_grid[:, :, 0] = effective_radius * np.cos(
- theta_coordinate_to_profile
- )
- transformed_grid[:, :, 1] = effective_radius * np.sin(
- theta_coordinate_to_profile
- )
-
- return transformed_grid
-
- def rotated_grid_from_reference_frame_from(self, grid: np.ndarray) -> np.ndarray:
- """
- Rotate a grid of (y,x) coordinates which have been transformed to the elliptical reference frame of a profile
- back to the original unrotated coordinate frame.
-
- This performs the following steps:
-
- 1) Rotate the coordinates from the elliptical reference frame back to the original unrotated coordinate frame
- using the profile's `angle`.
-
- 2) Translate the coordinates from the centre of the profile back to the (0.0, 0.0) origin by adding the
- profile centre.
-
- Parameters
- ----------
- grid
- The (y, x) coordinates in the reference frame of an elliptical profile.
- """
- cos_angle = np.cos(np.radians(self.angle))
- sin_angle = np.sin(np.radians(self.angle))
-
- transformed_grid = np.zeros(grid.shape)
-
- transformed_grid[:, :, 0] = np.add(
- np.multiply(grid[:, :, 0], cos_angle),
- -np.multiply(grid[:, :, 1], sin_angle),
- )
- transformed_grid[:, :, 1] = np.add(
- np.multiply(grid[:, :, 0], sin_angle), np.multiply(grid[:, :, 1], cos_angle)
- )
-
- return transformed_grid
-
- def elliptical_radii_grid_from(self, grid: np.ndarray) -> np.ndarray:
- """
- Convert a grid of (y,x) coordinates to a grid of elliptical radii.
-
- Parameters
- ----------
- grid
- The (y, x) coordinates in the reference frame of the elliptical profile.
- """
-
- return (
- (grid[:, :, 1] ** 2.0) + (grid[:, :, 0] / self.axis_ratio) ** 2.0
- ) ** 0.5
-
-
-class LightProfile(GeometryProfile):
- def __init__(
- self,
- centre: Tuple[float, float] = (0.0, 0.0),
- axis_ratio: float = 1.0,
- angle: float = 0.0,
- intensity: float = 0.1,
- effective_radius: float = 0.6,
- ):
- """
- Abstract base class for a light profile, which describes the emission of a galaxy as a
- function of radius.
-
- Parameters
- ----------
- centre
- The (y,x) coordinates of the profile centre.
- axis_ratio
- The axis-ratio of the ellipse (minor axis / major axis).
- angle
- The rotation angle in degrees counter-clockwise from the positive x-axis.
- intensity
- Overall intensity normalisation of the light profile.
- effective_radius
- The circular radius containing half the light of this profile.
- """
-
- super().__init__(centre=centre, axis_ratio=axis_ratio, angle=angle)
-
- self.intensity = intensity
- self.effective_radius = effective_radius
-
-
-class LightBulgey(LightProfile):
- def __init__(
- self,
- centre: Tuple[float, float] = (0.0, 0.0),
- axis_ratio: float = 1.0,
- angle: float = 0.0,
- intensity: float = 0.1,
- effective_radius: float = 0.6,
- ):
- """
- The De Vaucouleurs light profile often used in Astronomy to represent the bulge of galaxies.
-
- Parameters
- ----------
- centre
- The (y,x) coordinates of the profile centre.
- axis_ratio
- The axis-ratio of the ellipse (minor axis / major axis).
- angle
- The rotation angle in degrees counter-clockwise from the positive x-axis.
- intensity
- Overall intensity normalisation of the light profile.
- effective_radius
- The circular radius containing half the light of this profile.
- """
-
- super().__init__(
- centre=centre,
- axis_ratio=axis_ratio,
- angle=angle,
- intensity=intensity,
- effective_radius=effective_radius,
- )
-
- def image_from_grid(self, grid: np.ndarray) -> np.ndarray:
- """
- Returns the image of the De Vaucouleurs light profile on a grid of Cartesian (y,x) coordinates, which are
- first translated to the profile's reference frame.
-
- Parameters
- ----------
- grid
- The (y, x) coordinates where the image is computed.
- """
- grid_transformed = self.transformed_to_reference_frame_grid_from(grid=grid)
- grid_elliptical_radii = self.elliptical_radii_grid_from(grid=grid_transformed)
-
- return self.intensity * np.exp(
- -7.66924
- * ((grid_elliptical_radii / self.effective_radius) ** (1.0 / 7.66924) - 1.0)
- )
-
-
-class LightDisky(LightProfile):
- def __init__(
- self,
- centre: Tuple[float, float] = (0.0, 0.0),
- axis_ratio: float = 1.0,
- angle: float = 0.0,
- intensity: float = 0.1,
- effective_radius: float = 0.6,
- ):
- """
- The Exponential light profile often used in Astronomy to represent the disk of galaxies.
-
- Parameters
- ----------
- centre
- The (y,x) coordinates of the profile centre.
- axis_ratio
- The axis-ratio of the ellipse (minor axis / major axis).
- angle
- The rotation angle in degrees counter-clockwise from the positive x-axis.
- intensity
- Overall intensity normalisation of the light profile.
- effective_radius
- The circular radius containing half the light of this profile.
- """
-
- super().__init__(
- centre=centre,
- axis_ratio=axis_ratio,
- angle=angle,
- intensity=intensity,
- effective_radius=effective_radius,
- )
-
- def image_from_grid(self, grid: np.ndarray) -> np.ndarray:
- """
- Returns the image of the light profile on a grid of Cartesian (y,x) coordinates.
-
- Parameters
- ----------
- grid
- The (y, x) coordinates where the image is computed.
- """
- grid_transformed = self.transformed_to_reference_frame_grid_from(grid=grid)
- grid_elliptical_radii = self.elliptical_radii_grid_from(grid=grid_transformed)
-
- return self.intensity * np.exp(
- -1.67838
- * ((grid_elliptical_radii / self.effective_radius) ** (1.0 / 1.67838) - 1.0)
- )
-
-
-"""
-Here is an example of an image of a `LightDisky` profile, using the `image_from_grid` method and the
-grid aligned with the image of the galaxy.
-"""
-light_profile = LightDisky(
- centre=(0.01, 0.01), axis_ratio=0.7, angle=45.0, intensity=1.0, effective_radius=2.0
-)
-light_image = light_profile.image_from_grid(grid=grid)
-
-plot_array(array=light_image, title="Image of an Exponential light profile.")
-
-"""
-__Model Data__
-
-To produce the `model_data`, we now convolve the overall image with the Point Spread Function (PSF) of our observations.
-This blurs the image to simulate the effects of the telescope optics and pixelization used to capture the image.
-"""
-model_data = signal.convolve2d(light_image, psf, mode="same")
-
-plot_array(array=model_data, title="Model Data of the Light Profile.")
-
-"""
-By subtracting the model image from the data, we create a 2D residual map, similar to the residual maps we made in
-the 1D Gaussian examples but now for 2D imaging data. It's evident that the random model we used here does not fit
-the galaxy well.
-"""
-residual_map = data - model_data
-
-plot_array(array=residual_map, title="Residual Map of fit")
-
-"""
-Just like in the 1D `Gaussian` fitting examples, we can use the noise map to compute the normalized residuals and
-chi-squared map for the model.
-"""
-normalized_residual_map = residual_map / noise_map
-chi_squared_map = (normalized_residual_map) ** 2.0
-
-plot_array(
- array=normalized_residual_map,
- title="Normalized Residual Map of fit",
-)
-plot_array(array=chi_squared_map, title="Chi Squared Map of fit")
-
-"""
-Finally, we compute the `log_likelihood` of this model, which will be used next to fit the model to the data using
-a non-linear search.
-"""
-chi_squared = np.sum(chi_squared_map)
-noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
-
-log_likelihood = -0.5 * (chi_squared + noise_normalization)
-
-print(log_likelihood)
-
-"""
-__Model__
-
-We now define a model where one of the light profiles defined earlier is encapsulated within a `Model` object.
-This allows us to treat the light profile as a model component that can be used in conjunction with other components.
-
-Although we are currently using only one light profile in this example, we structure it within a `Collection` for
-potential extension with multiple light profiles.
-"""
-model = af.Collection(light_0=af.Model(LightDisky))
-
-"""
-The `model` operates the same as the model components we've utilized previously.
-
-Fitting 2D data is more time-consuming than 1D Gaussian data. I therefore employ the prior tuning method described in
-tutorial 4 to speed up the fit, by adjusting the priors to approximate values close to the correct solution.
-This adjustment is based on thorough fits that were performed using a slow and detailed search in order to locate
-the global maxima of the likelihood.
-
-Additionally, I fix the center of the light profile to (0.0, 0.0), where it visually appears to be located.
-This reduces the number of free parameters and simplifies the complexity of the non-linear search.
-"""
-model.light_0.centre_0 = 0.0
-model.light_0.centre_1 = 0.0
-model.light_0.angle = af.UniformPrior(lower_limit=100.0, upper_limit=120.0)
-model.light_0.axis_ratio = af.UniformPrior(lower_limit=0.6, upper_limit=0.8)
-model.light_0.effective_radius = af.UniformPrior(lower_limit=0.0, upper_limit=1.0)
-
-"""
-The model info contains information on all of the model components and priors, including the updates above.
-"""
-print(model.info)
-
-"""
-__Analysis__
-
-We now define the `Analysis` class for this astronomy example, which will fit the `model` to the image data.
-
-Checkout all the docstrings in this class for details on how the fit is performed to 2D imaging data, including
-the role of the mask and PSF.
-"""
-
-
-class Analysis(af.Analysis):
- def __init__(
- self,
- data: np.ndarray,
- noise_map: np.ndarray,
- psf: np.ndarray,
- grid: np.ndarray,
- mask: np.ndarray,
- ):
- """
- The analysis class for the **PyAutoFit** example Astronomy project on galaxy fitting.
-
- This class contains imaging data of a galaxy and it fits it with a model which represents the light profile of
- the galaxy.
-
- The imaging data contains:
-
- 1) An image of the galaxy.
-
- 2) The noise in every pixel of that image.
-
- 3) The Point Spread Function (PSF) describing how the optics of the telescope blur the image.
-
- 4) A (y,x) grid of coordinates describing the locations of the image pixels in a unit system, which are used
- to perform calculations.
-
- 5) A mask that removes certain image pixels from the analysis.
-
- This project is a scaled down version of the Astronomy project **PyAutoGalaxy**, which is one of the
- original projects from which PyAutoFit is an offshoot!
-
- https://github.com/Jammy2211/PyAutoGalaxy
-
- Parameters
- ----------
- data
- The image containing the observation of the galaxy that is fitted.
- noise_map
- The RMS noise values of the image data, which is folded into the log likelihood calculation.
- psf
- The Point Spread Function of the telescope, which describes how the telescope blurs the image.
- grid
- The (y, x) coordinates of the image from which the calculation is performed and model image is
- computed using.
- mask
- The 2D mask that is applied to the image data.
- """
-
- super().__init__()
-
- self.data = data
- self.noise_map = noise_map
- self.psf = psf
- self.grid = grid
- self.mask = mask
-
- def log_likelihood_function(self, instance) -> float:
- """
- The `log_likelihood_function` of the galaxy example, which performs the following step:
-
- 1) Using the model passed into the function (whose parameters are set via the non-linear search) create
- a model image of this galaxy using the light profiles in the `instance`.
-
- 2) Convolve this model image with the Point Spread Function of the telescope, ensuring the telescope optics
- are included in the model image.
-
- 3) Subtract this model image from the data and compute its residuals, chi-squared and likelihood via the
- noise map.
-
- 4) Apply the mask to all these quantities, ensuring the edges of the galaxy are omitted from the fit.
-
- Parameters
- ----------
- instance
- An instance of the model set via the non-linear search.
-
- Returns
- -------
- float
- The log likelihood value of this particular model.
- """
-
- """
- The 'instance' that comes into this method contains the light profiles we setup in the `Model` and `Collection`,
- which can be seen by uncommenting the code below.
- """
-
- # print("Model Instance:")
- # print("Light Profile = ", instance.light)
- # print("Light Profile Centre= ", instance.light.centre)
-
- """
- Generate the model data from the instance using the functions of the light profile class above.
-
- See the docstring of the `model_data_from_instance` function for a description of how this works.
- """
-
- model_data = self.model_data_from_instance(instance=instance)
-
- """
- In this context of fitting 2D astronomical imaging data, the calculation of residual-map,
- normalized residual-map, chi-squared-map, and likelihood differs from previous examples such as fitting 1D
- Gaussians.
-
- The main difference lies in the incorporation of a mask, which excludes regions where the galaxy is not
- visible, typically the edges.
-
- To ensure that the mask influences these calculations correctly, we use the `where` parameter of numpy.
- This parameter allows us to compute the chi-squared and likelihood only where the mask indicates valid
- data (where the mask is 0, indicating unmasked regions). This approach ensures that the analysis focuses
- exclusively on the relevant parts of the galaxy image where meaningful comparisons between the model and
- data can be made.
- """
- residual_map = np.subtract(
- self.data,
- model_data,
- out=np.zeros_like(data),
- where=np.asarray(self.mask) == 0,
- )
-
- normalized_residual_map = np.divide(
- residual_map,
- self.noise_map,
- out=np.zeros_like(residual_map),
- where=np.asarray(self.mask) == 0,
- )
-
- chi_squared_map = np.square(
- np.divide(
- residual_map,
- self.noise_map,
- out=np.zeros_like(residual_map),
- where=np.asarray(mask) == 0,
- )
- )
-
- chi_squared = float(np.sum(chi_squared_map[np.asarray(self.mask) == 0]))
- noise_normalization = float(
- np.sum(np.log(2 * np.pi * noise_map[np.asarray(mask) == 0] ** 2.0))
- )
-
- log_likelihood = -0.5 * (chi_squared + noise_normalization)
-
- return log_likelihood
-
- def model_data_from_instance(self, instance):
- """
- Create the image of a galaxy, including blurring due to the Point Spread Function of the telescope.
-
- For the purpose of illustrating **PyAutoFit** you do not need to understand what this function is doing, the
- main thing to note is that it allows us to create a model image of a galaxy.
-
- Nevertheless, if you are curious, it inputs the (y,x) Cartesian grids into the light profiles and evaluates
- the image of the galaxy. Multiple light profiles may be summed to create the overall model image that is fitted.
-
- The mask is used to zero all values of the model image that are not included in the mask.
-
- Parameters
- ----------
- instance
- An instance of the model set via the non-linear search.
-
- Returns
- -------
- image
- The image of this galaxy light profile model.
- """
-
- overall_image = np.zeros(self.data.shape)
-
- for light_profile in instance:
- overall_image += light_profile.image_from_grid(grid=self.grid)
-
- model_data = signal.convolve2d(overall_image, self.psf, mode="same")
-
- model_data[self.mask == 1] = 0.0
-
- return model_data
-
-
-"""
-__Model Fit__
-
-We have successfully composed a model and analysis class for the astronomy example.
-
-We can now fit it to the data using a search. We use the same method as previous examples, nested sampling
-algorithm Dynesty.
-"""
-search = af.DynestyStatic(
- nlive=500,
- sample="rwalk", # This makes dynesty run faster, don't worry about what it means for now!
-)
-
-analysis = Analysis(data=data, noise_map=noise_map, psf=psf, grid=grid, mask=mask)
-
-print(
- """
- The non-linear search has begun running.
- This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
- """
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-print("The search has finished run - you may now continue the notebook.")
-
-
-"""
-__Result__
-
-The `result` object provides a concise summary of the outcomes from the non-linear search. While many of these values
-may not be directly interpretable for non-astronomers, some are meaningful even without specialized knowledge.
-
-For instance, the `angle` parameter indicates the rotation of the galaxy's light profile counterclockwise from the
-x-axis. In our fit, this value should approximate 110.0 degrees, aligning with the observed orientation of the
-galaxy in the image.
-
-Similarly, the `axis-ratio` represents the ratio of the minor axis to the major axis of the galaxy's light profile.
-A value near 0.7 is expected, reflecting the elliptical shape typical of galaxies.
-
-This illustrates why we perform model-fitting, we can take complex data and infer simple, interpretable properties
-from it which provide insight into the physical processes generating the data. This is the core goal of the scientific
-method and the use of models to explain observations.
-"""
-print(result.info)
-
-"""
-The best way to assess the quality of the fit is to use the visualization techniques introduced in previous tutorials.
-
-For 2D data, we visualize the model image, residual-map, normalized residual-map, and chi-squared-map as 2D arrays.
-These quantities still have the same meanings as their 1D counterparts. For example, normalized residual-map values
-still represent the sigma values of the residuals, with values greater than 3.0 indicating a poor fit to the data.
-"""
-model_data = analysis.model_data_from_instance(
- instance=result.max_log_likelihood_instance
-)
-
-residual_map = np.subtract(
- data, model_data, out=np.zeros_like(data), where=np.asarray(mask) == 0
-)
-
-normalized_residual_map = np.divide(
- residual_map,
- noise_map,
- out=np.zeros_like(residual_map),
- where=np.asarray(mask) == 0,
-)
-
-chi_squared_map = np.square(
- np.divide(
- residual_map,
- noise_map,
- out=np.zeros_like(residual_map),
- where=np.asarray(mask) == 0,
- )
-)
-
-plot_array(
- array=model_data,
- title="Model Data of the Light Profile.",
-)
-plot_array(
- array=residual_map,
- title="Residual Map of fit",
-)
-plot_array(
- array=normalized_residual_map,
- title="Normalized Residual Map of fit",
-)
-plot_array(
- array=chi_squared_map,
- title="Chi Squared Map of fit",
-)
-
-"""
-__Bulgey__
-
-The fit above utilized the disky light profile, which is typically suitable for disk-like late-type galaxies.
-
-Now, we will repeat the fit using the bulgey light profile, which is more suitable for bulge-like early-type galaxies.
-
-The fit with the higher log likelihood will provide insight into whether the galaxy is more likely to be an early-type
-or a late-type galaxy.
-"""
-result_disk = result
-
-model = af.Collection(light_0=af.Model(LightBulgey))
-
-# model.light_0.centre_0 = 0.0
-# model.light_0.centre_1 = 0.0
-# model.light_0.axis_ratio = af.UniformPrior(lower_limit=0.7, upper_limit=1.0)
-
-print(
- """
- The non-linear search has begun running.
- This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
- """
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-print("The search has finished run - you may now continue the notebook.")
-
-"""
-Print the result info of the bulgey fit.
-"""
-print(result.info)
-
-"""
-We perform the same visualization of the model image, residual-map, normalized residual-map, and chi-squared-map as
-before.
-
-The model image of the bulgey profile provides a better fit to the data than the disky profile, as the residuals are
-lower and the chi-squared-map values are closer to 0.0.
-
-This suggests that the galaxy is more likely to be an early-type galaxy with a bulge-like light profile.
-"""
-
-model_data = analysis.model_data_from_instance(
- instance=result.max_log_likelihood_instance
-)
-
-residual_map = np.subtract(
- data, model_data, out=np.zeros_like(data), where=np.asarray(mask) == 0
-)
-
-normalized_residual_map = np.divide(
- residual_map,
- noise_map,
- out=np.zeros_like(residual_map),
- where=np.asarray(mask) == 0,
-)
-
-chi_squared_map = np.square(
- np.divide(
- residual_map,
- noise_map,
- out=np.zeros_like(residual_map),
- where=np.asarray(mask) == 0,
- )
-)
-
-plot_array(
- array=model_data,
- title="Model Data of the Light Profile.",
-)
-plot_array(
- array=residual_map,
- title="Residual Map of fit",
-)
-plot_array(
- array=normalized_residual_map,
- title="Normalized Residual Map of fit",
-)
-plot_array(
- array=chi_squared_map,
- title="Chi Squared Map of fit",
-)
-
-"""
-To make certain of our interpretation, we should compare the log likelihoods of the two fits.
-
-The fit with the highest log likelihood is the preferred model, which (provided your non-linear search sampled
-parameter space accurately), is the bulgey profile.
-
-Therefore, the galaxy is likely an early-type galaxy with a bulge-like light profile.
-"""
-print("Disk Model Log Likelihood:")
-print(result_disk.log_likelihood)
-print("Bulge Model Log Likelihood:")
-print(result.log_likelihood)
-
-"""
-__Model Mismatch__
-
-The analysis above allowed us to determine whether the galaxy is more likely to be an early-type or late-type galaxy.
-
-However, after fitting the bulgey profile, you may not of expected it to be the highest log likelihood model. The
-model gave a relatively poor fit, with significant residuals and chi-squared values. It just turns out that
-the disky profile gave an even worse fit!
-
-This reflected the notion of "model mismatch" that we discussed in tutorial 4. One of the challenges of model fitting
-is that you may not have a model that is a brilliant representation of the data, and your search is successfully
-locating the global maxima even though the fit looks visibly poor.
-
-In Astronomy, what a scientist would do next is update their model to try and improve the fit. For example, they
-may extend the model to contain both the bulgey and disky profile, allowing the model to fit both components of the
-galaxy simultaneously.
-
-There are a whole range of approaches that Astronomers take to improve the model, which include fitting the
-galaxy with hundreds of 2D Gaussians, fitting it with a pixel grid of light and decomposing it into what
-are called basis functions. These approaches are not relevent to your understanding of **PyAutoFit**, but they
-are all implemented in **PyAutoGalaxy** via the **PyAutoFit** API for model composition.
-
-This is another great example of how **PyAutoFit** can be used to perform complex model-fitting tasks.
-
-__Extensions__
-
-To conclude, I will illustrate some of the extensions that can be made to this example and the challenges that arise
-when fitting more complex models.
-
-I don't provide code examples of how to tackle these extensions with model-fitting, but I encourage you to think about
-how **PyAutoFit** could be used to address these problems and go ahead and give them a go yourself if you're feeling
-adventurous!
-
-**Multiple Components**:
-
-The model above fitted a single light profile to the galaxy, which turned out to be a bulge-like component.
-
-However, galaxies often have multiple components, for example a disk and bulge, but also structures such as a bar or
-spiral arms. Here is a galaxy with a bulge, disk, bar and spiral arms:
-
-
-
-To model this galaxy, we would add many different Python classes with their own parameters and light profiles
-representing each component of the galaxy. They would then be combined into a single model, whose model image
-would be the summed image of each component, the total number of parameters would be in the twenties or thirties
-and the degeneracies between the parameters would be challenging to sample accurately.
-
-Here is a snippet of rougyly what our model composition would look like:
-
-model = af.Collection(
- bulge=LightBulge,
- disk=LightDisk,
- bar=LightBar,
- spiral_arms=LightSpiralArms
-)
-
-This is a great example of how quickly model-fitting can become complex and how keeping track of the speed and
-efficiency of the non-linear search becomes crucial. Furthermore, with so many different light profiles to fit,
-**PyAutoFit**'s model composition API becomes invaluable.
-
-
-**Multiple Galaxies**:
-
-There is no reason our imaging data need contain only one galaxy. The data could include multiple galaxies, each
-of which we want to fit with its own light profile.
-
-Two galaxies close together are called galaxy mergers, and here is a beautiful example of a pair of merging galaxies:
-
-
-
-This would pretty much double the model complexity, as each galaxy would have its own model and light profile.
-All the usual issues then become doubly important, such as ensuring the likelihood function is efficient, that
-the search can sample parameter space accurately and that the results are interpreted correctly.
-
-The model composition would again change, and might look something like:
-
-model = af.Collection(
- bulge_0=LightBulge,
- disk_0=LightDisk,
- bulge_1=LightBulge,
- disk_1=LightDisk
-)
-
-model.bulge_0.centre_0 = 2.0
-model.bulge_0.centre_1 = 0.0
-model.disk_0.centre_0 = 2.0
-model.disk_0.centre_1 = 0.0
-
-model.bulge_1.centre_0 = -2.0
-model.bulge_1.centre_1 = 0.0
-model.disk_1.centre_0 = -2.0
-model.disk_1.centre_1 = 0.0
-
-In order to keep the model slightly more simple, the centres of the light profiles have been fixed to where
-they peak in the image.
-
-**PyAutoFit** extensible model composition API actually has much better tools for composing complex models like this
-than the example above. You can find a concise run through of these in the model cookbook, but they will
-also be the focus on a tutorial in the next chapter of the **HowToFit** lectures.
-
-
-**Multiple Wavelengths**:
-
-Galaxies emit light in many wavelengths, for example ultraviolet, optical, infrared, radio and X-ray. Each wavelength
-provides different information about the galaxy, for example the ultraviolet light tells us about star formation,
-the optical light about the stars themselves and the infrared about dust.
-
-The image below shows observations of the same galaxy at different wavelengths:
-
-
-
-In tutorial 6, we learn how to perform fits to multiple datasets simultaneously, and how to change the model
-parameterization to account for the variation across datasets.
-
-Multi-wavelength modeling of galaxies is a great example of where this is useful, as it allows us to fit the galaxy
-with certain parameters shared across all wavelengths (e.g., its centre) and other parameters varied (e.g., its
-intensity and effective radius). A child project of **PyAutoFit**, called **PyAutoGalaxy**, uses this functionality
-to achieve exactly this.
-
-__Chapter Wrap Up__
-
-We have now completed the first chapter of **HowToFit**, which has taught you the basics of model-fitting.
-
-Its now time you take everything you've learnt and apply it to your own model-fitting problem. Think carefully
-about the key concepts of this chapter, for example how to compose a model, how to create an analysis class and
-how to overcome challenges that arise when fitting complex models.
-
-Once you are familiar with these concepts, and confident that you have some simple model-fitting problems under
-your belt, you should move on to the next chapter. This covers how to build a scientific workflow around your
-model-fitting, so that you can begin to scale up your model-fitting to more complex models, larger datasets and
-more difficult problems. Checkout the `start_here.ipynb` notebook in chapter 2 to get started!
-"""
+"""
+Tutorial 8: Astronomy Example
+=============================
+
+In this tutorial, we'll apply the tools we've learned in this chapter to tackle a read world problem and fit
+2D Hubble Space Telescope imaging data of galaxies.
+
+One of the most well-known results in astronomy is the Hubble sequence, which classifies galaxies based on their
+visual appearance. The sequence is divided into three broad groups:
+
+- **Early-type**: Galaxies that are round in shape with a smooth blob of light, often called a "bulge".
+
+- **Late-type**: Galaxies that are elliptical in shape with a flattened light distribution, often called a "disk".
+
+- **Irregular**: Galaxies that do not have a regular shape.
+
+Here is the Hubble Sequence, showing early-type galaxies on the left and late-type galaxies on the right:
+
+
+
+This example fits a Hubble Space Telescope image of a galaxy with two models: one representing bulge-like early-type
+galaxies and one representing disk-like late-type galaxies. This will help us determine whether the galaxy is an early
+or late-type galaxy.
+
+The aim of this example is to illustrate that everything you have learnt in this chapter can be applied to real-world
+problems, and to show you how many of the tools and challenges you have learnt about are used in practice.
+
+After fitting the Hubble Space Telescope data, we will explore some common model-fitting problems astronomers face
+when classifying galaxies and measuring their properties. This will be an open exercise for you to consider how these
+problems can be addressed.
+
+__Overview__
+
+In this tutorial, we will:
+
+- Use the tools from tutorials 1, 2, and 3 to set up a model that fits 2D astronomical images of galaxies with light
+ profile models.
+
+- Encounter and discuss the challenges of model fitting described in tutorial 4, along with strategies to overcome
+ these challenges.
+
+- Use the result properties introduced in tutorial 5 to interpret the model fit and determine whether the galaxy is an
+ early or late-type galaxy.
+
+ __Contents__
+
+- **Plot**: Convensional plotting functions for 2D data and grids of 2D coordinates.
+- **Data**: Load and plot Hubble Space Telescope imaging data of a galaxy.
+- **Mask**: Apply a mask to the data to remove regions of the image that are not relevant to the model fitting.
+- **PSF**: Load and plot the Point Spread Function (PSF) of the telescope.
+- **Grid**: Create a grid of (y,x) coordinates that overlap the observed galaxy data.
+- **Light Profiles**: Define light profile classes representing the light of galaxies.
+- **Model Data**: Create the model image of a galaxy by convolving its light profile with the PSF.
+- **Model**: Define a model with a light profile to fit the galaxy data.
+- **Analysis**: Define the log likelihood function, which compares the model image to the observed image.
+- **Model Fit**: Fit the model to the data and display the results.
+- **Result**: Interpret the model fit to determine whether the galaxy is an early or late-type galaxy.
+- **Bulgey**: Repeat the fit using a bulgey light profile to determine the galaxy's type.
+- **Model Mismatch**: Analyze the challenges from model mismatches in galaxy classification.
+- **Extensions**: Illustrate examples of how this problem can be extended and the challenges that arise.
+- **Chapter Wrap Up**: Summarize the completion of Chapter 1 and its applications to real astronomy.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+from os import path
+import numpy as np
+import matplotlib.pyplot as plt
+from scipy import signal
+from typing import Tuple
+
+import autofit as af
+
+"""
+__Plot__
+
+We will plot a lot of arrays of 2D data and grids of 2D coordinates in this example, so lets make a convenience
+functions.
+"""
+
+
+def plot_array(array, title=None, norm=None, filename=None):
+ plt.imshow(array, norm=norm)
+ plt.colorbar()
+ plt.title(title)
+ if filename is not None:
+ plt.savefig(filename)
+ plt.show()
+ plt.clf()
+ plt.close()
+
+
+def plot_grid(grid, title=None):
+ plt.scatter(x=grid[:, :, 0], y=grid[:, :, 1], s=1)
+ plt.title(title)
+ plt.show()
+ plt.clf()
+ plt.close()
+
+
+"""
+__Data__
+
+First, let's load and plot Hubble Space Telescope imaging data of a galaxy. This data includes:
+
+1) The image of the galaxy, which is the data we'll fit.
+2) The noise in each pixel of this image, which will be used to evaluate the log likelihood.
+
+The noise-map has a few strange off-centre features which are an artefact of the telescope. Don't worry about these
+features.
+"""
+dataset_path = path.join("dataset", "howtofit", "chapter_1", "astro", "simple")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = np.load(file=path.join(dataset_path, "data.npy"))
+plot_array(array=data, title="Image of Galaxy")
+
+noise_map = np.load(file=path.join(dataset_path, "noise_map.npy"))
+plot_array(array=noise_map, title="Noise Map of Galaxy")
+
+"""
+__Mask__
+
+When fitting 2D imaging data, it is common to apply a mask which removes regions of the image that are not relevant to
+the model fitting.
+
+For example, when fitting the galaxy, we remove the edges of the image where the galaxy's light is not visible.
+
+We load and plot the mask below to show you how it is applied to the data, and we will use it in
+the `log_likelihood_function` below to ensure these regions are not fitted.
+"""
+mask = np.load(file=path.join(dataset_path, "mask.npy"))
+plot_array(array=mask, title="Mask of Galaxy")
+
+"""
+In the image of the galaxy a bright blob of light is clearly visible, which is the galaxy we'll fit with a model.
+
+__PSF__
+
+Another important component of imaging data is the Point Spread Function (PSF), which describes how the light from
+galaxies is blurred when it enters the Hubble Space Telescope.
+
+This blurring occurs due to diffraction as light passes through the HST's optics. The PSF is represented as a
+two-dimensional array, which acts as a 2D convolution kernel.
+
+When fitting the data and in the `log_likelihood_function` below, the PSF is used to create the model data. This
+demonstrates how an `Analysis` class can be extended to include additional steps in the model fitting process.
+"""
+psf = np.load(file=path.join(dataset_path, "psf.npy"))
+plot_array(array=psf, title="Point Spread Function of Galaxy ?")
+
+"""
+__Grid__
+
+To perform certain calculations, we need a grid of (x,y) coordinates that overlap the observed galaxy data.
+
+We create a 2D grid of coordinates where the origin is (0.0, 0.0) and each pixel is 0.05 in size, matching the
+resolution of our image data.
+
+This grid includes only (y,x) coordinates within the circular mask applied to the data, as we only need to perform
+calculations within this masked region.
+"""
+grid = np.load(file=path.join(dataset_path, "grid.npy"))
+
+plot_grid(
+ grid=grid,
+ title="Cartesian grid of (x,y) coordinates aligned with dataset",
+)
+
+"""
+__Light Profiles__
+
+Our galaxy model must describe the light of each galaxy, which we refer to as a "light profile." Below, we define two
+light profile classes named `LightBulgey` and `LightDisky`.
+
+These Python classes serve as the model components representing each galaxy's light, similar to the `Gaussian` class
+in previous tutorials. The `__init__` constructor's input parameters (e.g., `centre`, `axis_ratio`, `angle`) are the
+model parameters that the non-linear search will fit.
+
+These classes also include functions that create an image from the light profile based on an input grid of (x,y) 2D
+coordinates, which we will use to generate an image of a light profile. There is a lot of maths going on in these
+functions which understanding in detail isn't necessary for you to use **PyAutoFit**. So, if the code below looks
+complicated, don't worry about it!
+
+The Python code below uses Python's class inheritance functionality, which novice users may not be familiar with.
+Inheritance is the part of the code which reads `class LightProfile(GeometryProfile):`. This means that
+the `LightProfile` class inherits the functions and attributes of the `GeometryProfile` class.
+
+This is a common object-oriented programming feature in Python, but is not something you need to be familiar with to
+use **PyAutoFit**. If the code below is confusing, don't worry about it for now, it won't impact your ability to use
+**PyAutoFit**.
+"""
+
+
+class GeometryProfile:
+ def __init__(
+ self,
+ centre: Tuple[float, float] = (0.0, 0.0),
+ axis_ratio: float = 1.0,
+ angle: float = 0.0,
+ ):
+ """
+ Abstract base class for the geometry of a profile representing the light or mass of a galaxy.
+
+ Using the centre, axis-ratio and position angle of the profile this class describes how to convert a
+ (y,x) grid of Cartesian coordinates to the elliptical geometry of the profile.
+
+ Parameters
+ ----------
+ centre
+ The (y,x) coordinates of the profile centre.
+ axis_ratio
+ The axis-ratio of the ellipse (minor axis / major axis).
+ angle
+ The rotation angle in degrees counter-clockwise from the positive x-axis.
+ """
+
+ self.centre = centre
+ self.axis_ratio = axis_ratio
+ self.angle = angle
+
+ def transformed_to_reference_frame_grid_from(self, grid: np.ndarray):
+ """
+ Transform a grid of (y,x) coordinates to the geometric reference frame of the profile via a translation using
+ its `centre` and a rotation using its `angle`.
+
+ This performs the following steps:
+
+ 1) Translate the input (y,x) coordinates from the (0.0, 0.0) origin to the centre of the profile by subtracting
+ the profile centre.
+
+ 2) Compute the radial distance of every translated coordinate from the centre.
+
+ 3) Rotate the coordinates from the above step counter-clockwise from the positive x-axis by the
+ profile's `angle`.
+
+ Parameters
+ ----------
+ grid
+ The (y, x) coordinate grid in its original reference frame.
+ """
+
+ shifted_grid = grid - self.centre
+ effective_radius = (
+ (shifted_grid[:, :, 0] ** 2.0 + shifted_grid[:, :, 1] ** 2.0)
+ ) ** 0.5
+
+ theta_coordinate_to_profile = np.arctan2(
+ shifted_grid[:, :, 1], shifted_grid[:, :, 0]
+ ) - np.radians(self.angle)
+
+ transformed_grid = np.zeros(grid.shape)
+
+ transformed_grid[:, :, 0] = effective_radius * np.cos(
+ theta_coordinate_to_profile
+ )
+ transformed_grid[:, :, 1] = effective_radius * np.sin(
+ theta_coordinate_to_profile
+ )
+
+ return transformed_grid
+
+ def rotated_grid_from_reference_frame_from(self, grid: np.ndarray) -> np.ndarray:
+ """
+ Rotate a grid of (y,x) coordinates which have been transformed to the elliptical reference frame of a profile
+ back to the original unrotated coordinate frame.
+
+ This performs the following steps:
+
+ 1) Rotate the coordinates from the elliptical reference frame back to the original unrotated coordinate frame
+ using the profile's `angle`.
+
+ 2) Translate the coordinates from the centre of the profile back to the (0.0, 0.0) origin by adding the
+ profile centre.
+
+ Parameters
+ ----------
+ grid
+ The (y, x) coordinates in the reference frame of an elliptical profile.
+ """
+ cos_angle = np.cos(np.radians(self.angle))
+ sin_angle = np.sin(np.radians(self.angle))
+
+ transformed_grid = np.zeros(grid.shape)
+
+ transformed_grid[:, :, 0] = np.add(
+ np.multiply(grid[:, :, 0], cos_angle),
+ -np.multiply(grid[:, :, 1], sin_angle),
+ )
+ transformed_grid[:, :, 1] = np.add(
+ np.multiply(grid[:, :, 0], sin_angle), np.multiply(grid[:, :, 1], cos_angle)
+ )
+
+ return transformed_grid
+
+ def elliptical_radii_grid_from(self, grid: np.ndarray) -> np.ndarray:
+ """
+ Convert a grid of (y,x) coordinates to a grid of elliptical radii.
+
+ Parameters
+ ----------
+ grid
+ The (y, x) coordinates in the reference frame of the elliptical profile.
+ """
+
+ return (
+ (grid[:, :, 1] ** 2.0) + (grid[:, :, 0] / self.axis_ratio) ** 2.0
+ ) ** 0.5
+
+
+class LightProfile(GeometryProfile):
+ def __init__(
+ self,
+ centre: Tuple[float, float] = (0.0, 0.0),
+ axis_ratio: float = 1.0,
+ angle: float = 0.0,
+ intensity: float = 0.1,
+ effective_radius: float = 0.6,
+ ):
+ """
+ Abstract base class for a light profile, which describes the emission of a galaxy as a
+ function of radius.
+
+ Parameters
+ ----------
+ centre
+ The (y,x) coordinates of the profile centre.
+ axis_ratio
+ The axis-ratio of the ellipse (minor axis / major axis).
+ angle
+ The rotation angle in degrees counter-clockwise from the positive x-axis.
+ intensity
+ Overall intensity normalisation of the light profile.
+ effective_radius
+ The circular radius containing half the light of this profile.
+ """
+
+ super().__init__(centre=centre, axis_ratio=axis_ratio, angle=angle)
+
+ self.intensity = intensity
+ self.effective_radius = effective_radius
+
+
+class LightBulgey(LightProfile):
+ def __init__(
+ self,
+ centre: Tuple[float, float] = (0.0, 0.0),
+ axis_ratio: float = 1.0,
+ angle: float = 0.0,
+ intensity: float = 0.1,
+ effective_radius: float = 0.6,
+ ):
+ """
+ The De Vaucouleurs light profile often used in Astronomy to represent the bulge of galaxies.
+
+ Parameters
+ ----------
+ centre
+ The (y,x) coordinates of the profile centre.
+ axis_ratio
+ The axis-ratio of the ellipse (minor axis / major axis).
+ angle
+ The rotation angle in degrees counter-clockwise from the positive x-axis.
+ intensity
+ Overall intensity normalisation of the light profile.
+ effective_radius
+ The circular radius containing half the light of this profile.
+ """
+
+ super().__init__(
+ centre=centre,
+ axis_ratio=axis_ratio,
+ angle=angle,
+ intensity=intensity,
+ effective_radius=effective_radius,
+ )
+
+ def image_from_grid(self, grid: np.ndarray) -> np.ndarray:
+ """
+ Returns the image of the De Vaucouleurs light profile on a grid of Cartesian (y,x) coordinates, which are
+ first translated to the profile's reference frame.
+
+ Parameters
+ ----------
+ grid
+ The (y, x) coordinates where the image is computed.
+ """
+ grid_transformed = self.transformed_to_reference_frame_grid_from(grid=grid)
+ grid_elliptical_radii = self.elliptical_radii_grid_from(grid=grid_transformed)
+
+ return self.intensity * np.exp(
+ -7.66924
+ * ((grid_elliptical_radii / self.effective_radius) ** (1.0 / 7.66924) - 1.0)
+ )
+
+
+class LightDisky(LightProfile):
+ def __init__(
+ self,
+ centre: Tuple[float, float] = (0.0, 0.0),
+ axis_ratio: float = 1.0,
+ angle: float = 0.0,
+ intensity: float = 0.1,
+ effective_radius: float = 0.6,
+ ):
+ """
+ The Exponential light profile often used in Astronomy to represent the disk of galaxies.
+
+ Parameters
+ ----------
+ centre
+ The (y,x) coordinates of the profile centre.
+ axis_ratio
+ The axis-ratio of the ellipse (minor axis / major axis).
+ angle
+ The rotation angle in degrees counter-clockwise from the positive x-axis.
+ intensity
+ Overall intensity normalisation of the light profile.
+ effective_radius
+ The circular radius containing half the light of this profile.
+ """
+
+ super().__init__(
+ centre=centre,
+ axis_ratio=axis_ratio,
+ angle=angle,
+ intensity=intensity,
+ effective_radius=effective_radius,
+ )
+
+ def image_from_grid(self, grid: np.ndarray) -> np.ndarray:
+ """
+ Returns the image of the light profile on a grid of Cartesian (y,x) coordinates.
+
+ Parameters
+ ----------
+ grid
+ The (y, x) coordinates where the image is computed.
+ """
+ grid_transformed = self.transformed_to_reference_frame_grid_from(grid=grid)
+ grid_elliptical_radii = self.elliptical_radii_grid_from(grid=grid_transformed)
+
+ return self.intensity * np.exp(
+ -1.67838
+ * ((grid_elliptical_radii / self.effective_radius) ** (1.0 / 1.67838) - 1.0)
+ )
+
+
+"""
+Here is an example of an image of a `LightDisky` profile, using the `image_from_grid` method and the
+grid aligned with the image of the galaxy.
+"""
+light_profile = LightDisky(
+ centre=(0.01, 0.01), axis_ratio=0.7, angle=45.0, intensity=1.0, effective_radius=2.0
+)
+light_image = light_profile.image_from_grid(grid=grid)
+
+plot_array(array=light_image, title="Image of an Exponential light profile.")
+
+"""
+__Model Data__
+
+To produce the `model_data`, we now convolve the overall image with the Point Spread Function (PSF) of our observations.
+This blurs the image to simulate the effects of the telescope optics and pixelization used to capture the image.
+"""
+model_data = signal.convolve2d(light_image, psf, mode="same")
+
+plot_array(array=model_data, title="Model Data of the Light Profile.")
+
+"""
+By subtracting the model image from the data, we create a 2D residual map, similar to the residual maps we made in
+the 1D Gaussian examples but now for 2D imaging data. It's evident that the random model we used here does not fit
+the galaxy well.
+"""
+residual_map = data - model_data
+
+plot_array(array=residual_map, title="Residual Map of fit")
+
+"""
+Just like in the 1D `Gaussian` fitting examples, we can use the noise map to compute the normalized residuals and
+chi-squared map for the model.
+"""
+normalized_residual_map = residual_map / noise_map
+chi_squared_map = (normalized_residual_map) ** 2.0
+
+plot_array(
+ array=normalized_residual_map,
+ title="Normalized Residual Map of fit",
+)
+plot_array(array=chi_squared_map, title="Chi Squared Map of fit")
+
+"""
+Finally, we compute the `log_likelihood` of this model, which will be used next to fit the model to the data using
+a non-linear search.
+"""
+chi_squared = np.sum(chi_squared_map)
+noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
+
+log_likelihood = -0.5 * (chi_squared + noise_normalization)
+
+print(log_likelihood)
+
+"""
+__Model__
+
+We now define a model where one of the light profiles defined earlier is encapsulated within a `Model` object.
+This allows us to treat the light profile as a model component that can be used in conjunction with other components.
+
+Although we are currently using only one light profile in this example, we structure it within a `Collection` for
+potential extension with multiple light profiles.
+"""
+model = af.Collection(light_0=af.Model(LightDisky))
+
+"""
+The `model` operates the same as the model components we've utilized previously.
+
+Fitting 2D data is more time-consuming than 1D Gaussian data. I therefore employ the prior tuning method described in
+tutorial 4 to speed up the fit, by adjusting the priors to approximate values close to the correct solution.
+This adjustment is based on thorough fits that were performed using a slow and detailed search in order to locate
+the global maxima of the likelihood.
+
+Additionally, I fix the center of the light profile to (0.0, 0.0), where it visually appears to be located.
+This reduces the number of free parameters and simplifies the complexity of the non-linear search.
+"""
+model.light_0.centre_0 = 0.0
+model.light_0.centre_1 = 0.0
+model.light_0.angle = af.UniformPrior(lower_limit=100.0, upper_limit=120.0)
+model.light_0.axis_ratio = af.UniformPrior(lower_limit=0.6, upper_limit=0.8)
+model.light_0.effective_radius = af.UniformPrior(lower_limit=0.0, upper_limit=1.0)
+
+"""
+The model info contains information on all of the model components and priors, including the updates above.
+"""
+print(model.info)
+
+"""
+__Analysis__
+
+We now define the `Analysis` class for this astronomy example, which will fit the `model` to the image data.
+
+Checkout all the docstrings in this class for details on how the fit is performed to 2D imaging data, including
+the role of the mask and PSF.
+"""
+
+
+class Analysis(af.Analysis):
+ def __init__(
+ self,
+ data: np.ndarray,
+ noise_map: np.ndarray,
+ psf: np.ndarray,
+ grid: np.ndarray,
+ mask: np.ndarray,
+ ):
+ """
+ The analysis class for the **PyAutoFit** example Astronomy project on galaxy fitting.
+
+ This class contains imaging data of a galaxy and it fits it with a model which represents the light profile of
+ the galaxy.
+
+ The imaging data contains:
+
+ 1) An image of the galaxy.
+
+ 2) The noise in every pixel of that image.
+
+ 3) The Point Spread Function (PSF) describing how the optics of the telescope blur the image.
+
+ 4) A (y,x) grid of coordinates describing the locations of the image pixels in a unit system, which are used
+ to perform calculations.
+
+ 5) A mask that removes certain image pixels from the analysis.
+
+ This project is a scaled down version of the Astronomy project **PyAutoGalaxy**, which is one of the
+ original projects from which PyAutoFit is an offshoot!
+
+ https://github.com/Jammy2211/PyAutoGalaxy
+
+ Parameters
+ ----------
+ data
+ The image containing the observation of the galaxy that is fitted.
+ noise_map
+ The RMS noise values of the image data, which is folded into the log likelihood calculation.
+ psf
+ The Point Spread Function of the telescope, which describes how the telescope blurs the image.
+ grid
+ The (y, x) coordinates of the image from which the calculation is performed and model image is
+ computed using.
+ mask
+ The 2D mask that is applied to the image data.
+ """
+
+ super().__init__()
+
+ self.data = data
+ self.noise_map = noise_map
+ self.psf = psf
+ self.grid = grid
+ self.mask = mask
+
+ def log_likelihood_function(self, instance) -> float:
+ """
+ The `log_likelihood_function` of the galaxy example, which performs the following step:
+
+ 1) Using the model passed into the function (whose parameters are set via the non-linear search) create
+ a model image of this galaxy using the light profiles in the `instance`.
+
+ 2) Convolve this model image with the Point Spread Function of the telescope, ensuring the telescope optics
+ are included in the model image.
+
+ 3) Subtract this model image from the data and compute its residuals, chi-squared and likelihood via the
+ noise map.
+
+ 4) Apply the mask to all these quantities, ensuring the edges of the galaxy are omitted from the fit.
+
+ Parameters
+ ----------
+ instance
+ An instance of the model set via the non-linear search.
+
+ Returns
+ -------
+ float
+ The log likelihood value of this particular model.
+ """
+
+ """
+ The 'instance' that comes into this method contains the light profiles we setup in the `Model` and `Collection`,
+ which can be seen by uncommenting the code below.
+ """
+
+ # print("Model Instance:")
+ # print("Light Profile = ", instance.light)
+ # print("Light Profile Centre= ", instance.light.centre)
+
+ """
+ Generate the model data from the instance using the functions of the light profile class above.
+
+ See the docstring of the `model_data_from_instance` function for a description of how this works.
+ """
+
+ model_data = self.model_data_from_instance(instance=instance)
+
+ """
+ In this context of fitting 2D astronomical imaging data, the calculation of residual-map,
+ normalized residual-map, chi-squared-map, and likelihood differs from previous examples such as fitting 1D
+ Gaussians.
+
+ The main difference lies in the incorporation of a mask, which excludes regions where the galaxy is not
+ visible, typically the edges.
+
+ To ensure that the mask influences these calculations correctly, we use the `where` parameter of numpy.
+ This parameter allows us to compute the chi-squared and likelihood only where the mask indicates valid
+ data (where the mask is 0, indicating unmasked regions). This approach ensures that the analysis focuses
+ exclusively on the relevant parts of the galaxy image where meaningful comparisons between the model and
+ data can be made.
+ """
+ residual_map = np.subtract(
+ self.data,
+ model_data,
+ out=np.zeros_like(data),
+ where=np.asarray(self.mask) == 0,
+ )
+
+ normalized_residual_map = np.divide(
+ residual_map,
+ self.noise_map,
+ out=np.zeros_like(residual_map),
+ where=np.asarray(self.mask) == 0,
+ )
+
+ chi_squared_map = np.square(
+ np.divide(
+ residual_map,
+ self.noise_map,
+ out=np.zeros_like(residual_map),
+ where=np.asarray(mask) == 0,
+ )
+ )
+
+ chi_squared = float(np.sum(chi_squared_map[np.asarray(self.mask) == 0]))
+ noise_normalization = float(
+ np.sum(np.log(2 * np.pi * noise_map[np.asarray(mask) == 0] ** 2.0))
+ )
+
+ log_likelihood = -0.5 * (chi_squared + noise_normalization)
+
+ return log_likelihood
+
+ def model_data_from_instance(self, instance):
+ """
+ Create the image of a galaxy, including blurring due to the Point Spread Function of the telescope.
+
+ For the purpose of illustrating **PyAutoFit** you do not need to understand what this function is doing, the
+ main thing to note is that it allows us to create a model image of a galaxy.
+
+ Nevertheless, if you are curious, it inputs the (y,x) Cartesian grids into the light profiles and evaluates
+ the image of the galaxy. Multiple light profiles may be summed to create the overall model image that is fitted.
+
+ The mask is used to zero all values of the model image that are not included in the mask.
+
+ Parameters
+ ----------
+ instance
+ An instance of the model set via the non-linear search.
+
+ Returns
+ -------
+ image
+ The image of this galaxy light profile model.
+ """
+
+ overall_image = np.zeros(self.data.shape)
+
+ for light_profile in instance:
+ overall_image += light_profile.image_from_grid(grid=self.grid)
+
+ model_data = signal.convolve2d(overall_image, self.psf, mode="same")
+
+ model_data[self.mask == 1] = 0.0
+
+ return model_data
+
+
+"""
+__Model Fit__
+
+We have successfully composed a model and analysis class for the astronomy example.
+
+We can now fit it to the data using a search. We use the same method as previous examples, nested sampling
+algorithm Dynesty.
+"""
+search = af.DynestyStatic(
+ nlive=500,
+ sample="rwalk", # This makes dynesty run faster, don't worry about what it means for now!
+)
+
+analysis = Analysis(data=data, noise_map=noise_map, psf=psf, grid=grid, mask=mask)
+
+print(
+ """
+ The non-linear search has begun running.
+ This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
+ """
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+print("The search has finished run - you may now continue the notebook.")
+
+
+"""
+__Result__
+
+The `result` object provides a concise summary of the outcomes from the non-linear search. While many of these values
+may not be directly interpretable for non-astronomers, some are meaningful even without specialized knowledge.
+
+For instance, the `angle` parameter indicates the rotation of the galaxy's light profile counterclockwise from the
+x-axis. In our fit, this value should approximate 110.0 degrees, aligning with the observed orientation of the
+galaxy in the image.
+
+Similarly, the `axis-ratio` represents the ratio of the minor axis to the major axis of the galaxy's light profile.
+A value near 0.7 is expected, reflecting the elliptical shape typical of galaxies.
+
+This illustrates why we perform model-fitting, we can take complex data and infer simple, interpretable properties
+from it which provide insight into the physical processes generating the data. This is the core goal of the scientific
+method and the use of models to explain observations.
+"""
+print(result.info)
+
+"""
+The best way to assess the quality of the fit is to use the visualization techniques introduced in previous tutorials.
+
+For 2D data, we visualize the model image, residual-map, normalized residual-map, and chi-squared-map as 2D arrays.
+These quantities still have the same meanings as their 1D counterparts. For example, normalized residual-map values
+still represent the sigma values of the residuals, with values greater than 3.0 indicating a poor fit to the data.
+"""
+model_data = analysis.model_data_from_instance(
+ instance=result.max_log_likelihood_instance
+)
+
+residual_map = np.subtract(
+ data, model_data, out=np.zeros_like(data), where=np.asarray(mask) == 0
+)
+
+normalized_residual_map = np.divide(
+ residual_map,
+ noise_map,
+ out=np.zeros_like(residual_map),
+ where=np.asarray(mask) == 0,
+)
+
+chi_squared_map = np.square(
+ np.divide(
+ residual_map,
+ noise_map,
+ out=np.zeros_like(residual_map),
+ where=np.asarray(mask) == 0,
+ )
+)
+
+plot_array(
+ array=model_data,
+ title="Model Data of the Light Profile.",
+)
+plot_array(
+ array=residual_map,
+ title="Residual Map of fit",
+)
+plot_array(
+ array=normalized_residual_map,
+ title="Normalized Residual Map of fit",
+)
+plot_array(
+ array=chi_squared_map,
+ title="Chi Squared Map of fit",
+)
+
+"""
+__Bulgey__
+
+The fit above utilized the disky light profile, which is typically suitable for disk-like late-type galaxies.
+
+Now, we will repeat the fit using the bulgey light profile, which is more suitable for bulge-like early-type galaxies.
+
+The fit with the higher log likelihood will provide insight into whether the galaxy is more likely to be an early-type
+or a late-type galaxy.
+"""
+result_disk = result
+
+model = af.Collection(light_0=af.Model(LightBulgey))
+
+# model.light_0.centre_0 = 0.0
+# model.light_0.centre_1 = 0.0
+# model.light_0.axis_ratio = af.UniformPrior(lower_limit=0.7, upper_limit=1.0)
+
+print(
+ """
+ The non-linear search has begun running.
+ This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
+ """
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+print("The search has finished run - you may now continue the notebook.")
+
+"""
+Print the result info of the bulgey fit.
+"""
+print(result.info)
+
+"""
+We perform the same visualization of the model image, residual-map, normalized residual-map, and chi-squared-map as
+before.
+
+The model image of the bulgey profile provides a better fit to the data than the disky profile, as the residuals are
+lower and the chi-squared-map values are closer to 0.0.
+
+This suggests that the galaxy is more likely to be an early-type galaxy with a bulge-like light profile.
+"""
+
+model_data = analysis.model_data_from_instance(
+ instance=result.max_log_likelihood_instance
+)
+
+residual_map = np.subtract(
+ data, model_data, out=np.zeros_like(data), where=np.asarray(mask) == 0
+)
+
+normalized_residual_map = np.divide(
+ residual_map,
+ noise_map,
+ out=np.zeros_like(residual_map),
+ where=np.asarray(mask) == 0,
+)
+
+chi_squared_map = np.square(
+ np.divide(
+ residual_map,
+ noise_map,
+ out=np.zeros_like(residual_map),
+ where=np.asarray(mask) == 0,
+ )
+)
+
+plot_array(
+ array=model_data,
+ title="Model Data of the Light Profile.",
+)
+plot_array(
+ array=residual_map,
+ title="Residual Map of fit",
+)
+plot_array(
+ array=normalized_residual_map,
+ title="Normalized Residual Map of fit",
+)
+plot_array(
+ array=chi_squared_map,
+ title="Chi Squared Map of fit",
+)
+
+"""
+To make certain of our interpretation, we should compare the log likelihoods of the two fits.
+
+The fit with the highest log likelihood is the preferred model, which (provided your non-linear search sampled
+parameter space accurately), is the bulgey profile.
+
+Therefore, the galaxy is likely an early-type galaxy with a bulge-like light profile.
+"""
+print("Disk Model Log Likelihood:")
+print(result_disk.log_likelihood)
+print("Bulge Model Log Likelihood:")
+print(result.log_likelihood)
+
+"""
+__Model Mismatch__
+
+The analysis above allowed us to determine whether the galaxy is more likely to be an early-type or late-type galaxy.
+
+However, after fitting the bulgey profile, you may not of expected it to be the highest log likelihood model. The
+model gave a relatively poor fit, with significant residuals and chi-squared values. It just turns out that
+the disky profile gave an even worse fit!
+
+This reflected the notion of "model mismatch" that we discussed in tutorial 4. One of the challenges of model fitting
+is that you may not have a model that is a brilliant representation of the data, and your search is successfully
+locating the global maxima even though the fit looks visibly poor.
+
+In Astronomy, what a scientist would do next is update their model to try and improve the fit. For example, they
+may extend the model to contain both the bulgey and disky profile, allowing the model to fit both components of the
+galaxy simultaneously.
+
+There are a whole range of approaches that Astronomers take to improve the model, which include fitting the
+galaxy with hundreds of 2D Gaussians, fitting it with a pixel grid of light and decomposing it into what
+are called basis functions. These approaches are not relevent to your understanding of **PyAutoFit**, but they
+are all implemented in **PyAutoGalaxy** via the **PyAutoFit** API for model composition.
+
+This is another great example of how **PyAutoFit** can be used to perform complex model-fitting tasks.
+
+__Extensions__
+
+To conclude, I will illustrate some of the extensions that can be made to this example and the challenges that arise
+when fitting more complex models.
+
+I don't provide code examples of how to tackle these extensions with model-fitting, but I encourage you to think about
+how **PyAutoFit** could be used to address these problems and go ahead and give them a go yourself if you're feeling
+adventurous!
+
+**Multiple Components**:
+
+The model above fitted a single light profile to the galaxy, which turned out to be a bulge-like component.
+
+However, galaxies often have multiple components, for example a disk and bulge, but also structures such as a bar or
+spiral arms. Here is a galaxy with a bulge, disk, bar and spiral arms:
+
+
+
+To model this galaxy, we would add many different Python classes with their own parameters and light profiles
+representing each component of the galaxy. They would then be combined into a single model, whose model image
+would be the summed image of each component, the total number of parameters would be in the twenties or thirties
+and the degeneracies between the parameters would be challenging to sample accurately.
+
+Here is a snippet of rougyly what our model composition would look like:
+
+model = af.Collection(
+ bulge=LightBulge,
+ disk=LightDisk,
+ bar=LightBar,
+ spiral_arms=LightSpiralArms
+)
+
+This is a great example of how quickly model-fitting can become complex and how keeping track of the speed and
+efficiency of the non-linear search becomes crucial. Furthermore, with so many different light profiles to fit,
+**PyAutoFit**'s model composition API becomes invaluable.
+
+
+**Multiple Galaxies**:
+
+There is no reason our imaging data need contain only one galaxy. The data could include multiple galaxies, each
+of which we want to fit with its own light profile.
+
+Two galaxies close together are called galaxy mergers, and here is a beautiful example of a pair of merging galaxies:
+
+
+
+This would pretty much double the model complexity, as each galaxy would have its own model and light profile.
+All the usual issues then become doubly important, such as ensuring the likelihood function is efficient, that
+the search can sample parameter space accurately and that the results are interpreted correctly.
+
+The model composition would again change, and might look something like:
+
+model = af.Collection(
+ bulge_0=LightBulge,
+ disk_0=LightDisk,
+ bulge_1=LightBulge,
+ disk_1=LightDisk
+)
+
+model.bulge_0.centre_0 = 2.0
+model.bulge_0.centre_1 = 0.0
+model.disk_0.centre_0 = 2.0
+model.disk_0.centre_1 = 0.0
+
+model.bulge_1.centre_0 = -2.0
+model.bulge_1.centre_1 = 0.0
+model.disk_1.centre_0 = -2.0
+model.disk_1.centre_1 = 0.0
+
+In order to keep the model slightly more simple, the centres of the light profiles have been fixed to where
+they peak in the image.
+
+**PyAutoFit** extensible model composition API actually has much better tools for composing complex models like this
+than the example above. You can find a concise run through of these in the model cookbook, but they will
+also be the focus on a tutorial in the next chapter of the **HowToFit** lectures.
+
+
+**Multiple Wavelengths**:
+
+Galaxies emit light in many wavelengths, for example ultraviolet, optical, infrared, radio and X-ray. Each wavelength
+provides different information about the galaxy, for example the ultraviolet light tells us about star formation,
+the optical light about the stars themselves and the infrared about dust.
+
+The image below shows observations of the same galaxy at different wavelengths:
+
+
+
+In tutorial 6, we learn how to perform fits to multiple datasets simultaneously, and how to change the model
+parameterization to account for the variation across datasets.
+
+Multi-wavelength modeling of galaxies is a great example of where this is useful, as it allows us to fit the galaxy
+with certain parameters shared across all wavelengths (e.g., its centre) and other parameters varied (e.g., its
+intensity and effective radius). A child project of **PyAutoFit**, called **PyAutoGalaxy**, uses this functionality
+to achieve exactly this.
+
+__Chapter Wrap Up__
+
+We have now completed the first chapter of **HowToFit**, which has taught you the basics of model-fitting.
+
+Its now time you take everything you've learnt and apply it to your own model-fitting problem. Think carefully
+about the key concepts of this chapter, for example how to compose a model, how to create an analysis class and
+how to overcome challenges that arise when fitting complex models.
+
+Once you are familiar with these concepts, and confident that you have some simple model-fitting problems under
+your belt, you should move on to the next chapter. This covers how to build a scientific workflow around your
+model-fitting, so that you can begin to scale up your model-fitting to more complex models, larger datasets and
+more difficult problems. Checkout the `start_here.ipynb` notebook in chapter 2 to get started!
+"""
diff --git a/scripts/howtofit/chapter_3_graphical_models/tutorial_1_individual_models.py b/scripts/howtofit/chapter_3_graphical_models/tutorial_1_individual_models.py
index ee523d71..3f599c2c 100644
--- a/scripts/howtofit/chapter_3_graphical_models/tutorial_1_individual_models.py
+++ b/scripts/howtofit/chapter_3_graphical_models/tutorial_1_individual_models.py
@@ -1,323 +1,319 @@
-"""
-Tutorial 1: Individual Models
-=============================
-
-In many examples, we fit one model to one dataset. For many problems, we may have a large dataset and are not
-interested in how well the model fits each individual dataset. Instead, we want to know how the model fits the full
-dataset, so that we can determine "global" trends of how the model fits the data.
-
-These tutorials show you how to compose and fit hierarchical models to large datasets, which fit many individual
-models to each dataset. However, all parameters in the model are linked together, enabling global inference of the
-model over the full dataset. This can extract a significant amount of extra information from large datasets, which
-fitting each dataset individually does not.
-
-Fitting a hierarchical model uses a "graphical model", which is a model that is simultaneously fitted to every
-dataset simultaneously. The graph expresses how the parameters of every individual model is paired with each dataset
-and how they are linked to every other model parameter. Complex graphical models fitting a diversity of different
-datasets and non-trivial model parameter linking is possible and common.
-
-This chapter will start by fitting a simple graphical model to a dataset of noisy 1D Gaussians. The Gaussians all
-share the same `centre`, meaning that a graphical model can be composed where there is only a single global `centre`
-shared by all Gaussians.
-
-However, before fitting a graphical model, we will first fit each Gaussian individually and combine the inference
-on the `centre` after every fit is complete. This will give us an estimate of the `centre` that we can compare to
-the result of the graphical model in tutorial 2.
-
-__Real World Example__
-
-Hierarchical models are often used to determine effective drug treatments across a sample of patients distributed over
-many hospitals. Trying to do this on each individual hospital dataset is not ideal, as the number of patients in each
-hospital is small and the treatment may be more or less effective in some hospitals than others. Hierarchical models
-can extract the global trends of how effective the treatment is across the full population of patients.
-
-In healthcare, there may also be many datasets available, with different formats that require slightly different models
-to fit them. The high levels of customization possible in model composition and defining the analysis class mean
-that fitting diverse datasets with hierarchical models is feasible. This also means that a common problem in healthcare
-data, missing data, can be treated in a statistically robust manner.
-
-__Contents__
-
-This tutorial is split into the following sections:
-
-- **Real World Example**: A healthcare example illustrating the value of hierarchical models.
-- **Example Source Code (`af.ex`)**: The example objects used in this tutorial.
-- **Model**: Define a simple Gaussian model in a Collection.
-- **Data**: Load and set up 5 noisy 1D Gaussian datasets.
-- **Model Fits (one-by-one)**: Fit each dataset individually using a separate non-linear search.
-- **Results**: Analyze the fit results and error estimates for each dataset.
-- **Estimating the Centre**: Combine centre estimates using a weighted average approach.
-- **Posterior Multiplication**: Discuss KDE-based posterior multiplication as an alternative method.
-- **Wrap Up**: Summary and transition to the graphical model approach in the next tutorial.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-from os import path
-import numpy as np
-
-import autofit as af
-import autofit.plot as aplt
-
-"""
-__Example Source Code (`af.ex`)__
-
-The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
-
- - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
- `visualize` functions.
-
- - `Gaussian`: a model component representing a 1D Gaussian profile.
-
- - `plot_profile_1d`: a function for plotting 1D profile datasets including their noise.
-
-These are functionally identical to the `Analysis`, `Gaussian` and `plot_profile_1d` objects and functions you have
-seen and used elsewhere throughout the workspace.
-
-__Model__
-
-Our model is a single `Gaussian`.
-
-We put this in a `Collection` so that when we extend the model in later tutorials we use the same API throughout
-all tutorials.
-"""
-model = af.Collection(gaussian=af.ex.Gaussian)
-
-"""
-__Data__
-
-We quickly set up the name of each dataset, which is used below for loading the datasets.
-
-The dataset contains 10 Gaussians, but for speed we'll fit just 5. You can change this to 10 to see how the result
-changes with more datasets.
-"""
-total_datasets = 5
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(
- path.join("dataset", "example_1d", "gaussian_x1__low_snr", "dataset_0")
-):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-dataset_name_list = []
-
-for dataset_index in range(total_datasets):
- dataset_name_list.append(f"dataset_{dataset_index}")
-
-"""
-For each 1D Gaussian dataset we now set up the correct path, load it, and plot it.
-
-Notice how much lower the signal-to-noise is than you are used too, you probably find it difficult to estimate
-the centre of some of the Gaussians by eye!
-"""
-for dataset_name in dataset_name_list:
- """
- Load the dataset from the `autofit_workspace/dataset` folder.
- """
-
- dataset_path = path.join(
- "dataset", "example_1d", "gaussian_x1__low_snr", dataset_name
- )
-
- data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
- noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
- )
-
- af.ex.plot_profile_1d(
- xvalues=np.arange(data.shape[0]),
- profile_1d=data,
- title=dataset_name,
- ylabel="Data Values",
- color="k",
- )
-
-"""
-__Model Fits (one-by-one)__
-
-For every dataset we now create an `Analysis` and fit it with a `Gaussian`.
-
-The `Result` is stored in the list `result_list`.
-"""
-result_list = []
-
-for dataset_name in dataset_name_list:
- """
- Load the dataset from the `autofit_workspace/dataset` folder.
- """
- dataset_path = path.join(
- "dataset", "example_1d", "gaussian_x1__low_snr", dataset_name
- )
-
- data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
- noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
- )
-
- """
- For each dataset create a corresponding `Analysis` class.
- """
- analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
- """
- Create the `DynestyStatic` non-linear search and use it to fit the data.
-
- We use custom dynesty settings which ensure the posterior is explored fully and that our error estimates are robust.
- """
- dynesty = af.DynestyStatic(
- name="global_model",
- path_prefix=path.join(
- "howtofit", "chapter_graphical_models", "tutorial_1_individual_models"
- ),
- unique_tag=dataset_name,
- nlive=200,
- dlogz=1e-4,
- sample="rwalk",
- walks=10,
- )
-
- print(
- f"The non-linear search has begun running, checkout \n"
- f"autofit_workspace/output/howtofit/chapter_graphica_models/tutorial_1_individual_models/{dataset_name} for live \n"
- f"output of the results. This Jupyter notebook cell with progress once search has completed, this could take a \n"
- f"few minutes!"
- )
-
- result_list.append(dynesty.fit(model=model, analysis=analysis))
-
-
-"""
-__Results__
-
-Checkout the output folder, you should see five new sets of results corresponding to our Gaussian datasets.
-
-In the `model.results` file of each fit, it will be clear that the `centre` value of every fit (and the other
-parameters) have much larger errors than other **PyAutoFit** examples due to the low signal to noise of the data.
-
-The `result_list` allows us to plot the median PDF value and 3.0 confidence intervals of the `centre` estimate from
-the model-fit to each dataset.
-"""
-import matplotlib.pyplot as plt
-
-samples_list = [result.samples for result in result_list]
-
-mp_instances = [samps.median_pdf() for samps in samples_list]
-ue3_instances = [samp.errors_at_upper_sigma(sigma=3.0) for samp in samples_list]
-le3_instances = [samp.errors_at_lower_sigma(sigma=3.0) for samp in samples_list]
-
-mp_centres = [instance.gaussian.centre for instance in mp_instances]
-ue3_centres = [instance.gaussian.centre for instance in ue3_instances]
-le3_centres = [instance.gaussian.centre for instance in le3_instances]
-
-plt.errorbar(
- x=[f"Gaussian {index}" for index in range(total_datasets)],
- y=mp_centres,
- marker=".",
- linestyle="",
- yerr=[le3_centres, ue3_centres],
-)
-plt.xticks(rotation=90)
-plt.show()
-plt.close()
-
-"""
-These model-fits are consistent with a range of `centre` values.
-
-We can show this by plotting the 1D and 2D PDF's of each model fit
-"""
-
-for samples in samples_list:
- aplt.corner_cornerpy(samples=samples)
-
-"""
-We can also print the values of each centre estimate, including their estimates at 3.0 sigma.
-
-Note that above we used the samples to estimate the size of the errors on the parameters. Below, we use the samples to
-get the value of the parameter at these sigma confidence intervals.
-"""
-u1_instances = [samp.values_at_upper_sigma(sigma=1.0) for samp in samples_list]
-l1_instances = [samp.values_at_lower_sigma(sigma=1.0) for samp in samples_list]
-
-u1_centres = [instance.gaussian.centre for instance in u1_instances]
-l1_centres = [instance.gaussian.centre for instance in l1_instances]
-
-u3_instances = [samp.values_at_upper_sigma(sigma=3.0) for samp in samples_list]
-l3_instances = [samp.values_at_lower_sigma(sigma=3.0) for samp in samples_list]
-
-u3_centres = [instance.gaussian.centre for instance in u3_instances]
-l3_centres = [instance.gaussian.centre for instance in l3_instances]
-
-for index in range(total_datasets):
- print(f"Centre estimate of Gaussian dataset {index}:\n")
- print(
- f"{mp_centres[index]} ({l1_centres[index]} {u1_centres[index]}) [1.0 sigma confidence interval]"
- )
- print(
- f"{mp_centres[index]} ({l3_centres[index]} {u3_centres[index]}) [3.0 sigma confidence interval] \n"
- )
-
-
-"""
-__Estimating the Centre__
-
-So how might we estimate our global `centre` value?
-
-A simple approach takes the weighted average of the value inferred by all five fits above.
-"""
-ue1_instances = [samp.values_at_upper_sigma(sigma=1.0) for samp in samples_list]
-le1_instances = [samp.values_at_lower_sigma(sigma=1.0) for samp in samples_list]
-
-ue1_centres = [instance.gaussian.centre for instance in ue1_instances]
-le1_centres = [instance.gaussian.centre for instance in le1_instances]
-
-error_list = [ue1 - le1 for ue1, le1 in zip(ue1_centres, le1_centres)]
-
-values = np.asarray(mp_centres)
-sigmas = np.asarray(error_list)
-
-weights = 1 / sigmas**2.0
-weight_averaged = np.sum(1.0 / sigmas**2)
-
-weighted_centre = np.sum(values * weights) / np.sum(weights, axis=0)
-weighted_error = 1.0 / np.sqrt(weight_averaged)
-
-print(
- f"Weighted Average Centre Estimate = {weighted_centre} ({weighted_error}) [1.0 sigma confidence intervals]"
-)
-
-"""
-__Posterior Multiplication__
-
-An alternative and more accurate way to combine each individual inferred centre is multiply their posteriors together.
-
-In order to do this, a smooth 1D profile must be fit to the posteriors via a Kernel Density Estimator (KDE).
-
-[There is currently no support for posterior multiplication and an example illustrating this is currently missing
-from this tutorial. However, I will discuss KDE multiplication throughout these tutorials to give the reader context
-for how this approach to parameter estimation compares to graphical models.]
-
-__Wrap Up__
-
-Lets wrap up the tutorial. The methods used above combine the results of different fits and estimate a global
-value of `centre` alongside estimates of its error.
-
-In this tutorial, we fitted just 5 datasets. Of course, we could easily fit more datasets, and we would find that
-as we added more datasets our estimate of the global centre would become more precise.
-
-In the next tutorial, we will compare this result to one inferred via a graphical model.
-"""
+"""
+Tutorial 1: Individual Models
+=============================
+
+In many examples, we fit one model to one dataset. For many problems, we may have a large dataset and are not
+interested in how well the model fits each individual dataset. Instead, we want to know how the model fits the full
+dataset, so that we can determine "global" trends of how the model fits the data.
+
+These tutorials show you how to compose and fit hierarchical models to large datasets, which fit many individual
+models to each dataset. However, all parameters in the model are linked together, enabling global inference of the
+model over the full dataset. This can extract a significant amount of extra information from large datasets, which
+fitting each dataset individually does not.
+
+Fitting a hierarchical model uses a "graphical model", which is a model that is simultaneously fitted to every
+dataset simultaneously. The graph expresses how the parameters of every individual model is paired with each dataset
+and how they are linked to every other model parameter. Complex graphical models fitting a diversity of different
+datasets and non-trivial model parameter linking is possible and common.
+
+This chapter will start by fitting a simple graphical model to a dataset of noisy 1D Gaussians. The Gaussians all
+share the same `centre`, meaning that a graphical model can be composed where there is only a single global `centre`
+shared by all Gaussians.
+
+However, before fitting a graphical model, we will first fit each Gaussian individually and combine the inference
+on the `centre` after every fit is complete. This will give us an estimate of the `centre` that we can compare to
+the result of the graphical model in tutorial 2.
+
+__Real World Example__
+
+Hierarchical models are often used to determine effective drug treatments across a sample of patients distributed over
+many hospitals. Trying to do this on each individual hospital dataset is not ideal, as the number of patients in each
+hospital is small and the treatment may be more or less effective in some hospitals than others. Hierarchical models
+can extract the global trends of how effective the treatment is across the full population of patients.
+
+In healthcare, there may also be many datasets available, with different formats that require slightly different models
+to fit them. The high levels of customization possible in model composition and defining the analysis class mean
+that fitting diverse datasets with hierarchical models is feasible. This also means that a common problem in healthcare
+data, missing data, can be treated in a statistically robust manner.
+
+__Contents__
+
+This tutorial is split into the following sections:
+
+- **Real World Example**: A healthcare example illustrating the value of hierarchical models.
+- **Example Source Code (`af.ex`)**: The example objects used in this tutorial.
+- **Model**: Define a simple Gaussian model in a Collection.
+- **Data**: Load and set up 5 noisy 1D Gaussian datasets.
+- **Model Fits (one-by-one)**: Fit each dataset individually using a separate non-linear search.
+- **Results**: Analyze the fit results and error estimates for each dataset.
+- **Estimating the Centre**: Combine centre estimates using a weighted average approach.
+- **Posterior Multiplication**: Discuss KDE-based posterior multiplication as an alternative method.
+- **Wrap Up**: Summary and transition to the graphical model approach in the next tutorial.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+from os import path
+import numpy as np
+
+import autofit as af
+import autofit.plot as aplt
+
+"""
+__Example Source Code (`af.ex`)__
+
+The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
+
+ - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
+ `visualize` functions.
+
+ - `Gaussian`: a model component representing a 1D Gaussian profile.
+
+ - `plot_profile_1d`: a function for plotting 1D profile datasets including their noise.
+
+These are functionally identical to the `Analysis`, `Gaussian` and `plot_profile_1d` objects and functions you have
+seen and used elsewhere throughout the workspace.
+
+__Model__
+
+Our model is a single `Gaussian`.
+
+We put this in a `Collection` so that when we extend the model in later tutorials we use the same API throughout
+all tutorials.
+"""
+model = af.Collection(gaussian=af.ex.Gaussian)
+
+"""
+__Data__
+
+We quickly set up the name of each dataset, which is used below for loading the datasets.
+
+The dataset contains 10 Gaussians, but for speed we'll fit just 5. You can change this to 10 to see how the result
+changes with more datasets.
+"""
+total_datasets = 5
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(
+ path.join("dataset", "example_1d", "gaussian_x1__low_snr", "dataset_0")
+):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+dataset_name_list = []
+
+for dataset_index in range(total_datasets):
+ dataset_name_list.append(f"dataset_{dataset_index}")
+
+"""
+For each 1D Gaussian dataset we now set up the correct path, load it, and plot it.
+
+Notice how much lower the signal-to-noise is than you are used too, you probably find it difficult to estimate
+the centre of some of the Gaussians by eye!
+"""
+for dataset_name in dataset_name_list:
+ """
+ Load the dataset from the `autofit_workspace/dataset` folder.
+ """
+
+ dataset_path = path.join(
+ "dataset", "example_1d", "gaussian_x1__low_snr", dataset_name
+ )
+
+ data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+ noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+ )
+
+ af.ex.plot_profile_1d(
+ xvalues=np.arange(data.shape[0]),
+ profile_1d=data,
+ title=dataset_name,
+ ylabel="Data Values",
+ color="k",
+ )
+
+"""
+__Model Fits (one-by-one)__
+
+For every dataset we now create an `Analysis` and fit it with a `Gaussian`.
+
+The `Result` is stored in the list `result_list`.
+"""
+result_list = []
+
+for dataset_name in dataset_name_list:
+ """
+ Load the dataset from the `autofit_workspace/dataset` folder.
+ """
+ dataset_path = path.join(
+ "dataset", "example_1d", "gaussian_x1__low_snr", dataset_name
+ )
+
+ data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+ noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+ )
+
+ """
+ For each dataset create a corresponding `Analysis` class.
+ """
+ analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+ """
+ Create the `DynestyStatic` non-linear search and use it to fit the data.
+
+ We use custom dynesty settings which ensure the posterior is explored fully and that our error estimates are robust.
+ """
+ dynesty = af.DynestyStatic(
+ name="global_model",
+ path_prefix=path.join(
+ "howtofit", "chapter_graphical_models", "tutorial_1_individual_models"
+ ),
+ unique_tag=dataset_name,
+ nlive=200,
+ dlogz=1e-4,
+ sample="rwalk",
+ walks=10,
+ )
+
+ print(
+ f"The non-linear search has begun running, checkout \n"
+ f"autofit_workspace/output/howtofit/chapter_graphica_models/tutorial_1_individual_models/{dataset_name} for live \n"
+ f"output of the results. This Jupyter notebook cell with progress once search has completed, this could take a \n"
+ f"few minutes!"
+ )
+
+ result_list.append(dynesty.fit(model=model, analysis=analysis))
+
+
+"""
+__Results__
+
+Checkout the output folder, you should see five new sets of results corresponding to our Gaussian datasets.
+
+In the `model.results` file of each fit, it will be clear that the `centre` value of every fit (and the other
+parameters) have much larger errors than other **PyAutoFit** examples due to the low signal to noise of the data.
+
+The `result_list` allows us to plot the median PDF value and 3.0 confidence intervals of the `centre` estimate from
+the model-fit to each dataset.
+"""
+import matplotlib.pyplot as plt
+
+samples_list = [result.samples for result in result_list]
+
+mp_instances = [samps.median_pdf() for samps in samples_list]
+ue3_instances = [samp.errors_at_upper_sigma(sigma=3.0) for samp in samples_list]
+le3_instances = [samp.errors_at_lower_sigma(sigma=3.0) for samp in samples_list]
+
+mp_centres = [instance.gaussian.centre for instance in mp_instances]
+ue3_centres = [instance.gaussian.centre for instance in ue3_instances]
+le3_centres = [instance.gaussian.centre for instance in le3_instances]
+
+plt.errorbar(
+ x=[f"Gaussian {index}" for index in range(total_datasets)],
+ y=mp_centres,
+ marker=".",
+ linestyle="",
+ yerr=[le3_centres, ue3_centres],
+)
+plt.xticks(rotation=90)
+plt.show()
+plt.close()
+
+"""
+These model-fits are consistent with a range of `centre` values.
+
+We can show this by plotting the 1D and 2D PDF's of each model fit
+"""
+
+for samples in samples_list:
+ aplt.corner_cornerpy(samples=samples)
+
+"""
+We can also print the values of each centre estimate, including their estimates at 3.0 sigma.
+
+Note that above we used the samples to estimate the size of the errors on the parameters. Below, we use the samples to
+get the value of the parameter at these sigma confidence intervals.
+"""
+u1_instances = [samp.values_at_upper_sigma(sigma=1.0) for samp in samples_list]
+l1_instances = [samp.values_at_lower_sigma(sigma=1.0) for samp in samples_list]
+
+u1_centres = [instance.gaussian.centre for instance in u1_instances]
+l1_centres = [instance.gaussian.centre for instance in l1_instances]
+
+u3_instances = [samp.values_at_upper_sigma(sigma=3.0) for samp in samples_list]
+l3_instances = [samp.values_at_lower_sigma(sigma=3.0) for samp in samples_list]
+
+u3_centres = [instance.gaussian.centre for instance in u3_instances]
+l3_centres = [instance.gaussian.centre for instance in l3_instances]
+
+for index in range(total_datasets):
+ print(f"Centre estimate of Gaussian dataset {index}:\n")
+ print(
+ f"{mp_centres[index]} ({l1_centres[index]} {u1_centres[index]}) [1.0 sigma confidence interval]"
+ )
+ print(
+ f"{mp_centres[index]} ({l3_centres[index]} {u3_centres[index]}) [3.0 sigma confidence interval] \n"
+ )
+
+
+"""
+__Estimating the Centre__
+
+So how might we estimate our global `centre` value?
+
+A simple approach takes the weighted average of the value inferred by all five fits above.
+"""
+ue1_instances = [samp.values_at_upper_sigma(sigma=1.0) for samp in samples_list]
+le1_instances = [samp.values_at_lower_sigma(sigma=1.0) for samp in samples_list]
+
+ue1_centres = [instance.gaussian.centre for instance in ue1_instances]
+le1_centres = [instance.gaussian.centre for instance in le1_instances]
+
+error_list = [ue1 - le1 for ue1, le1 in zip(ue1_centres, le1_centres)]
+
+values = np.asarray(mp_centres)
+sigmas = np.asarray(error_list)
+
+weights = 1 / sigmas**2.0
+weight_averaged = np.sum(1.0 / sigmas**2)
+
+weighted_centre = np.sum(values * weights) / np.sum(weights, axis=0)
+weighted_error = 1.0 / np.sqrt(weight_averaged)
+
+print(
+ f"Weighted Average Centre Estimate = {weighted_centre} ({weighted_error}) [1.0 sigma confidence intervals]"
+)
+
+"""
+__Posterior Multiplication__
+
+An alternative and more accurate way to combine each individual inferred centre is multiply their posteriors together.
+
+In order to do this, a smooth 1D profile must be fit to the posteriors via a Kernel Density Estimator (KDE).
+
+[There is currently no support for posterior multiplication and an example illustrating this is currently missing
+from this tutorial. However, I will discuss KDE multiplication throughout these tutorials to give the reader context
+for how this approach to parameter estimation compares to graphical models.]
+
+__Wrap Up__
+
+Lets wrap up the tutorial. The methods used above combine the results of different fits and estimate a global
+value of `centre` alongside estimates of its error.
+
+In this tutorial, we fitted just 5 datasets. Of course, we could easily fit more datasets, and we would find that
+as we added more datasets our estimate of the global centre would become more precise.
+
+In the next tutorial, we will compare this result to one inferred via a graphical model.
+"""
diff --git a/scripts/howtofit/chapter_3_graphical_models/tutorial_2_graphical_model.py b/scripts/howtofit/chapter_3_graphical_models/tutorial_2_graphical_model.py
index 249d41dc..531024f1 100644
--- a/scripts/howtofit/chapter_3_graphical_models/tutorial_2_graphical_model.py
+++ b/scripts/howtofit/chapter_3_graphical_models/tutorial_2_graphical_model.py
@@ -1,287 +1,283 @@
-"""
-Tutorial 2: Graphical Models
-============================
-
-We have fitted a dataset containing 5 noisy 1D Gaussian which had a shared `centre` value. We estimated
-the `centre` by fitting each dataset individually and combining the value of the `centre` inferred by each fit into
-an overall estimate, using a weighted average.
-
-Graphical models use a different approach. They are a single model that is fitted to the entire dataset simultaneously.
-The model includes specific model component for every individual 1D Gaussian in the sample. However, the graphical
-model also has shared parameters between these individual model components.
-
-This example fits a graphical model using the same sample fitted in the previous tutorial, consisting of many 1D
-Gaussians. However, whereas previously the `centre` of each Gaussian was a free parameter in each fit, in the graphical
-model there is only a single parameter for the `centre` shared by all 1D Gaussians.
-
-This graphical model creates a non-linear parameter space with parameters for every Gaussian in our sample. For 5
-Gaussians each with their own model parameters but a single shared centre:
-
- - Each Gaussian has 2 free parameters from the components that are not shared (`normalization`, `sigma`).
- - There is one additional free parameter, which is the `centre` shared by all 5 Gaussians.
-
-__Contents__
-
-This tutorial is split into the following sections:
-
-- **Example Source Code (`af.ex`)**: The example objects used in this tutorial.
-- **Dataset**: Load the 5 noisy 1D Gaussian datasets for simultaneous fitting.
-- **Analysis**: Create Analysis objects for each dataset.
-- **Model**: Set up the graphical model with a shared prior for the centre parameter.
-- **Analysis Factors**: Pair each model with its corresponding Analysis class at factor graph nodes.
-- **Factor Graph**: Combine the Analysis Factors into a factor graph representing the graphical model.
-- **Search**: Configure and run the non-linear search to fit the factor graph.
-- **Result**: Inspect and compare the graphical model results to the individual fits.
-- **Wrap Up**: Summary and discussion of the benefits of graphical models.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import numpy as np
-from os import path
-
-import autofit as af
-
-"""
-__Example Source Code (`af.ex`)__
-
-The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
-
- - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
- `visualize` functions.
-
- - `Gaussian`: a model component representing a 1D Gaussian profile.
-
- - `plot_profile_1d`: a function for plotting 1D profile datasets including their noise.
-
-These are functionally identical to the `Analysis`, `Gaussian` and `plot_profile_1d` objects and functions you
-have seen and used elsewhere throughout the workspace.
-
-__Dataset__
-
-For each dataset we now set up the correct path and load it.
-
-Whereas in the previous tutorial we fitted each dataset one-by-one, in this tutorial we instead store each dataset
-in a list so that we can set up a single model-fit that fits the 5 datasets simultaneously.
-"""
-total_datasets = 5
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(
- path.join("dataset", "example_1d", "gaussian_x1__low_snr", "dataset_0")
-):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-dataset_name_list = []
-data_list = []
-noise_map_list = []
-
-for dataset_index in range(total_datasets):
- dataset_name = f"dataset_{dataset_index}"
-
- dataset_path = path.join(
- "dataset", "example_1d", "gaussian_x1__low_snr", dataset_name
- )
-
- data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
- noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
- )
-
- dataset_name_list.append(dataset_name)
- data_list.append(data)
- noise_map_list.append(noise_map)
-
-"""
-By plotting the Gaussians we can remind ourselves that determining their centres by eye is difficult.
-"""
-for dataset_name, data in zip(dataset_name_list, data_list):
- af.ex.plot_profile_1d(
- xvalues=np.arange(data.shape[0]),
- profile_1d=data,
- title=dataset_name,
- ylabel="Data Values",
- color="k",
- )
-
-"""
-__Analysis__
-
-For each dataset we now create a corresponding `Analysis` class.
-"""
-analysis_list = []
-
-for data, noise_map in zip(data_list, noise_map_list):
- analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
- analysis_list.append(analysis)
-
-"""
-__Model__
-
-We now compose the graphical model that we fit, using the `Model` object you are now familiar with.
-
-We begin by setting up a shared prior for `centre`.
-
-We set up this up as a single `UniformPrior` which is passed to separate `Model`'s for each `Gaussian` below.
-"""
-centre_shared_prior = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-
-"""
-We now set up a list of `Model`'s, each of which contain a `Gaussian` that is used to fit each of the datasets
-loaded above.
-
-All of these models use the `centre_shared_prior`, meaning that all model-components use the same value of `centre`
-for every individual model component.
-
-For a fit using five Gaussians, this reduces the dimensionality of parameter space from N=15 (e.g. 3 parameters per
-Gaussian) to N=11 (e.g. 5 `sigma`'s 5 `normalizations` and 1 `centre`).
-"""
-model_list = []
-
-for model_index in range(len(data_list)):
- gaussian = af.Model(af.ex.Gaussian)
-
- gaussian.centre = centre_shared_prior # This prior is used by all 3 Gaussians!
- gaussian.normalization = af.LogUniformPrior(lower_limit=1e-6, upper_limit=1e6)
- gaussian.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=25.0)
-
- model_list.append(gaussian)
-
-"""
-__Analysis Factors__
-
-Above, we composed a model consisting of three `Gaussian`'s with a shared `centre` prior. We also loaded three datasets
-which we intend to fit with each of these `Gaussians`, setting up each in an `Analysis` class that defines how the
-model is used to fit the data.
-
-We now simply pair each model-component to each `Analysis` class, so that:
-
-- `gaussian_0` fits `data_0` via `analysis_0`.
-- `gaussian_1` fits `data_1` via `analysis_1`.
-- `gaussian_2` fits `data_2` via `analysis_2`.
-
-The point where a `Model` and `Analysis` class meet is called an `AnalysisFactor`.
-
-This term denotes that we are composing a graphical model, which is commonly called a 'factor graph'. A factor
-defines a node on this graph where we have some data, a model, and we fit the two together. The 'links' between these
-different nodes then define the global model we are fitting.
-"""
-analysis_factor_list = []
-
-for model, analysis in zip(model_list, analysis_list):
- analysis_factor = af.AnalysisFactor(prior_model=model, analysis=analysis)
-
- analysis_factor_list.append(analysis_factor)
-
-"""
-__Factor Graph__
-
-We now combine our `AnalysisFactor`'s to compose a factor graph.
-
-What is a factor graph? A factor graph defines the graphical model's graph. For example, it defines the different
-model components that make up our model (e.g. the individual `Gaussian` classes) and how their parameters are linked or
-shared (e.g. that each `Gaussian` has its own unique `normalization` and `sigma`, but a shared `centre` parameter).
-
-This is what our factor graph looks like (visualization of graphs not implemented yet):
-
-The factor graph above is made up of two components:
-
-- Nodes: these are points on the graph where we have a unique set of data and a model that is made up of a subset of
-our overall graphical model. This is effectively the `AnalysisFactor` objects we created above.
-
-- Links: these define the model components and parameters that are shared across different nodes and thus retain the
-same values when fitting different datasets.
-"""
-factor_graph = af.FactorGraphModel(*analysis_factor_list)
-
-"""
-The fit will use the factor graph's `global_prior_model`, which uses the models contained in every analysis factor
-to contrast the overall global model that is fitted.
-
-Printing the `info` attribute of this model reveals the overall structure of the model, which is grouped in terms
-of the analysis factors and therefore datasets.
-"""
-print(factor_graph.global_prior_model.info)
-
-"""
-__Search__
-
-We can now create a non-linear search and use it to the fit the factor graph, using its `global_prior_model` property.
-"""
-search = af.DynestyStatic(
- path_prefix=path.join("howtofit", "chapter_graphical_models"),
- name="tutorial_2_graphical_model",
- nlive=200,
- dlogz=1e-4,
- sample="rwalk",
- walks=10,
-)
-
-result = search.fit(model=factor_graph.global_prior_model, analysis=factor_graph)
-
-"""
-__Result__
-
-The result's `info` attribute shows that the result is expressed following the same struture of analysis factors
-that the `global_prior_model.info` attribute revealed above.
-"""
-print(result.info)
-
-"""
-We can now inspect the inferred value of `centre`, and compare this to the value we estimated in the previous tutorial
-via a weighted average or posterior multiplicaition using KDE.(feature missing currently).
-
-(The errors of the weighted average and KDE below is what was estimated for a run on my PC, yours may be slightly
-different!)
-"""
-print(
- f"Weighted Average Centre Estimate = 48.535531422571886 (4.139907734505303) [1.0 sigma confidence intervals] \n"
-)
-
-centre = result.samples.median_pdf()[0].centre
-
-u1_error = result.samples.values_at_upper_sigma(sigma=1.0)[0].centre
-l1_error = result.samples.values_at_lower_sigma(sigma=1.0)[0].centre
-
-u3_error = result.samples.values_at_upper_sigma(sigma=3.0)[0].centre
-l3_error = result.samples.values_at_lower_sigma(sigma=3.0)[0].centre
-
-print("Inferred value of the shared centre via a graphical model fit: \n")
-print(f"{centre} ({l1_error} {u1_error}) [1.0 sigma confidence intervals]")
-print(f"{centre} ({l3_error} {u3_error}) [3.0 sigma confidence intervals]")
-
-"""
-The graphical model's centre estimate and errors are pretty much exactly the same as the weighted average or KDE!
-
-Whats the point of fitting a graphical model if the much simpler approach of the previous tutorial gives the
-same answer?
-
-The answer, is model complexity. Graphical models become more powerful as we make our model more complex,
-our non-linear parameter space higher dimensionality and the degeneracies between different parameters on the graph
-more significant.
-
-We will demonstrate this in the next tutorial.
-
-__Wrap Up__
-
-In this tutorial, we showed that for our extremely simple model the graphical model gives pretty much the
-same estimate of the 1D Gaussian centre's as simpler approaches followed in the previous tutorial.
-
-We will next show the strengths of graphical models by fitting more complex models.
-"""
+"""
+Tutorial 2: Graphical Models
+============================
+
+We have fitted a dataset containing 5 noisy 1D Gaussian which had a shared `centre` value. We estimated
+the `centre` by fitting each dataset individually and combining the value of the `centre` inferred by each fit into
+an overall estimate, using a weighted average.
+
+Graphical models use a different approach. They are a single model that is fitted to the entire dataset simultaneously.
+The model includes specific model component for every individual 1D Gaussian in the sample. However, the graphical
+model also has shared parameters between these individual model components.
+
+This example fits a graphical model using the same sample fitted in the previous tutorial, consisting of many 1D
+Gaussians. However, whereas previously the `centre` of each Gaussian was a free parameter in each fit, in the graphical
+model there is only a single parameter for the `centre` shared by all 1D Gaussians.
+
+This graphical model creates a non-linear parameter space with parameters for every Gaussian in our sample. For 5
+Gaussians each with their own model parameters but a single shared centre:
+
+ - Each Gaussian has 2 free parameters from the components that are not shared (`normalization`, `sigma`).
+ - There is one additional free parameter, which is the `centre` shared by all 5 Gaussians.
+
+__Contents__
+
+This tutorial is split into the following sections:
+
+- **Example Source Code (`af.ex`)**: The example objects used in this tutorial.
+- **Dataset**: Load the 5 noisy 1D Gaussian datasets for simultaneous fitting.
+- **Analysis**: Create Analysis objects for each dataset.
+- **Model**: Set up the graphical model with a shared prior for the centre parameter.
+- **Analysis Factors**: Pair each model with its corresponding Analysis class at factor graph nodes.
+- **Factor Graph**: Combine the Analysis Factors into a factor graph representing the graphical model.
+- **Search**: Configure and run the non-linear search to fit the factor graph.
+- **Result**: Inspect and compare the graphical model results to the individual fits.
+- **Wrap Up**: Summary and discussion of the benefits of graphical models.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import numpy as np
+from os import path
+
+import autofit as af
+
+"""
+__Example Source Code (`af.ex`)__
+
+The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
+
+ - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
+ `visualize` functions.
+
+ - `Gaussian`: a model component representing a 1D Gaussian profile.
+
+ - `plot_profile_1d`: a function for plotting 1D profile datasets including their noise.
+
+These are functionally identical to the `Analysis`, `Gaussian` and `plot_profile_1d` objects and functions you
+have seen and used elsewhere throughout the workspace.
+
+__Dataset__
+
+For each dataset we now set up the correct path and load it.
+
+Whereas in the previous tutorial we fitted each dataset one-by-one, in this tutorial we instead store each dataset
+in a list so that we can set up a single model-fit that fits the 5 datasets simultaneously.
+"""
+total_datasets = 5
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(
+ path.join("dataset", "example_1d", "gaussian_x1__low_snr", "dataset_0")
+):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+dataset_name_list = []
+data_list = []
+noise_map_list = []
+
+for dataset_index in range(total_datasets):
+ dataset_name = f"dataset_{dataset_index}"
+
+ dataset_path = path.join(
+ "dataset", "example_1d", "gaussian_x1__low_snr", dataset_name
+ )
+
+ data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+ noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+ )
+
+ dataset_name_list.append(dataset_name)
+ data_list.append(data)
+ noise_map_list.append(noise_map)
+
+"""
+By plotting the Gaussians we can remind ourselves that determining their centres by eye is difficult.
+"""
+for dataset_name, data in zip(dataset_name_list, data_list):
+ af.ex.plot_profile_1d(
+ xvalues=np.arange(data.shape[0]),
+ profile_1d=data,
+ title=dataset_name,
+ ylabel="Data Values",
+ color="k",
+ )
+
+"""
+__Analysis__
+
+For each dataset we now create a corresponding `Analysis` class.
+"""
+analysis_list = []
+
+for data, noise_map in zip(data_list, noise_map_list):
+ analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+ analysis_list.append(analysis)
+
+"""
+__Model__
+
+We now compose the graphical model that we fit, using the `Model` object you are now familiar with.
+
+We begin by setting up a shared prior for `centre`.
+
+We set up this up as a single `UniformPrior` which is passed to separate `Model`'s for each `Gaussian` below.
+"""
+centre_shared_prior = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+
+"""
+We now set up a list of `Model`'s, each of which contain a `Gaussian` that is used to fit each of the datasets
+loaded above.
+
+All of these models use the `centre_shared_prior`, meaning that all model-components use the same value of `centre`
+for every individual model component.
+
+For a fit using five Gaussians, this reduces the dimensionality of parameter space from N=15 (e.g. 3 parameters per
+Gaussian) to N=11 (e.g. 5 `sigma`'s 5 `normalizations` and 1 `centre`).
+"""
+model_list = []
+
+for model_index in range(len(data_list)):
+ gaussian = af.Model(af.ex.Gaussian)
+
+ gaussian.centre = centre_shared_prior # This prior is used by all 3 Gaussians!
+ gaussian.normalization = af.LogUniformPrior(lower_limit=1e-6, upper_limit=1e6)
+ gaussian.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=25.0)
+
+ model_list.append(gaussian)
+
+"""
+__Analysis Factors__
+
+Above, we composed a model consisting of three `Gaussian`'s with a shared `centre` prior. We also loaded three datasets
+which we intend to fit with each of these `Gaussians`, setting up each in an `Analysis` class that defines how the
+model is used to fit the data.
+
+We now simply pair each model-component to each `Analysis` class, so that:
+
+- `gaussian_0` fits `data_0` via `analysis_0`.
+- `gaussian_1` fits `data_1` via `analysis_1`.
+- `gaussian_2` fits `data_2` via `analysis_2`.
+
+The point where a `Model` and `Analysis` class meet is called an `AnalysisFactor`.
+
+This term denotes that we are composing a graphical model, which is commonly called a 'factor graph'. A factor
+defines a node on this graph where we have some data, a model, and we fit the two together. The 'links' between these
+different nodes then define the global model we are fitting.
+"""
+analysis_factor_list = []
+
+for model, analysis in zip(model_list, analysis_list):
+ analysis_factor = af.AnalysisFactor(prior_model=model, analysis=analysis)
+
+ analysis_factor_list.append(analysis_factor)
+
+"""
+__Factor Graph__
+
+We now combine our `AnalysisFactor`'s to compose a factor graph.
+
+What is a factor graph? A factor graph defines the graphical model's graph. For example, it defines the different
+model components that make up our model (e.g. the individual `Gaussian` classes) and how their parameters are linked or
+shared (e.g. that each `Gaussian` has its own unique `normalization` and `sigma`, but a shared `centre` parameter).
+
+This is what our factor graph looks like (visualization of graphs not implemented yet):
+
+The factor graph above is made up of two components:
+
+- Nodes: these are points on the graph where we have a unique set of data and a model that is made up of a subset of
+our overall graphical model. This is effectively the `AnalysisFactor` objects we created above.
+
+- Links: these define the model components and parameters that are shared across different nodes and thus retain the
+same values when fitting different datasets.
+"""
+factor_graph = af.FactorGraphModel(*analysis_factor_list)
+
+"""
+The fit will use the factor graph's `global_prior_model`, which uses the models contained in every analysis factor
+to contrast the overall global model that is fitted.
+
+Printing the `info` attribute of this model reveals the overall structure of the model, which is grouped in terms
+of the analysis factors and therefore datasets.
+"""
+print(factor_graph.global_prior_model.info)
+
+"""
+__Search__
+
+We can now create a non-linear search and use it to the fit the factor graph, using its `global_prior_model` property.
+"""
+search = af.DynestyStatic(
+ path_prefix=path.join("howtofit", "chapter_graphical_models"),
+ name="tutorial_2_graphical_model",
+ nlive=200,
+ dlogz=1e-4,
+ sample="rwalk",
+ walks=10,
+)
+
+result = search.fit(model=factor_graph.global_prior_model, analysis=factor_graph)
+
+"""
+__Result__
+
+The result's `info` attribute shows that the result is expressed following the same struture of analysis factors
+that the `global_prior_model.info` attribute revealed above.
+"""
+print(result.info)
+
+"""
+We can now inspect the inferred value of `centre`, and compare this to the value we estimated in the previous tutorial
+via a weighted average or posterior multiplicaition using KDE.(feature missing currently).
+
+(The errors of the weighted average and KDE below is what was estimated for a run on my PC, yours may be slightly
+different!)
+"""
+print(
+ f"Weighted Average Centre Estimate = 48.535531422571886 (4.139907734505303) [1.0 sigma confidence intervals] \n"
+)
+
+centre = result.samples.median_pdf()[0].centre
+
+u1_error = result.samples.values_at_upper_sigma(sigma=1.0)[0].centre
+l1_error = result.samples.values_at_lower_sigma(sigma=1.0)[0].centre
+
+u3_error = result.samples.values_at_upper_sigma(sigma=3.0)[0].centre
+l3_error = result.samples.values_at_lower_sigma(sigma=3.0)[0].centre
+
+print("Inferred value of the shared centre via a graphical model fit: \n")
+print(f"{centre} ({l1_error} {u1_error}) [1.0 sigma confidence intervals]")
+print(f"{centre} ({l3_error} {u3_error}) [3.0 sigma confidence intervals]")
+
+"""
+The graphical model's centre estimate and errors are pretty much exactly the same as the weighted average or KDE!
+
+Whats the point of fitting a graphical model if the much simpler approach of the previous tutorial gives the
+same answer?
+
+The answer, is model complexity. Graphical models become more powerful as we make our model more complex,
+our non-linear parameter space higher dimensionality and the degeneracies between different parameters on the graph
+more significant.
+
+We will demonstrate this in the next tutorial.
+
+__Wrap Up__
+
+In this tutorial, we showed that for our extremely simple model the graphical model gives pretty much the
+same estimate of the 1D Gaussian centre's as simpler approaches followed in the previous tutorial.
+
+We will next show the strengths of graphical models by fitting more complex models.
+"""
diff --git a/scripts/howtofit/chapter_3_graphical_models/tutorial_3_graphical_benefits.py b/scripts/howtofit/chapter_3_graphical_models/tutorial_3_graphical_benefits.py
index 32281db9..63e1eb98 100644
--- a/scripts/howtofit/chapter_3_graphical_models/tutorial_3_graphical_benefits.py
+++ b/scripts/howtofit/chapter_3_graphical_models/tutorial_3_graphical_benefits.py
@@ -1,485 +1,481 @@
-"""
-Tutorial 3: Graphical Benefits
-==============================
-
-In the previous tutorials, we fitted a dataset containing 5 noisy 1D Gaussian which had a shared `centre` value and
-compared different approaches to estimate the shared `centre`. This included a simple approach fitting each dataset
-one-by-one and estimating the centre via a weighted average or posterior multiplication and a more complicated
-approach using a graphical model.
-
-The estimates were consistent with one another, making it hard to justify the use of the more complicated graphical
-model. However, the model fitted in the previous tutorial was extremely simple, and by making it slightly more complex
-we will show the benefits of the graphical model.
-
-__The Model__
-
-In this tutorial, each dataset now contains two Gaussians, and they all have the same shared centres, located at
-pixels 40 and 60.
-
-__Contents__
-
-This tutorial is split into the following sections:
-
-- **The Model**: Describe the two-Gaussian model fitted in this tutorial.
-- **Example Source Code (`af.ex`)**: The example objects used in this tutorial.
-- **Dataset**: Load datasets where each contains two Gaussians with shared centres.
-- **Analysis**: Create Analysis objects for each dataset.
-- **Model (one-by-one)**: Set up individual models with two Gaussians for one-by-one fitting.
-- **Model Fits (one-by-one)**: Fit each dataset individually using separate non-linear searches.
-- **Centre Estimates (Weighted Average)**: Compute centre estimates and errors using a weighted average.
-- **Discussion**: Analyze the limitations of the one-by-one fitting approach.
-- **Model (Graphical)**: Set up the graphical model with shared centre priors across datasets.
-- **Analysis Factors**: Create Analysis Factors pairing models with Analysis objects.
-- **Factor Graph**: Combine Analysis Factors into a factor graph.
-- **Search**: Configure and run the non-linear search for the graphical model.
-- **Result**: Inspect the graphical model results and compare to individual fits.
-- **Discussion**: Discuss the benefits of graphical models over one-by-one fitting.
-- **Posterior Multiplication**: Discuss KDE-based posterior multiplication as an alternative method.
-- **Wrap Up**: Summary comparing the different methods and transition to hierarchical models.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import numpy as np
-from os import path
-
-import autofit as af
-import autofit.plot as aplt
-
-"""
-__Example Source Code (`af.ex`)__
-
-The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
-
- - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
- `visualize` functions.
-
- - `Gaussian`: a model component representing a 1D Gaussian profile.
-
- - `plot_profile_1d`: a function for plotting 1D profile datasets including their noise.
-
-These are functionally identical to the `Analysis`, `Gaussian` and `plot_profile_1d` objects and functions you have seen
-and used elsewhere throughout the workspace.
-
-__Dataset__
-
-For each dataset we now set up the correct path and load it.
-
-Note that we are loading a new dataset called `gaussian_x2__offset_centres`.
-"""
-total_datasets = 5
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(
- path.join("dataset", "example_1d", "gaussian_x2__offset_centres", "dataset_0")
-):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-dataset_name_list = []
-data_list = []
-noise_map_list = []
-
-for dataset_index in range(total_datasets):
- dataset_name = f"dataset_{dataset_index}"
-
- dataset_path = path.join(
- "dataset", "example_1d", "gaussian_x2__offset_centres", dataset_name
- )
-
- data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
- noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
- )
-
- dataset_name_list.append(dataset_name)
- data_list.append(data)
- noise_map_list.append(noise_map)
-
-"""
-By plotting the datasets we see that each dataset contains two Gaussians.
-
-Their centres are offset from one another and not located at pixel 50, like in the previous tutorials.
-
-As discussed above, the Gaussians in every dataset are in facted centred at pixels 40 and 60.
-"""
-for dataset_name, data in zip(dataset_name_list, data_list):
- af.ex.plot_profile_1d(
- xvalues=np.arange(data.shape[0]),
- profile_1d=data,
- title=dataset_name,
- ylabel="Data Values",
- color="k",
- )
-
-"""
-__Analysis__
-
-For each dataset we now create a corresponding `Analysis` class.
-"""
-analysis_list = []
-
-for data, noise_map in zip(data_list, noise_map_list):
- analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
- analysis_list.append(analysis)
-
-"""
-__Model (one-by-one)__
-
-We are first going to fit each dataset one by one.
-
-Our model therefore now has two 1D `Gaussian`'s.
-
-To remove solutions where the Gaussians flip locations and fit the other Gaussian, we set uniform priors on the
-`centre`'s which ensures one Gaussian stays on the left side of the data (fitting the Gaussian at pixel 40)
-whilst the other stays on the right (fitting the Gaussian at pixel 60).
-"""
-gaussian_0 = af.Model(af.ex.Gaussian)
-
-gaussian_0.centre = af.UniformPrior(lower_limit=0.0, upper_limit=50.0)
-
-gaussian_1 = af.Model(af.ex.Gaussian)
-
-gaussian_1.centre = af.UniformPrior(lower_limit=50.0, upper_limit=100.0)
-
-model = af.Collection(gaussian_0=gaussian_0, gaussian_1=gaussian_1)
-
-"""
-__Model Fits (one-by-one)__
-
-For every dataset we now create an `Analysis` and fit it with a `Gaussian`.
-
-The `Result` is stored in the list `result_list`.
-"""
-result_list = []
-
-for i, analysis in enumerate(analysis_list):
- """
- Create the `DynestyStatic` non-linear search and use it to fit the data.
-
- We use custom dynesty settings which ensure the posterior is explored fully and that our error estimates are robust.
- """
- search = af.DynestyStatic(
- name=f"individual_fit_{i}",
- path_prefix=path.join(
- "howtofit", "chapter_graphical_models", "tutorial_3_graphical_benefits"
- ),
- nlive=200,
- dlogz=1e-4,
- sample="rwalk",
- walks=10,
- )
-
- print(
- f"The non-linear search has begun running, checkout \n"
- f"autofit_workspace/output/howtofit/chapter_graphical_models/tutorial_3_graphical_benefits/{dataset_name} for live \n"
- f"output of the results. This Jupyter notebook cell with progress once search has completed, this could take a \n"
- f"few minutes!"
- )
-
- result_list.append(search.fit(model=model, analysis=analysis))
-
-"""
-__Centre Estimates (Weighted Average)__
-
-We can now compute the centre estimate of both Gaussians, including their errors, from the individual model fits
-performed above.
-"""
-samples_list = [result.samples for result in result_list]
-
-mp_instances = [samps.median_pdf() for samps in samples_list]
-
-mp_centres_0 = [instance.gaussian_0.centre for instance in mp_instances]
-mp_centres_1 = [instance.gaussian_1.centre for instance in mp_instances]
-
-ue1_instances = [samp.values_at_upper_sigma(sigma=1.0) for samp in samples_list]
-le1_instances = [samp.values_at_lower_sigma(sigma=1.0) for samp in samples_list]
-
-ue1_centres_0 = [instance.gaussian_0.centre for instance in ue1_instances]
-le1_centres_0 = [instance.gaussian_0.centre for instance in le1_instances]
-
-error_0_list = [ue1 - le1 for ue1, le1 in zip(ue1_centres_0, le1_centres_0)]
-
-values_0 = np.asarray(mp_centres_0)
-sigmas_0 = np.asarray(error_0_list)
-
-weights_0 = 1 / sigmas_0**2.0
-weight_averaged_0 = np.sum(1.0 / sigmas_0**2)
-
-weighted_centre_0 = np.sum(values_0 * weights_0) / np.sum(weights_0, axis=0)
-weighted_error_0 = 1.0 / np.sqrt(weight_averaged_0)
-
-ue1_centres_1 = [instance.gaussian_1.centre for instance in ue1_instances]
-le1_centres_1 = [instance.gaussian_1.centre for instance in le1_instances]
-
-error_1_list = [ue1 - le1 for ue1, le1 in zip(ue1_centres_1, le1_centres_1)]
-
-values_1 = np.asarray(mp_centres_1)
-sigmas_1 = np.asarray(error_1_list)
-
-weights_1 = 1 / sigmas_1**2.0
-weight_averaged_1 = np.sum(1.0 / sigmas_1**2)
-
-weighted_centre_1 = np.sum(values_1 * weights_1) / np.sum(weights_1, axis=0)
-weighted_error_1 = 1.0 / np.sqrt(weight_averaged_1)
-
-
-print(
- f"Centre 0 via Weighted Average: {weighted_centre_0} ({weighted_error_0}) [1.0 sigma confidence intervals]"
-)
-print(
- f"Centre 1 via Weighted Average: {weighted_centre_1} ({weighted_error_1}) [1.0 sigma confidence intervals]"
-)
-
-"""
-The estimate of the centres is not accurate, with both estimates well offset from the input values of 40 and 60.
-
-We will next show that the graphical model offers a notable improvement, but first lets consider why this
-approach is suboptimal.
-
-The most important difference between this model and the model fitted in the previous tutorial is that there are now
-two shared parameters we are trying to estimate, which are degenerate with one another.
-
-We can see this by inspecting the probability distribution function (PDF) of the fit, placing particular focus on the
-2D degeneracy between the Gaussians centres.
-"""
-aplt.corner_cornerpy(samples=result_list[0].samples)
-
-"""
-The problem is that the simple approach of taking a weighted average does not capture the curved banana-like shape
-of the PDF between the two centres. This leads to significant error over estimation and biased inferences on the centre.
-
-__Discussion__
-
-Let us now consider other downsides of fitting each dataset one-by-one, from a statistical perspective. We
-will contrast these to the graphical model later in the tutorial:
-
-1) By fitting each dataset one-by-one this means that each model-fit fails to fully exploit the information we know
-about the global model. We know that there are only two single shared values of `centre` across the full dataset
-that we want to estimate. However, each individual fit has its own `centre` value which is able to assume different
-values than the `centre` values used to fit the other datasets. This means that large degeneracies between the two
-centres are present in each model-fit.
-
-By not fitting our model as a global model, we do not maximize the amount of information that we can extract from the
-dataset as a whole. If a model fits dataset 1 poorly, this should be reflected in how we interpret how well the model
-fits datasets 2 and 3. Our non-linear search should have a global view of how well the model fits the whole dataset.
-This is the crucial aspect of fitting each dataset individually that we miss, and what a graphical model addresses.
-
-2) When we combined the result to estimate the global `centre` value via a weighted average, we marginalized over
-the samples in 1D. As showed above, when there are strong degeneracies between models parameters the information on
-the covariance between these parameters is lost when computing the global `centre`. This increases the inferred
-uncertainties. A graphical model performs no such 1D marginalization and therefore fully samples the
-parameter covariances.
-
-3) In Bayesian inference it is important that we define priors on all of the model parameters. By estimating the
-global `centre` after the model-fits are completed it is unclear what prior the global `centre` actually has! We
-actually defined the prior five times -- once for each fit -- which is not a well defined prior. In a graphical model
-the prior is clearly defined.
-
-What would have happened if we had estimate the shared centres via 2D posterior multiplication using a KDE? We
-will discuss this at the end of the tutorial after fitting a graphical model.
-
-__Model (Graphical)__
-
-We now compose a graphical model and fit it.
-
-Our model now consists of two Gaussians with two `centre_shared_prior` variables, such that the same centres are
-used for each Gaussians across all datasets.
-
-We again restrict one Gaussian's centre between pixels 0 -> 50 and the other 50 -> 100 to remove solutions where
-the Gaussians flip location.
-"""
-centre_0_shared_prior = af.UniformPrior(lower_limit=0.0, upper_limit=50.0)
-centre_1_shared_prior = af.UniformPrior(lower_limit=50.0, upper_limit=100.0)
-
-"""
-We now set up a list of `Model`'s, each of which contain two `Gaussian`'s that are used to fit each of the datasets
-loaded above.
-
-All of these `Model`'s use the `centre_shared_prior`'s abpve. This means all model-components use the same value
-of `centre` for every model composed and fitted.
-
-For a fit to five datasets (each using two Gaussians), this reduces the dimensionality of parameter space
-from N=30 (e.g. 6 parameters per pair of Gaussians) to N=22 (e.g. 10 `sigma`'s 10 `normalizations` and 2 `centre`'s).
-"""
-model_list = []
-
-for model_index in range(len(data_list)):
- gaussian_0 = af.Model(af.ex.Gaussian)
- gaussian_1 = af.Model(af.ex.Gaussian)
-
- gaussian_0.centre = centre_0_shared_prior # This prior is used by all Gaussians!
- gaussian_1.centre = centre_1_shared_prior # This prior is used by all Gaussians!
-
- model = af.Collection(gaussian_0=gaussian_0, gaussian_1=gaussian_1)
-
- model_list.append(model)
-
-"""
-__Analysis Factors__
-
-We again create the graphical model using `AnalysisFactor` objects.
-"""
-analysis_factor_list = []
-
-for model, analysis in zip(model_list, analysis_list):
- analysis_factor = af.AnalysisFactor(prior_model=model, analysis=analysis)
-
- analysis_factor_list.append(analysis_factor)
-
-"""
-__Factor Graph__
-
-The analysis factors are then used to create the factor graph.
-"""
-factor_graph = af.FactorGraphModel(*analysis_factor_list)
-
-"""
-The factor graph model can again be printed via the `info` attribute, which shows that there are two shared
-parameters across the datasets.
-"""
-print(factor_graph.global_prior_model.info)
-
-"""
-__Search__
-
-We can now create a non-linear search and use it to the fit the factor graph, again using its `global_prior_model`
-property.
-"""
-search = af.DynestyStatic(
- path_prefix=path.join("howtofit", "chapter_graphical_models"),
- name="tutorial_3_graphical_benefits",
- sample="rwalk",
-)
-
-result = search.fit(model=factor_graph.global_prior_model, analysis=factor_graph)
-
-"""
-__Result__
-
-The result's `info` attribute shows that the result is expressed following the same structure of analysis factors
-that the `global_prior_model.info` attribute revealed above.
-"""
-print(result.info)
-
-"""
-We can now inspect the inferred `centre` values and compare this to the values estimated above via a weighted average.
-
-(The errors of the weighted average is what was estimated for a run on my PC, yours may be slightly different!)
-"""
-centre_0 = result.samples.median_pdf()[0].gaussian_0.centre
-
-u1_error_0 = result.samples.values_at_upper_sigma(sigma=1.0)[0].gaussian_0.centre
-l1_error_0 = result.samples.values_at_lower_sigma(sigma=1.0)[0].gaussian_0.centre
-
-u3_error_0 = result.samples.values_at_upper_sigma(sigma=3.0)[0].gaussian_0.centre
-l3_error_0 = result.samples.values_at_lower_sigma(sigma=3.0)[0].gaussian_0.centre
-
-centre_1 = result.samples.median_pdf()[0].gaussian_1.centre
-
-u1_error_1 = result.samples.values_at_upper_sigma(sigma=1.0)[0].gaussian_1.centre
-l1_error_1 = result.samples.values_at_lower_sigma(sigma=1.0)[0].gaussian_1.centre
-
-u3_error_1 = result.samples.values_at_upper_sigma(sigma=3.0)[0].gaussian_1.centre
-l3_error_1 = result.samples.values_at_lower_sigma(sigma=3.0)[0].gaussian_1.centre
-
-
-print(
- f"Centre 0 via Weighted Average: 29.415828686393333 (15.265325182888517) [1.0 sigma confidence intervals] \n"
-)
-print(
- f"Centre 1 via Weighted Average: 54.13825075629124 (2.3460686758693234) [1.0 sigma confidence intervals] \n"
-)
-
-print(
- f"Inferred value of Gaussian 0's shared centre via a graphical fit to {total_datasets} datasets: \n"
-)
-print(
- f"{centre_0} ({l1_error_0} {u1_error_0}) ({u1_error_0 - l1_error_0}) [1.0 sigma confidence intervals]"
-)
-print(
- f"{centre_0} ({l3_error_0} {u3_error_0}) ({u3_error_0 - l3_error_0}) [3.0 sigma confidence intervals]"
-)
-
-print(
- f"Inferred value of Gaussian 1's shared centre via a graphical fit to {total_datasets} datasets: \n"
-)
-print(
- f"{centre_1} ({l1_error_1} {u1_error_1}) ({u1_error_1 - l1_error_1}) [1.0 sigma confidence intervals]"
-)
-print(
- f"{centre_1} ({l3_error_1} {u3_error_1}) ({u3_error_1 - l3_error_1}) [3.0 sigma confidence intervals]"
-)
-
-"""
-As expected, using a graphical model allows us to infer a more precise and accurate model.
-
-__Discussion__
-
-Unlike a fit to each dataset one-by-one, the graphical model:
-
-1) Infers a PDF on the global centre that fully accounts for the degeneracies between the models fitted to different
-datasets. This reduces significantly the large 2D degeneracies between the two centres we saw when inspecting the PDFs
-of each individual fit.
-
-2) Fully exploits the information we know about the global model, for example that the centre of every Gaussian in every
-dataset is aligned. Now, the fit of the Gaussian in dataset 1 informs the fits in datasets 2 and 3, and visa versa.
-
-3) Has a well defined prior on the global centre, instead of 5 independent priors on the centre of each dataset.
-
-__Posterior Multiplication__
-
-What if we had combined the results of the individual model fits using 2D posterior multiplication via a KDE?
-
-This would produce an inaccurate estimate of the error, because each posterior contains the prior on the centre five
-times which given the properties of this model should not be repeated.
-
-However, it is possible to convert each posterior to a likelihood (by dividing by its prior), combining these 5
-likelihoods to form a joint likelihood via 2D KDE multiplication and then insert just one prior back (again using a 2D
-KDE) at the end to get a posterior which does not have repeated priors.
-
-This posterior, in theory, should be equivalent to the graphical model, giving the same accurate estimates of the
-centres with precise errors. The process extracts the same information, fully accounting for the 2D structure of the
-PDF between the two centres for each fit.
-
-However, in practise, this will likely not work well. Every time we use a KDE to represent and multiply a posterior, we
-make an approximation which will impact our inferred errors. The removal of the prior before combining the likelihood
-and reinserting it after also introduces approximations, especially because the fit performed by the non-linear search
-is informed by the prior.
-
-Crucially, whilst posterior multiplication can work in two dimensions, for models with many more dimensions and
-degeneracies between parameters that are in 3D, 4D of more dimensions it will introduce more and more numerical
-inaccuracies.
-
-A graphical model fully samples all of the information a large dataset contains about the model, without making
-such large approximation. Therefore, irrespective of how complex the model gets, it extracts significantly more
-information contained in the dataset.
-
-__Wrap Up__
-
-In this tutorial, we demonstrated the strengths of a graphical model over fitting each dataset one-by-one.
-
-We argued that irrespective of how one may try to combine the results of many individual fits, the approximations that
-are made will always lead to a suboptimal estimation of the model parameters and fail to fully extract all information
-from the dataset.
-
-We argued that for high dimensional complex models a graphical model is the only way to fully extract all of the
-information contained in the dataset.
-
-In the next tutorial, we will consider a natural extension of a graphical model called a hierarchical model.
-"""
+"""
+Tutorial 3: Graphical Benefits
+==============================
+
+In the previous tutorials, we fitted a dataset containing 5 noisy 1D Gaussian which had a shared `centre` value and
+compared different approaches to estimate the shared `centre`. This included a simple approach fitting each dataset
+one-by-one and estimating the centre via a weighted average or posterior multiplication and a more complicated
+approach using a graphical model.
+
+The estimates were consistent with one another, making it hard to justify the use of the more complicated graphical
+model. However, the model fitted in the previous tutorial was extremely simple, and by making it slightly more complex
+we will show the benefits of the graphical model.
+
+__The Model__
+
+In this tutorial, each dataset now contains two Gaussians, and they all have the same shared centres, located at
+pixels 40 and 60.
+
+__Contents__
+
+This tutorial is split into the following sections:
+
+- **The Model**: Describe the two-Gaussian model fitted in this tutorial.
+- **Example Source Code (`af.ex`)**: The example objects used in this tutorial.
+- **Dataset**: Load datasets where each contains two Gaussians with shared centres.
+- **Analysis**: Create Analysis objects for each dataset.
+- **Model (one-by-one)**: Set up individual models with two Gaussians for one-by-one fitting.
+- **Model Fits (one-by-one)**: Fit each dataset individually using separate non-linear searches.
+- **Centre Estimates (Weighted Average)**: Compute centre estimates and errors using a weighted average.
+- **Discussion**: Analyze the limitations of the one-by-one fitting approach.
+- **Model (Graphical)**: Set up the graphical model with shared centre priors across datasets.
+- **Analysis Factors**: Create Analysis Factors pairing models with Analysis objects.
+- **Factor Graph**: Combine Analysis Factors into a factor graph.
+- **Search**: Configure and run the non-linear search for the graphical model.
+- **Result**: Inspect the graphical model results and compare to individual fits.
+- **Discussion**: Discuss the benefits of graphical models over one-by-one fitting.
+- **Posterior Multiplication**: Discuss KDE-based posterior multiplication as an alternative method.
+- **Wrap Up**: Summary comparing the different methods and transition to hierarchical models.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import numpy as np
+from os import path
+
+import autofit as af
+import autofit.plot as aplt
+
+"""
+__Example Source Code (`af.ex`)__
+
+The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
+
+ - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
+ `visualize` functions.
+
+ - `Gaussian`: a model component representing a 1D Gaussian profile.
+
+ - `plot_profile_1d`: a function for plotting 1D profile datasets including their noise.
+
+These are functionally identical to the `Analysis`, `Gaussian` and `plot_profile_1d` objects and functions you have seen
+and used elsewhere throughout the workspace.
+
+__Dataset__
+
+For each dataset we now set up the correct path and load it.
+
+Note that we are loading a new dataset called `gaussian_x2__offset_centres`.
+"""
+total_datasets = 5
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(
+ path.join("dataset", "example_1d", "gaussian_x2__offset_centres", "dataset_0")
+):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+dataset_name_list = []
+data_list = []
+noise_map_list = []
+
+for dataset_index in range(total_datasets):
+ dataset_name = f"dataset_{dataset_index}"
+
+ dataset_path = path.join(
+ "dataset", "example_1d", "gaussian_x2__offset_centres", dataset_name
+ )
+
+ data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+ noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+ )
+
+ dataset_name_list.append(dataset_name)
+ data_list.append(data)
+ noise_map_list.append(noise_map)
+
+"""
+By plotting the datasets we see that each dataset contains two Gaussians.
+
+Their centres are offset from one another and not located at pixel 50, like in the previous tutorials.
+
+As discussed above, the Gaussians in every dataset are in facted centred at pixels 40 and 60.
+"""
+for dataset_name, data in zip(dataset_name_list, data_list):
+ af.ex.plot_profile_1d(
+ xvalues=np.arange(data.shape[0]),
+ profile_1d=data,
+ title=dataset_name,
+ ylabel="Data Values",
+ color="k",
+ )
+
+"""
+__Analysis__
+
+For each dataset we now create a corresponding `Analysis` class.
+"""
+analysis_list = []
+
+for data, noise_map in zip(data_list, noise_map_list):
+ analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+ analysis_list.append(analysis)
+
+"""
+__Model (one-by-one)__
+
+We are first going to fit each dataset one by one.
+
+Our model therefore now has two 1D `Gaussian`'s.
+
+To remove solutions where the Gaussians flip locations and fit the other Gaussian, we set uniform priors on the
+`centre`'s which ensures one Gaussian stays on the left side of the data (fitting the Gaussian at pixel 40)
+whilst the other stays on the right (fitting the Gaussian at pixel 60).
+"""
+gaussian_0 = af.Model(af.ex.Gaussian)
+
+gaussian_0.centre = af.UniformPrior(lower_limit=0.0, upper_limit=50.0)
+
+gaussian_1 = af.Model(af.ex.Gaussian)
+
+gaussian_1.centre = af.UniformPrior(lower_limit=50.0, upper_limit=100.0)
+
+model = af.Collection(gaussian_0=gaussian_0, gaussian_1=gaussian_1)
+
+"""
+__Model Fits (one-by-one)__
+
+For every dataset we now create an `Analysis` and fit it with a `Gaussian`.
+
+The `Result` is stored in the list `result_list`.
+"""
+result_list = []
+
+for i, analysis in enumerate(analysis_list):
+ """
+ Create the `DynestyStatic` non-linear search and use it to fit the data.
+
+ We use custom dynesty settings which ensure the posterior is explored fully and that our error estimates are robust.
+ """
+ search = af.DynestyStatic(
+ name=f"individual_fit_{i}",
+ path_prefix=path.join(
+ "howtofit", "chapter_graphical_models", "tutorial_3_graphical_benefits"
+ ),
+ nlive=200,
+ dlogz=1e-4,
+ sample="rwalk",
+ walks=10,
+ )
+
+ print(
+ f"The non-linear search has begun running, checkout \n"
+ f"autofit_workspace/output/howtofit/chapter_graphical_models/tutorial_3_graphical_benefits/{dataset_name} for live \n"
+ f"output of the results. This Jupyter notebook cell with progress once search has completed, this could take a \n"
+ f"few minutes!"
+ )
+
+ result_list.append(search.fit(model=model, analysis=analysis))
+
+"""
+__Centre Estimates (Weighted Average)__
+
+We can now compute the centre estimate of both Gaussians, including their errors, from the individual model fits
+performed above.
+"""
+samples_list = [result.samples for result in result_list]
+
+mp_instances = [samps.median_pdf() for samps in samples_list]
+
+mp_centres_0 = [instance.gaussian_0.centre for instance in mp_instances]
+mp_centres_1 = [instance.gaussian_1.centre for instance in mp_instances]
+
+ue1_instances = [samp.values_at_upper_sigma(sigma=1.0) for samp in samples_list]
+le1_instances = [samp.values_at_lower_sigma(sigma=1.0) for samp in samples_list]
+
+ue1_centres_0 = [instance.gaussian_0.centre for instance in ue1_instances]
+le1_centres_0 = [instance.gaussian_0.centre for instance in le1_instances]
+
+error_0_list = [ue1 - le1 for ue1, le1 in zip(ue1_centres_0, le1_centres_0)]
+
+values_0 = np.asarray(mp_centres_0)
+sigmas_0 = np.asarray(error_0_list)
+
+weights_0 = 1 / sigmas_0**2.0
+weight_averaged_0 = np.sum(1.0 / sigmas_0**2)
+
+weighted_centre_0 = np.sum(values_0 * weights_0) / np.sum(weights_0, axis=0)
+weighted_error_0 = 1.0 / np.sqrt(weight_averaged_0)
+
+ue1_centres_1 = [instance.gaussian_1.centre for instance in ue1_instances]
+le1_centres_1 = [instance.gaussian_1.centre for instance in le1_instances]
+
+error_1_list = [ue1 - le1 for ue1, le1 in zip(ue1_centres_1, le1_centres_1)]
+
+values_1 = np.asarray(mp_centres_1)
+sigmas_1 = np.asarray(error_1_list)
+
+weights_1 = 1 / sigmas_1**2.0
+weight_averaged_1 = np.sum(1.0 / sigmas_1**2)
+
+weighted_centre_1 = np.sum(values_1 * weights_1) / np.sum(weights_1, axis=0)
+weighted_error_1 = 1.0 / np.sqrt(weight_averaged_1)
+
+
+print(
+ f"Centre 0 via Weighted Average: {weighted_centre_0} ({weighted_error_0}) [1.0 sigma confidence intervals]"
+)
+print(
+ f"Centre 1 via Weighted Average: {weighted_centre_1} ({weighted_error_1}) [1.0 sigma confidence intervals]"
+)
+
+"""
+The estimate of the centres is not accurate, with both estimates well offset from the input values of 40 and 60.
+
+We will next show that the graphical model offers a notable improvement, but first lets consider why this
+approach is suboptimal.
+
+The most important difference between this model and the model fitted in the previous tutorial is that there are now
+two shared parameters we are trying to estimate, which are degenerate with one another.
+
+We can see this by inspecting the probability distribution function (PDF) of the fit, placing particular focus on the
+2D degeneracy between the Gaussians centres.
+"""
+aplt.corner_cornerpy(samples=result_list[0].samples)
+
+"""
+The problem is that the simple approach of taking a weighted average does not capture the curved banana-like shape
+of the PDF between the two centres. This leads to significant error over estimation and biased inferences on the centre.
+
+__Discussion__
+
+Let us now consider other downsides of fitting each dataset one-by-one, from a statistical perspective. We
+will contrast these to the graphical model later in the tutorial:
+
+1) By fitting each dataset one-by-one this means that each model-fit fails to fully exploit the information we know
+about the global model. We know that there are only two single shared values of `centre` across the full dataset
+that we want to estimate. However, each individual fit has its own `centre` value which is able to assume different
+values than the `centre` values used to fit the other datasets. This means that large degeneracies between the two
+centres are present in each model-fit.
+
+By not fitting our model as a global model, we do not maximize the amount of information that we can extract from the
+dataset as a whole. If a model fits dataset 1 poorly, this should be reflected in how we interpret how well the model
+fits datasets 2 and 3. Our non-linear search should have a global view of how well the model fits the whole dataset.
+This is the crucial aspect of fitting each dataset individually that we miss, and what a graphical model addresses.
+
+2) When we combined the result to estimate the global `centre` value via a weighted average, we marginalized over
+the samples in 1D. As showed above, when there are strong degeneracies between models parameters the information on
+the covariance between these parameters is lost when computing the global `centre`. This increases the inferred
+uncertainties. A graphical model performs no such 1D marginalization and therefore fully samples the
+parameter covariances.
+
+3) In Bayesian inference it is important that we define priors on all of the model parameters. By estimating the
+global `centre` after the model-fits are completed it is unclear what prior the global `centre` actually has! We
+actually defined the prior five times -- once for each fit -- which is not a well defined prior. In a graphical model
+the prior is clearly defined.
+
+What would have happened if we had estimate the shared centres via 2D posterior multiplication using a KDE? We
+will discuss this at the end of the tutorial after fitting a graphical model.
+
+__Model (Graphical)__
+
+We now compose a graphical model and fit it.
+
+Our model now consists of two Gaussians with two `centre_shared_prior` variables, such that the same centres are
+used for each Gaussians across all datasets.
+
+We again restrict one Gaussian's centre between pixels 0 -> 50 and the other 50 -> 100 to remove solutions where
+the Gaussians flip location.
+"""
+centre_0_shared_prior = af.UniformPrior(lower_limit=0.0, upper_limit=50.0)
+centre_1_shared_prior = af.UniformPrior(lower_limit=50.0, upper_limit=100.0)
+
+"""
+We now set up a list of `Model`'s, each of which contain two `Gaussian`'s that are used to fit each of the datasets
+loaded above.
+
+All of these `Model`'s use the `centre_shared_prior`'s abpve. This means all model-components use the same value
+of `centre` for every model composed and fitted.
+
+For a fit to five datasets (each using two Gaussians), this reduces the dimensionality of parameter space
+from N=30 (e.g. 6 parameters per pair of Gaussians) to N=22 (e.g. 10 `sigma`'s 10 `normalizations` and 2 `centre`'s).
+"""
+model_list = []
+
+for model_index in range(len(data_list)):
+ gaussian_0 = af.Model(af.ex.Gaussian)
+ gaussian_1 = af.Model(af.ex.Gaussian)
+
+ gaussian_0.centre = centre_0_shared_prior # This prior is used by all Gaussians!
+ gaussian_1.centre = centre_1_shared_prior # This prior is used by all Gaussians!
+
+ model = af.Collection(gaussian_0=gaussian_0, gaussian_1=gaussian_1)
+
+ model_list.append(model)
+
+"""
+__Analysis Factors__
+
+We again create the graphical model using `AnalysisFactor` objects.
+"""
+analysis_factor_list = []
+
+for model, analysis in zip(model_list, analysis_list):
+ analysis_factor = af.AnalysisFactor(prior_model=model, analysis=analysis)
+
+ analysis_factor_list.append(analysis_factor)
+
+"""
+__Factor Graph__
+
+The analysis factors are then used to create the factor graph.
+"""
+factor_graph = af.FactorGraphModel(*analysis_factor_list)
+
+"""
+The factor graph model can again be printed via the `info` attribute, which shows that there are two shared
+parameters across the datasets.
+"""
+print(factor_graph.global_prior_model.info)
+
+"""
+__Search__
+
+We can now create a non-linear search and use it to the fit the factor graph, again using its `global_prior_model`
+property.
+"""
+search = af.DynestyStatic(
+ path_prefix=path.join("howtofit", "chapter_graphical_models"),
+ name="tutorial_3_graphical_benefits",
+ sample="rwalk",
+)
+
+result = search.fit(model=factor_graph.global_prior_model, analysis=factor_graph)
+
+"""
+__Result__
+
+The result's `info` attribute shows that the result is expressed following the same structure of analysis factors
+that the `global_prior_model.info` attribute revealed above.
+"""
+print(result.info)
+
+"""
+We can now inspect the inferred `centre` values and compare this to the values estimated above via a weighted average.
+
+(The errors of the weighted average is what was estimated for a run on my PC, yours may be slightly different!)
+"""
+centre_0 = result.samples.median_pdf()[0].gaussian_0.centre
+
+u1_error_0 = result.samples.values_at_upper_sigma(sigma=1.0)[0].gaussian_0.centre
+l1_error_0 = result.samples.values_at_lower_sigma(sigma=1.0)[0].gaussian_0.centre
+
+u3_error_0 = result.samples.values_at_upper_sigma(sigma=3.0)[0].gaussian_0.centre
+l3_error_0 = result.samples.values_at_lower_sigma(sigma=3.0)[0].gaussian_0.centre
+
+centre_1 = result.samples.median_pdf()[0].gaussian_1.centre
+
+u1_error_1 = result.samples.values_at_upper_sigma(sigma=1.0)[0].gaussian_1.centre
+l1_error_1 = result.samples.values_at_lower_sigma(sigma=1.0)[0].gaussian_1.centre
+
+u3_error_1 = result.samples.values_at_upper_sigma(sigma=3.0)[0].gaussian_1.centre
+l3_error_1 = result.samples.values_at_lower_sigma(sigma=3.0)[0].gaussian_1.centre
+
+
+print(
+ f"Centre 0 via Weighted Average: 29.415828686393333 (15.265325182888517) [1.0 sigma confidence intervals] \n"
+)
+print(
+ f"Centre 1 via Weighted Average: 54.13825075629124 (2.3460686758693234) [1.0 sigma confidence intervals] \n"
+)
+
+print(
+ f"Inferred value of Gaussian 0's shared centre via a graphical fit to {total_datasets} datasets: \n"
+)
+print(
+ f"{centre_0} ({l1_error_0} {u1_error_0}) ({u1_error_0 - l1_error_0}) [1.0 sigma confidence intervals]"
+)
+print(
+ f"{centre_0} ({l3_error_0} {u3_error_0}) ({u3_error_0 - l3_error_0}) [3.0 sigma confidence intervals]"
+)
+
+print(
+ f"Inferred value of Gaussian 1's shared centre via a graphical fit to {total_datasets} datasets: \n"
+)
+print(
+ f"{centre_1} ({l1_error_1} {u1_error_1}) ({u1_error_1 - l1_error_1}) [1.0 sigma confidence intervals]"
+)
+print(
+ f"{centre_1} ({l3_error_1} {u3_error_1}) ({u3_error_1 - l3_error_1}) [3.0 sigma confidence intervals]"
+)
+
+"""
+As expected, using a graphical model allows us to infer a more precise and accurate model.
+
+__Discussion__
+
+Unlike a fit to each dataset one-by-one, the graphical model:
+
+1) Infers a PDF on the global centre that fully accounts for the degeneracies between the models fitted to different
+datasets. This reduces significantly the large 2D degeneracies between the two centres we saw when inspecting the PDFs
+of each individual fit.
+
+2) Fully exploits the information we know about the global model, for example that the centre of every Gaussian in every
+dataset is aligned. Now, the fit of the Gaussian in dataset 1 informs the fits in datasets 2 and 3, and visa versa.
+
+3) Has a well defined prior on the global centre, instead of 5 independent priors on the centre of each dataset.
+
+__Posterior Multiplication__
+
+What if we had combined the results of the individual model fits using 2D posterior multiplication via a KDE?
+
+This would produce an inaccurate estimate of the error, because each posterior contains the prior on the centre five
+times which given the properties of this model should not be repeated.
+
+However, it is possible to convert each posterior to a likelihood (by dividing by its prior), combining these 5
+likelihoods to form a joint likelihood via 2D KDE multiplication and then insert just one prior back (again using a 2D
+KDE) at the end to get a posterior which does not have repeated priors.
+
+This posterior, in theory, should be equivalent to the graphical model, giving the same accurate estimates of the
+centres with precise errors. The process extracts the same information, fully accounting for the 2D structure of the
+PDF between the two centres for each fit.
+
+However, in practise, this will likely not work well. Every time we use a KDE to represent and multiply a posterior, we
+make an approximation which will impact our inferred errors. The removal of the prior before combining the likelihood
+and reinserting it after also introduces approximations, especially because the fit performed by the non-linear search
+is informed by the prior.
+
+Crucially, whilst posterior multiplication can work in two dimensions, for models with many more dimensions and
+degeneracies between parameters that are in 3D, 4D of more dimensions it will introduce more and more numerical
+inaccuracies.
+
+A graphical model fully samples all of the information a large dataset contains about the model, without making
+such large approximation. Therefore, irrespective of how complex the model gets, it extracts significantly more
+information contained in the dataset.
+
+__Wrap Up__
+
+In this tutorial, we demonstrated the strengths of a graphical model over fitting each dataset one-by-one.
+
+We argued that irrespective of how one may try to combine the results of many individual fits, the approximations that
+are made will always lead to a suboptimal estimation of the model parameters and fail to fully extract all information
+from the dataset.
+
+We argued that for high dimensional complex models a graphical model is the only way to fully extract all of the
+information contained in the dataset.
+
+In the next tutorial, we will consider a natural extension of a graphical model called a hierarchical model.
+"""
diff --git a/scripts/howtofit/chapter_3_graphical_models/tutorial_4_hierachical_models.py b/scripts/howtofit/chapter_3_graphical_models/tutorial_4_hierachical_models.py
index 6373ab78..27e5b961 100644
--- a/scripts/howtofit/chapter_3_graphical_models/tutorial_4_hierachical_models.py
+++ b/scripts/howtofit/chapter_3_graphical_models/tutorial_4_hierachical_models.py
@@ -1,353 +1,349 @@
-"""
-Tutorial 4: Hierarchical
-========================
-
-In the previous tutorial, we fitted a graphical model with the aim of determining an estimate of shared parameters,
-the `centre`'s of a dataset of 1D Gaussians. We did this by fitting all datasets simultaneously. When there are shared
-parameters in a model, this is a powerful and effective tool, but things may not always be so simple.
-
-A common extension to the problem is one where we expect that the shared parameter(s) of the model do not have exactly
-the same value in every dataset. Instead, our expectation is that the parameter(s) are drawn from a common
-parent distribution (e.g. a Gaussian distribution). It is the parameters of this distribution that we consider shared
-across the dataset (e.g. the means and scatter of the Gaussian distribution). These are the parameters we ultimately
-wish to infer to understand the global behaviour of our model.
-
-This is called a hierarchical model, which we fit in this tutorial. The `centre` of each 1D Gaussian is now no
-longer the same in each dataset and they are instead drawn from a shared parent Gaussian distribution
-(with `mean=50.0` and `sigma=10.0`). The hierarchical model will recover the `mean` and `sigma` values of the parent
-distribution'.
-
-__Contents__
-
-This tutorial is split into the following sections:
-
-- **Example Source Code (`af.ex`)**: The example objects used in this tutorial.
-- **Dataset**: Load the hierarchical Gaussian datasets with variable centres.
-- **Analysis**: Create Analysis objects for each dataset.
-- **Model Individual Factors**: Set up individual Gaussian models with independent priors.
-- **Analysis Factors**: Compose Analysis Factors pairing models with Analysis objects.
-- **Model**: Create a HierarchicalFactor with a parent Gaussian distribution for the centres.
-- **Factor Graph**: Compose the factor graph including the hierarchical factor.
-- **Search**: Configure and run the non-linear search for the hierarchical model.
-- **Result**: Inspect the inferred hierarchical distribution parameters.
-- **Comparison to One-by-One Fits**: Compare the hierarchical model results to simpler individual fits.
-- **Benefits of Graphical Model**: Discuss how datasets inform one another through the hierarchical model.
-- **Wrap Up**: Summary and introduction to expectation propagation.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import numpy as np
-from os import path
-
-import autofit as af
-
-"""
-__Example Source Code (`af.ex`)__
-
-The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
-
- - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
- `visualize` functions.
-
- - `Gaussian`: a model component representing a 1D Gaussian profile.
-
- - `plot_profile_1d`: a function for plotting 1D profile datasets including their noise.
-
-These are functionally identical to the `Analysis`, `Gaussian` and `plot_profile_1d` objects and functions you
-have seen and used elsewhere throughout the workspace.
-
-__Dataset__
-
-For each dataset we now set up the correct path and load it.
-
-We are loading a new Gaussian dataset, where the Gaussians have different centres which were drawn from a parent
-Gaussian distribution with a mean centre value of 50.0 and sigma of 10.0.
-"""
-total_datasets = 5
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(
- path.join("dataset", "example_1d", "gaussian_x1__hierarchical", "dataset_0")
-):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-dataset_name_list = []
-data_list = []
-noise_map_list = []
-
-for dataset_index in range(total_datasets):
- dataset_name = f"dataset_{dataset_index}"
-
- dataset_path = path.join(
- "dataset", "example_1d", "gaussian_x1__hierarchical", dataset_name
- )
-
- data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
- noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
- )
-
- dataset_name_list.append(dataset_name)
- data_list.append(data)
- noise_map_list.append(noise_map)
-
-"""
-By plotting the Gaussians we can just about make out that their centres are not all at pixel 50, and are spread out
-around it (albeit its difficult to be sure, due to the low signal-to-noise of the data).
-"""
-for dataset_name, data in zip(dataset_name_list, data_list):
- af.ex.plot_profile_1d(
- xvalues=np.arange(data.shape[0]),
- profile_1d=data,
- title=dataset_name,
- ylabel="Data Values",
- color="k",
- )
-
-"""
-__Analysis__
-
-For each dataset we now create a corresponding `Analysis` class, like in the previous tutorial.
-"""
-analysis_list = []
-
-for data, noise_map in zip(data_list, noise_map_list):
- analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
- analysis_list.append(analysis)
-
-
-"""
-__Model Individual Factors__
-
-We first set up a model for each `Gaussian` which is individually fitted to each 1D dataset, which forms the
-factors on the factor graph we compose.
-
-This uses a nearly identical for loop to the previous tutorials, however a shared `centre` is no longer used and each
-`Gaussian` is given its own prior for the `centre`.
-
-We will see next how this `centre` is used to construct the hierarchical model.
-"""
-model_list = []
-
-for model_index in range(len(data_list)):
- gaussian = af.Model(af.ex.Gaussian)
-
- gaussian.centre = af.TruncatedGaussianPrior(
- mean=50.0, sigma=20.0, lower_limit=0.0, upper_limit=100.0
- )
- gaussian.normalization = af.TruncatedGaussianPrior(
- mean=3.0, sigma=5.0, lower_limit=0.0
- )
- gaussian.sigma = af.TruncatedGaussianPrior(mean=10.0, sigma=10.0, lower_limit=0.0)
-
- model_list.append(gaussian)
-
-"""
-__Analysis Factors__
-
-Now we have our `Analysis` classes and model components, we can compose our `AnalysisFactor`'s.
-
-These are composed in the same way as for the graphical model in the previous tutorial.
-"""
-analysis_factor_list = []
-
-for model, analysis in zip(model_list, analysis_list):
- analysis_factor = af.AnalysisFactor(prior_model=model, analysis=analysis)
-
- analysis_factor_list.append(analysis_factor)
-
-"""
-__Model__
-
-We now compose the hierarchical model that we fit, using the individual Gaussian model components created above.
-
-We first create a `HierarchicalFactor`, which represents the parent Gaussian distribution from which we will assume
-that the `centre` of each individual `Gaussian` dataset is drawn.
-
-For this parent `Gaussian`, we have to place priors on its `mean` and `sigma`, given that they are parameters in our
-model we are ultimately fitting for.
-"""
-hierarchical_factor = af.HierarchicalFactor(
- af.GaussianPrior,
- mean=af.TruncatedGaussianPrior(
- mean=50.0, sigma=10, lower_limit=0.0, upper_limit=100.0
- ),
- sigma=af.TruncatedGaussianPrior(
- mean=10.0, sigma=5.0, lower_limit=0.0, upper_limit=100.0
- ),
-)
-
-"""
-We now add each of the individual model `Gaussian`'s `centre` parameters to the `hierarchical_factor`.
-
-This composes the hierarchical model whereby the individual `centre` of every `Gaussian` in our dataset is now assumed
-to be drawn from a shared parent distribution. It is the `mean` and `sigma` of this distribution we are hoping to
-estimate.
-"""
-for model in model_list:
- hierarchical_factor.add_drawn_variable(model.centre)
-
-"""
-__Factor Graph__
-
-We now create the factor graph for this model, using the list of `AnalysisFactor`'s and the hierarchical factor.
-
-Note that the `hierarchical_factor` is passed in below, which was not the case in previous tutorials.
-"""
-factor_graph = af.FactorGraphModel(*analysis_factor_list, hierarchical_factor)
-
-"""
-The factor graph model `info` attribute shows that the hierarchical factor's parameters are included in the model.
-"""
-print(factor_graph.global_prior_model.info)
-
-"""
-__Search__
-
-We can now create a non-linear search and used it to the fit the hierarchical model, again using
-its `global_prior_model` property.
-"""
-search = af.DynestyStatic(
- path_prefix=path.join("howtofit", "chapter_graphical_models"),
- name="tutorial_4_hierarchical",
- sample="rwalk",
-)
-
-result = search.fit(model=factor_graph.global_prior_model, analysis=factor_graph)
-
-"""
-__Result__
-
-The result's `info` attribute shows the result, including the hierarchical factor's parameters.
-"""
-print(result.info)
-
-"""
-We can now inspect the inferred value of hierarchical factor's mean and sigma.
-
-We see that they are consistent with the input values of `mean=50.0` and `sigma=10.0`.
-
-The hierarchical factor results are at the end of the samples list, hence why we extract them using `[-1]` and [-2]`
-below.
-"""
-samples = result.samples
-
-mean = samples.median_pdf(as_instance=False)[-2]
-
-u1_error = samples.values_at_upper_sigma(sigma=1.0)[-2]
-l1_error = samples.values_at_lower_sigma(sigma=1.0)[-2]
-
-u3_error = samples.values_at_upper_sigma(sigma=3.0)[-2]
-l3_error = samples.values_at_lower_sigma(sigma=3.0)[-2]
-
-print(
- "Inferred value of the mean of the parent hierarchical distribution for the centre: \n"
-)
-print(f"{mean} ({l1_error} {u1_error}) [1.0 sigma confidence intervals]")
-print(f"{mean} ({l3_error} {u3_error}) [3.0 sigma confidence intervals]")
-
-scatter = samples.median_pdf(as_instance=False)[-1]
-
-u1_error = samples.values_at_upper_sigma(sigma=1.0)[-1]
-l1_error = samples.values_at_lower_sigma(sigma=1.0)[-1]
-
-u3_error = samples.values_at_upper_sigma(sigma=3.0)[-1]
-l3_error = samples.values_at_lower_sigma(sigma=3.0)[-1]
-
-print(
- "Inferred value of the scatter (the sigma value of the Gassuain) of the parent hierarchical distribution for the centre: \n"
-)
-print(f"{scatter} ({l1_error} {u1_error}) [1.0 sigma confidence intervals]")
-print(f"{scatter} ({l3_error} {u3_error}) [3.0 sigma confidence intervals]")
-
-"""
-__Comparison to One-by-One Fits__
-
-We can compare the inferred values above to the values inferred for individual fits in the
-tutorial `tutorial_optional_hierarchical_individual.py`.
-
-This fits the hierarchical model is a much simpler way -- fitting each dataset one-by-one and then fitting the
-parent Gaussian distribution to those results.
-
-For the results below, inferred on my laptop, we can see that the correct mean and scatter of the parent Gaussian is
-inferred but the errors are much larger than the graphical model fit above.
-"""
-print(
- "Inferred value of the mean of the parent hierarchical distribution for one-by-one fits: \n"
-)
-print(
- "50.00519854538594 (35.825675441265815 65.56274024242403) [1.0 sigma confidence intervals]"
-)
-print(
- "50.00519854538594 (1.3226539914914734 96.92151898283811) [3.0 sigma confidence intervals]"
-)
-
-print(
- "Inferred value of the scatter of the parent hierarchical distribution for one-by-one fits: \n"
-)
-print(
- "15.094393493747617 (4.608862348173649 31.346751522582483) [1.0 sigma confidence intervals]"
-)
-print(
- "15.094393493747617 (0.060533647989089806 49.05537884440667) [3.0 sigma confidence intervals]"
-)
-
-"""
-__Benefits of Graphical Model__
-
-We compared the results inferred in this script via a graphical model to a simpler approach which fits each dataset
-one-by-one and infers the hierarchical parent distribution's parameters afterwards.
-
-The graphical model provides a more accurate and precise estimate of the parent distribution's parameters. This is
-because the fit to each dataset informs the hierarchical distribution's parameters, which in turn improves
-constraints on the other datasets. In a hierarchical fit, we describe this as "the datasets talking to one another".
-
-For example, by itself, dataset_0 may give weak constraints on the centre spanning the range 20 -> 85 at 1 sigma
-confidence. Now, consider if simultaneously all of the other datasets provide strong constraints on the
-hierarchical's distribution's parameters, such that its `mean = 50 +- 5.0` and `sigma = 10.0 +- 2.0` at 1 sigma
-confidence.
-
-This will significantly change our inferred parameters for dataset 0, as the other datasets inform us
-that solutions where the centre is well below approximately 40 are less likely, because they are inconsistent with
-the parent hierarchical distribution's parameters!
-
-For complex graphical models with many hierarchical factors, this phenomena of the "datasets talking to one another"
-is crucial in breaking degeneracies between parameters and maximally extracting information from extremely large
-datasets.
-
-__Wrap Up__
-
-By composing and fitting hierarchical models in the graphical modeling framework we can fit for global trends
-within large datasets. The tools applied in this tutorial and the previous tutorial can be easily extended to
-compose complex graphical models, with multiple shared parameters and hierarchical factors.
-
-However, there is a clear challenge scaling the graphical modeling framework up in this way: model complexity. As the
-model becomes more complex, an inadequate sampling of parameter space will lead one to infer local maxima. Furthermore,
-one will soon hit computational limits on how many datasets can feasibly be fitted simultaneously, both in terms of
-CPU time and memory limitations.
-
-Therefore, the next tutorial introduces expectation propagation, a framework that inspects the factor graph of a
-graphical model and partitions the model-fit into many separate fits on each graph node. When a fit is complete,
-it passes the information learned about the model to neighboring nodes.
-
-Therefore, graphs comprising hundreds of model components (and tens of thousands of parameters) can be fitted as
-many bite-sized model fits, where the model fitted at each node consists of just tens of parameters. This makes
-graphical models scalable to largest datasets and most complex models!
-"""
+"""
+Tutorial 4: Hierarchical
+========================
+
+In the previous tutorial, we fitted a graphical model with the aim of determining an estimate of shared parameters,
+the `centre`'s of a dataset of 1D Gaussians. We did this by fitting all datasets simultaneously. When there are shared
+parameters in a model, this is a powerful and effective tool, but things may not always be so simple.
+
+A common extension to the problem is one where we expect that the shared parameter(s) of the model do not have exactly
+the same value in every dataset. Instead, our expectation is that the parameter(s) are drawn from a common
+parent distribution (e.g. a Gaussian distribution). It is the parameters of this distribution that we consider shared
+across the dataset (e.g. the means and scatter of the Gaussian distribution). These are the parameters we ultimately
+wish to infer to understand the global behaviour of our model.
+
+This is called a hierarchical model, which we fit in this tutorial. The `centre` of each 1D Gaussian is now no
+longer the same in each dataset and they are instead drawn from a shared parent Gaussian distribution
+(with `mean=50.0` and `sigma=10.0`). The hierarchical model will recover the `mean` and `sigma` values of the parent
+distribution'.
+
+__Contents__
+
+This tutorial is split into the following sections:
+
+- **Example Source Code (`af.ex`)**: The example objects used in this tutorial.
+- **Dataset**: Load the hierarchical Gaussian datasets with variable centres.
+- **Analysis**: Create Analysis objects for each dataset.
+- **Model Individual Factors**: Set up individual Gaussian models with independent priors.
+- **Analysis Factors**: Compose Analysis Factors pairing models with Analysis objects.
+- **Model**: Create a HierarchicalFactor with a parent Gaussian distribution for the centres.
+- **Factor Graph**: Compose the factor graph including the hierarchical factor.
+- **Search**: Configure and run the non-linear search for the hierarchical model.
+- **Result**: Inspect the inferred hierarchical distribution parameters.
+- **Comparison to One-by-One Fits**: Compare the hierarchical model results to simpler individual fits.
+- **Benefits of Graphical Model**: Discuss how datasets inform one another through the hierarchical model.
+- **Wrap Up**: Summary and introduction to expectation propagation.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import numpy as np
+from os import path
+
+import autofit as af
+
+"""
+__Example Source Code (`af.ex`)__
+
+The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
+
+ - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
+ `visualize` functions.
+
+ - `Gaussian`: a model component representing a 1D Gaussian profile.
+
+ - `plot_profile_1d`: a function for plotting 1D profile datasets including their noise.
+
+These are functionally identical to the `Analysis`, `Gaussian` and `plot_profile_1d` objects and functions you
+have seen and used elsewhere throughout the workspace.
+
+__Dataset__
+
+For each dataset we now set up the correct path and load it.
+
+We are loading a new Gaussian dataset, where the Gaussians have different centres which were drawn from a parent
+Gaussian distribution with a mean centre value of 50.0 and sigma of 10.0.
+"""
+total_datasets = 5
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(
+ path.join("dataset", "example_1d", "gaussian_x1__hierarchical", "dataset_0")
+):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+dataset_name_list = []
+data_list = []
+noise_map_list = []
+
+for dataset_index in range(total_datasets):
+ dataset_name = f"dataset_{dataset_index}"
+
+ dataset_path = path.join(
+ "dataset", "example_1d", "gaussian_x1__hierarchical", dataset_name
+ )
+
+ data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+ noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+ )
+
+ dataset_name_list.append(dataset_name)
+ data_list.append(data)
+ noise_map_list.append(noise_map)
+
+"""
+By plotting the Gaussians we can just about make out that their centres are not all at pixel 50, and are spread out
+around it (albeit its difficult to be sure, due to the low signal-to-noise of the data).
+"""
+for dataset_name, data in zip(dataset_name_list, data_list):
+ af.ex.plot_profile_1d(
+ xvalues=np.arange(data.shape[0]),
+ profile_1d=data,
+ title=dataset_name,
+ ylabel="Data Values",
+ color="k",
+ )
+
+"""
+__Analysis__
+
+For each dataset we now create a corresponding `Analysis` class, like in the previous tutorial.
+"""
+analysis_list = []
+
+for data, noise_map in zip(data_list, noise_map_list):
+ analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+ analysis_list.append(analysis)
+
+
+"""
+__Model Individual Factors__
+
+We first set up a model for each `Gaussian` which is individually fitted to each 1D dataset, which forms the
+factors on the factor graph we compose.
+
+This uses a nearly identical for loop to the previous tutorials, however a shared `centre` is no longer used and each
+`Gaussian` is given its own prior for the `centre`.
+
+We will see next how this `centre` is used to construct the hierarchical model.
+"""
+model_list = []
+
+for model_index in range(len(data_list)):
+ gaussian = af.Model(af.ex.Gaussian)
+
+ gaussian.centre = af.TruncatedGaussianPrior(
+ mean=50.0, sigma=20.0, lower_limit=0.0, upper_limit=100.0
+ )
+ gaussian.normalization = af.TruncatedGaussianPrior(
+ mean=3.0, sigma=5.0, lower_limit=0.0
+ )
+ gaussian.sigma = af.TruncatedGaussianPrior(mean=10.0, sigma=10.0, lower_limit=0.0)
+
+ model_list.append(gaussian)
+
+"""
+__Analysis Factors__
+
+Now we have our `Analysis` classes and model components, we can compose our `AnalysisFactor`'s.
+
+These are composed in the same way as for the graphical model in the previous tutorial.
+"""
+analysis_factor_list = []
+
+for model, analysis in zip(model_list, analysis_list):
+ analysis_factor = af.AnalysisFactor(prior_model=model, analysis=analysis)
+
+ analysis_factor_list.append(analysis_factor)
+
+"""
+__Model__
+
+We now compose the hierarchical model that we fit, using the individual Gaussian model components created above.
+
+We first create a `HierarchicalFactor`, which represents the parent Gaussian distribution from which we will assume
+that the `centre` of each individual `Gaussian` dataset is drawn.
+
+For this parent `Gaussian`, we have to place priors on its `mean` and `sigma`, given that they are parameters in our
+model we are ultimately fitting for.
+"""
+hierarchical_factor = af.HierarchicalFactor(
+ af.GaussianPrior,
+ mean=af.TruncatedGaussianPrior(
+ mean=50.0, sigma=10, lower_limit=0.0, upper_limit=100.0
+ ),
+ sigma=af.TruncatedGaussianPrior(
+ mean=10.0, sigma=5.0, lower_limit=0.0, upper_limit=100.0
+ ),
+)
+
+"""
+We now add each of the individual model `Gaussian`'s `centre` parameters to the `hierarchical_factor`.
+
+This composes the hierarchical model whereby the individual `centre` of every `Gaussian` in our dataset is now assumed
+to be drawn from a shared parent distribution. It is the `mean` and `sigma` of this distribution we are hoping to
+estimate.
+"""
+for model in model_list:
+ hierarchical_factor.add_drawn_variable(model.centre)
+
+"""
+__Factor Graph__
+
+We now create the factor graph for this model, using the list of `AnalysisFactor`'s and the hierarchical factor.
+
+Note that the `hierarchical_factor` is passed in below, which was not the case in previous tutorials.
+"""
+factor_graph = af.FactorGraphModel(*analysis_factor_list, hierarchical_factor)
+
+"""
+The factor graph model `info` attribute shows that the hierarchical factor's parameters are included in the model.
+"""
+print(factor_graph.global_prior_model.info)
+
+"""
+__Search__
+
+We can now create a non-linear search and used it to the fit the hierarchical model, again using
+its `global_prior_model` property.
+"""
+search = af.DynestyStatic(
+ path_prefix=path.join("howtofit", "chapter_graphical_models"),
+ name="tutorial_4_hierarchical",
+ sample="rwalk",
+)
+
+result = search.fit(model=factor_graph.global_prior_model, analysis=factor_graph)
+
+"""
+__Result__
+
+The result's `info` attribute shows the result, including the hierarchical factor's parameters.
+"""
+print(result.info)
+
+"""
+We can now inspect the inferred value of hierarchical factor's mean and sigma.
+
+We see that they are consistent with the input values of `mean=50.0` and `sigma=10.0`.
+
+The hierarchical factor results are at the end of the samples list, hence why we extract them using `[-1]` and [-2]`
+below.
+"""
+samples = result.samples
+
+mean = samples.median_pdf(as_instance=False)[-2]
+
+u1_error = samples.values_at_upper_sigma(sigma=1.0)[-2]
+l1_error = samples.values_at_lower_sigma(sigma=1.0)[-2]
+
+u3_error = samples.values_at_upper_sigma(sigma=3.0)[-2]
+l3_error = samples.values_at_lower_sigma(sigma=3.0)[-2]
+
+print(
+ "Inferred value of the mean of the parent hierarchical distribution for the centre: \n"
+)
+print(f"{mean} ({l1_error} {u1_error}) [1.0 sigma confidence intervals]")
+print(f"{mean} ({l3_error} {u3_error}) [3.0 sigma confidence intervals]")
+
+scatter = samples.median_pdf(as_instance=False)[-1]
+
+u1_error = samples.values_at_upper_sigma(sigma=1.0)[-1]
+l1_error = samples.values_at_lower_sigma(sigma=1.0)[-1]
+
+u3_error = samples.values_at_upper_sigma(sigma=3.0)[-1]
+l3_error = samples.values_at_lower_sigma(sigma=3.0)[-1]
+
+print(
+ "Inferred value of the scatter (the sigma value of the Gassuain) of the parent hierarchical distribution for the centre: \n"
+)
+print(f"{scatter} ({l1_error} {u1_error}) [1.0 sigma confidence intervals]")
+print(f"{scatter} ({l3_error} {u3_error}) [3.0 sigma confidence intervals]")
+
+"""
+__Comparison to One-by-One Fits__
+
+We can compare the inferred values above to the values inferred for individual fits in the
+tutorial `tutorial_optional_hierarchical_individual.py`.
+
+This fits the hierarchical model is a much simpler way -- fitting each dataset one-by-one and then fitting the
+parent Gaussian distribution to those results.
+
+For the results below, inferred on my laptop, we can see that the correct mean and scatter of the parent Gaussian is
+inferred but the errors are much larger than the graphical model fit above.
+"""
+print(
+ "Inferred value of the mean of the parent hierarchical distribution for one-by-one fits: \n"
+)
+print(
+ "50.00519854538594 (35.825675441265815 65.56274024242403) [1.0 sigma confidence intervals]"
+)
+print(
+ "50.00519854538594 (1.3226539914914734 96.92151898283811) [3.0 sigma confidence intervals]"
+)
+
+print(
+ "Inferred value of the scatter of the parent hierarchical distribution for one-by-one fits: \n"
+)
+print(
+ "15.094393493747617 (4.608862348173649 31.346751522582483) [1.0 sigma confidence intervals]"
+)
+print(
+ "15.094393493747617 (0.060533647989089806 49.05537884440667) [3.0 sigma confidence intervals]"
+)
+
+"""
+__Benefits of Graphical Model__
+
+We compared the results inferred in this script via a graphical model to a simpler approach which fits each dataset
+one-by-one and infers the hierarchical parent distribution's parameters afterwards.
+
+The graphical model provides a more accurate and precise estimate of the parent distribution's parameters. This is
+because the fit to each dataset informs the hierarchical distribution's parameters, which in turn improves
+constraints on the other datasets. In a hierarchical fit, we describe this as "the datasets talking to one another".
+
+For example, by itself, dataset_0 may give weak constraints on the centre spanning the range 20 -> 85 at 1 sigma
+confidence. Now, consider if simultaneously all of the other datasets provide strong constraints on the
+hierarchical's distribution's parameters, such that its `mean = 50 +- 5.0` and `sigma = 10.0 +- 2.0` at 1 sigma
+confidence.
+
+This will significantly change our inferred parameters for dataset 0, as the other datasets inform us
+that solutions where the centre is well below approximately 40 are less likely, because they are inconsistent with
+the parent hierarchical distribution's parameters!
+
+For complex graphical models with many hierarchical factors, this phenomena of the "datasets talking to one another"
+is crucial in breaking degeneracies between parameters and maximally extracting information from extremely large
+datasets.
+
+__Wrap Up__
+
+By composing and fitting hierarchical models in the graphical modeling framework we can fit for global trends
+within large datasets. The tools applied in this tutorial and the previous tutorial can be easily extended to
+compose complex graphical models, with multiple shared parameters and hierarchical factors.
+
+However, there is a clear challenge scaling the graphical modeling framework up in this way: model complexity. As the
+model becomes more complex, an inadequate sampling of parameter space will lead one to infer local maxima. Furthermore,
+one will soon hit computational limits on how many datasets can feasibly be fitted simultaneously, both in terms of
+CPU time and memory limitations.
+
+Therefore, the next tutorial introduces expectation propagation, a framework that inspects the factor graph of a
+graphical model and partitions the model-fit into many separate fits on each graph node. When a fit is complete,
+it passes the information learned about the model to neighboring nodes.
+
+Therefore, graphs comprising hundreds of model components (and tens of thousands of parameters) can be fitted as
+many bite-sized model fits, where the model fitted at each node consists of just tens of parameters. This makes
+graphical models scalable to largest datasets and most complex models!
+"""
diff --git a/scripts/howtofit/chapter_3_graphical_models/tutorial_5_expectation_propagation.py b/scripts/howtofit/chapter_3_graphical_models/tutorial_5_expectation_propagation.py
index ff077863..d1bcaed4 100644
--- a/scripts/howtofit/chapter_3_graphical_models/tutorial_5_expectation_propagation.py
+++ b/scripts/howtofit/chapter_3_graphical_models/tutorial_5_expectation_propagation.py
@@ -1,363 +1,359 @@
-"""
-Tutorial 5: Expectation Propagation
-===================================
-
-In the previous tutorials, we fitted graphical models to dataset comprising many noisy 1D Gaussians. These had a shared
-and global value of their `centre`, or assumed their centres were hierarchically drawn from a parent Gaussian
-distribution. This provides the basis of composing and fitting complex graphical models to large datasets.
-
-We concluded by discussing that there is a ceiling scaling these graphical models up to extremely large datasets. One
-would soon find that the parameter space is too complex to sample, and computational limits would ultimately cap how
-many datasets one could feasibly fit.
-
-This tutorial introduces expectation propagation (EP), the solution to this problem, which inspects a factor graph
-and partitions the model-fit into many simpler fits of sub-components of the graph to individual datasets. This
-overcomes the challenge of model complexity, and mitigates computational restrictions that may occur if one tries to
-fit every dataset simultaneously.
-
-This tutorial fits a global model with a shared parameter and does not use a hierarchical model. The optional tutorial
-`tutorial_optional_hierarchical_ep` shows an example fit of a hierarchical model with EP.
-
-__Contents__
-
-This tutorial is split into the following sections:
-
-- **Example Source Code (`af.ex`)**: The example objects used in this tutorial.
-- **Dataset**: Load the noisy 1D Gaussian datasets with a shared centre.
-- **Analysis**: Create Analysis objects for each dataset.
-- **Model**: Set up the model with a shared centre prior across all datasets.
-- **Analysis Factors**: Create Analysis Factors with individual searches for each dataset.
-- **Factor Graph**: Compose the factor graph for the EP framework.
-- **Expectation Propagation**: Explain the EP message passing algorithm.
-- **Cyclic Fitting**: Describe the iterative EP convergence process.
-- **Result**: Access the result of the EP fit.
-- **Output**: Describe the output directory structure and files generated by the EP fit.
-- **Results**: Use the MeanField object to infer parameter estimates and errors.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import numpy as np
-from os import path
-
-import autofit as af
-
-"""
-__Example Source Code (`af.ex`)__
-
-The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
-
- - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
- `visualize` functions.
-
- - `Gaussian`: a model component representing a 1D Gaussian profile.
-
- - `plot_profile_1d`: a function for plotting 1D profile datasets including their noise.
-
-These are functionally identical to the `Analysis`, `Gaussian` and `plot_profile_1d` objects and functions you have seen
-and used elsewhere throughout the workspace.
-
-__Dataset__
-
-For each dataset we now set up the correct path and load it.
-
-We first fit the 1D Gaussians which all share the same centre, thus not requiring a hierarchical model.
-
-An example for fitting the hierarchical model with EP is given at the end of this tutorial.
-"""
-total_datasets = 3
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(
- path.join("dataset", "example_1d", "gaussian_x1__low_snr", "dataset_0")
-):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-dataset_name_list = []
-data_list = []
-noise_map_list = []
-
-for dataset_index in range(total_datasets):
- dataset_name = f"dataset_{dataset_index}"
-
- dataset_path = path.join(
- "dataset", "example_1d", "gaussian_x1__low_snr", dataset_name
- )
-
- data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
- noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
- )
-
- dataset_name_list.append(dataset_name)
- data_list.append(data)
- noise_map_list.append(noise_map)
-
-"""
-By plotting the Gaussians we can remind ourselves that determining their centres by eye is difficult.
-"""
-for dataset_name, data in zip(dataset_name_list, data_list):
- af.ex.plot_profile_1d(
- xvalues=np.arange(data.shape[0]),
- profile_1d=data,
- title=dataset_name,
- ylabel="Data Values",
- color="k",
- )
-
-"""
-__Analysis__
-
-For each dataset we now create a corresponding `Analysis` class, like in the previous tutorial.
-"""
-analysis_list = []
-
-for data, noise_map in zip(data_list, noise_map_list):
- analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
- analysis_list.append(analysis)
-
-"""
-__Model__
-
-We now compose the graphical model that we fit, using the `Model` and `Collection` objects you are now familiar with.
-
-We will assume all Gaussians share the same centre, therefore we set up a shared prior for `centre`.
-"""
-centre_shared_prior = af.GaussianPrior(mean=50.0, sigma=30.0)
-
-model_list = []
-
-for model_index in range(len(data_list)):
- gaussian = af.Model(af.ex.Gaussian)
-
- gaussian.centre = af.GaussianPrior(mean=50.0, sigma=30.0)
- gaussian.normalization = af.TruncatedGaussianPrior(
- mean=3.0, sigma=5.0, lower_limit=0.0
- )
- gaussian.sigma = af.TruncatedGaussianPrior(mean=10.0, sigma=10.0, lower_limit=0.0)
-
- model = af.Collection(gaussian=gaussian)
-
- model_list.append(model)
-
-"""
-__Analysis Factors__
-
-Now we have our `Analysis` classes and graphical model, we can compose our `AnalysisFactor`'s.
-
-However, unlike the previous tutorials, each `AnalysisFactor` is now assigned its own `search`. This is because the EP
-framework performs a model-fit to each node on the factor graph (e.g. each `AnalysisFactor`). Therefore, each node
-requires its own non-linear search, and in this tutorial we use `dynesty`. For complex graphs consisting of many
-nodes, one could easily use different searches for different nodes on the factor graph.
-
-Each `AnalysisFactor` is also given a `name`, corresponding to the name of the dataset it fits. These names are used
-to name the folders containing the results in the output directory.
-"""
-paths = af.DirectoryPaths(
- name=path.join(
- "howtofit", "chapter_graphical_models", "tutorial_5_expectation_propagation"
- )
-)
-
-search = af.DynestyStatic(paths=paths, nlive=100, sample="rwalk")
-
-analysis_factor_list = []
-
-dataset_index = 0
-
-for model, analysis in zip(model_list, analysis_list):
- dataset_name = f"dataset_{dataset_index}"
- dataset_index += 1
-
- analysis_factor = af.AnalysisFactor(
- prior_model=model, analysis=analysis, optimiser=search, name=dataset_name
- )
-
- analysis_factor_list.append(analysis_factor)
-
-
-class LinearRegressionAnalysis(af.Analysis):
- def log_likelihood_function(self, instance, xp=np):
- return -1
-
-
-fwhm_list = [2 * np.sqrt(2 * np.log(2)) * model.gaussian.sigma for model in model_list]
-
-linear_regression_factor = af.AnalysisFactor(
- prior_model=af.Collection(
- m=af.GaussianPrior(mean=0.0, sigma=1.0),
- c=af.GaussianPrior(mean=0.0, sigma=1.0),
- *fwhm_list,
- ),
- analysis=LinearRegressionAnalysis(),
- optimiser=search,
- name="linear_regression",
-)
-
-"""
-__Factor Graph__
-
-We combine our `AnalysisFactors` into one, to compose the factor graph.
-"""
-factor_graph = af.FactorGraphModel(
- *analysis_factor_list,
- linear_regression_factor,
-)
-
-"""
-The factor graph model `info` attribute shows the model which we fit via expectaton propagation (note that we do
-not use `global_prior_model` below when performing the fit).
-"""
-print(factor_graph.global_prior_model.info)
-
-"""
-__Expectation Propagation__
-
-In the previous tutorials, we used the `global_prior_model` of the `factor_graph` to fit the global model. In this
-tutorial, we instead fit the `factor_graph` using the EP framework, which fits the graphical model composed in this
-tutorial as follows:
-
-1) Go to the first node on the factor graph (e.g. `analysis_factor_list[0]`) and fit its model to its dataset. This is
-simply a fit of the `Gaussian` model to the first 1D Gaussian dataset, the model-fit we are used to performing by now.
-
-2) Once the model-fit is complete, inspect the model for parameters that are shared with other nodes on the factor
-graph. In this example, the `centre` of the `Gaussian` fitted to the first dataset is global, and therefore connects
-to the other nodes on the factor graph (the `AnalysisFactor`'s) of the second and first `Gaussian` datasets.
-
-3) The EP framework now creates a 'message' that is to be passed to the connecting nodes on the factor graph. This
-message informs them of the results of the model-fit, so they can update their priors on the `Gaussian`'s centre
-accordingly and, more importantly, update their posterior inference and therefore estimate of the global centre.
-
-For example, the model fitted to the first Gaussian dataset includes the global centre. Therefore, after the model is
-fitted, the EP framework creates a 'message' informing the factor graph about its inference on that Gaussians's centre,
-thereby updating our overall inference on this shared parameter. This is termed 'message passing'.
-
-__Cyclic Fitting__
-
-After every `AnalysisFactor` has been fitted (e.g. after each fit to each of the 5 datasets in this example), we have a
-new estimate of the shared parameter `centre`. This updates our priors on the shared parameter `centre`, which needs
-to be reflected in each model-fit we perform on each `AnalysisFactor`.
-
-The EP framework therefore performs a second iteration of model-fits. It again cycles through each `AnalysisFactor`
-and refits the model, using updated priors on shared parameters like the `centre`. At the end of each fit, we again
-create messages that update our knowledge about other parameters on the graph.
-
-This process is repeated multiple times, until a convergence criteria is met whereby continued cycles are expected to
-produce the same estimate of the shared parameter `centre`.
-
-When we fit the factor graph a `name` is passed, which determines the folder all results of the factor graph are
-stored in.
-"""
-laplace = af.LaplaceOptimiser()
-
-factor_graph_result = factor_graph.optimise(
- optimiser=laplace, paths=paths, ep_history=af.EPHistory(kl_tol=0.05), max_steps=5
-)
-
-"""
-__Result__
-
-An `info` attribute for the result of a factor graph fitted via EP does not exist yet, its on the to do list!
-
-The result can be seen in the `graph.result` file output to hard-disk.
-"""
-### print(factor_graph_result.info)##
-
-"""
-__Output__
-
-The results of the factor graph, using the EP framework and message passing, are contained in the folder
-`output/howtofit/chapter_graphical_models/tutorial_5_expectation_propagation`.
-
-The following folders and files are worth of note:
-
- - `graph.info`: this provides an overall summary of the graphical model that is fitted, including every parameter,
- how parameters are shared across `AnalysisFactor`'s and the priors associated to each individual parameter.
-
- - The 3 folders titled `gaussian_x1_#__low_snr` correspond to the three `AnalysisFactor`'s and therefore signify
- repeated non-linear searches that are performed to fit each dataset.
-
- - Inside each of these folders are `optimization_#` folders, corresponding to each model-fit performed over cycles of
- the EP fit. A careful inspection of the `model.info` files inside each folder reveals how the priors are updated
- over each cycle, whereas the `model.results` file should indicate the improved estimate of model parameters over each
- cycle.
-
-__Results__
-
-The `MeanField` object represent the posterior of the entire factor graph and is used to infer estimates of the
-values and error of each parameter in the graph.
-"""
-mean_field = factor_graph_result.updated_ep_mean_field.mean_field
-print(mean_field)
-print()
-
-"""
-The object has a `variables` property which lists every variable in the factor graph, which is essentially all of the
-free parameters on the graph.
-
-This includes the parameters specific to each data (E.g. each node on the graph) as well as the shared centre.
-"""
-print(mean_field.variables)
-print()
-
-# """
-# The variables above use the priors on each parameter as their key.
-#
-# Therefore to estimate mean-field quantities of the shared centre, we can simply use the `centre_shared_prior` defined
-# above.
-#
-# Each parameter estimate is given by the mean of its value in the `MeanField`. Below, we use the `centred_shared_prior`
-# as a key to the `MeanField.mean` dictionary to print the estimated value of the shared centre.
-# """
-# print(f"Centre Mean Parameter Estimate = {mean_field.mean[centre_shared_prior]}")
-# print()
-#
-# """
-# If we want the parameter estimate of another parameter in the model, we can use the `model_list` that we composed
-# above to pass a parameter prior to the mean field dictionary.
-# """
-# print(
-# f"Normalization Gaussian Dataset 0 Mean = {mean_field.mean[model_list[0].gaussian.normalization]}"
-# )
-#
-# """
-# The mean-field mean dictionary contains the estimate value of every parameter.
-# """
-# print(f"All Parameter Estimates = {mean_field.mean}")
-# print()
-#
-# """
-# The mean-field also contains a `variance` dictionary, which has the same keys as the `mean` dictionary above.
-#
-# This is the easier way to estimate the error on every parameter, for example that of the shared centre.
-# """
-# print(f"Centre Variance = {mean_field.variance[centre_shared_prior]}")
-# print()
-#
-# """
-# The standard deviation (or error at one sigma confidence interval) is given by the square root of the variance.
-# """
-# print(f"Centre 1 Sigma = {np.sqrt(mean_field.variance[centre_shared_prior])}")
-# print()
-#
-# """
-# The mean field object also contains a dictionary of the s.d./variance**0.5.
-# """
-# print(f"Centre SD/sqrt(variance) = {mean_field.scale[centre_shared_prior]}")
-# print()
+"""
+Tutorial 5: Expectation Propagation
+===================================
+
+In the previous tutorials, we fitted graphical models to dataset comprising many noisy 1D Gaussians. These had a shared
+and global value of their `centre`, or assumed their centres were hierarchically drawn from a parent Gaussian
+distribution. This provides the basis of composing and fitting complex graphical models to large datasets.
+
+We concluded by discussing that there is a ceiling scaling these graphical models up to extremely large datasets. One
+would soon find that the parameter space is too complex to sample, and computational limits would ultimately cap how
+many datasets one could feasibly fit.
+
+This tutorial introduces expectation propagation (EP), the solution to this problem, which inspects a factor graph
+and partitions the model-fit into many simpler fits of sub-components of the graph to individual datasets. This
+overcomes the challenge of model complexity, and mitigates computational restrictions that may occur if one tries to
+fit every dataset simultaneously.
+
+This tutorial fits a global model with a shared parameter and does not use a hierarchical model. The optional tutorial
+`tutorial_optional_hierarchical_ep` shows an example fit of a hierarchical model with EP.
+
+__Contents__
+
+This tutorial is split into the following sections:
+
+- **Example Source Code (`af.ex`)**: The example objects used in this tutorial.
+- **Dataset**: Load the noisy 1D Gaussian datasets with a shared centre.
+- **Analysis**: Create Analysis objects for each dataset.
+- **Model**: Set up the model with a shared centre prior across all datasets.
+- **Analysis Factors**: Create Analysis Factors with individual searches for each dataset.
+- **Factor Graph**: Compose the factor graph for the EP framework.
+- **Expectation Propagation**: Explain the EP message passing algorithm.
+- **Cyclic Fitting**: Describe the iterative EP convergence process.
+- **Result**: Access the result of the EP fit.
+- **Output**: Describe the output directory structure and files generated by the EP fit.
+- **Results**: Use the MeanField object to infer parameter estimates and errors.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import numpy as np
+from os import path
+
+import autofit as af
+
+"""
+__Example Source Code (`af.ex`)__
+
+The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
+
+ - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
+ `visualize` functions.
+
+ - `Gaussian`: a model component representing a 1D Gaussian profile.
+
+ - `plot_profile_1d`: a function for plotting 1D profile datasets including their noise.
+
+These are functionally identical to the `Analysis`, `Gaussian` and `plot_profile_1d` objects and functions you have seen
+and used elsewhere throughout the workspace.
+
+__Dataset__
+
+For each dataset we now set up the correct path and load it.
+
+We first fit the 1D Gaussians which all share the same centre, thus not requiring a hierarchical model.
+
+An example for fitting the hierarchical model with EP is given at the end of this tutorial.
+"""
+total_datasets = 3
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(
+ path.join("dataset", "example_1d", "gaussian_x1__low_snr", "dataset_0")
+):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+dataset_name_list = []
+data_list = []
+noise_map_list = []
+
+for dataset_index in range(total_datasets):
+ dataset_name = f"dataset_{dataset_index}"
+
+ dataset_path = path.join(
+ "dataset", "example_1d", "gaussian_x1__low_snr", dataset_name
+ )
+
+ data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+ noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+ )
+
+ dataset_name_list.append(dataset_name)
+ data_list.append(data)
+ noise_map_list.append(noise_map)
+
+"""
+By plotting the Gaussians we can remind ourselves that determining their centres by eye is difficult.
+"""
+for dataset_name, data in zip(dataset_name_list, data_list):
+ af.ex.plot_profile_1d(
+ xvalues=np.arange(data.shape[0]),
+ profile_1d=data,
+ title=dataset_name,
+ ylabel="Data Values",
+ color="k",
+ )
+
+"""
+__Analysis__
+
+For each dataset we now create a corresponding `Analysis` class, like in the previous tutorial.
+"""
+analysis_list = []
+
+for data, noise_map in zip(data_list, noise_map_list):
+ analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+ analysis_list.append(analysis)
+
+"""
+__Model__
+
+We now compose the graphical model that we fit, using the `Model` and `Collection` objects you are now familiar with.
+
+We will assume all Gaussians share the same centre, therefore we set up a shared prior for `centre`.
+"""
+centre_shared_prior = af.GaussianPrior(mean=50.0, sigma=30.0)
+
+model_list = []
+
+for model_index in range(len(data_list)):
+ gaussian = af.Model(af.ex.Gaussian)
+
+ gaussian.centre = af.GaussianPrior(mean=50.0, sigma=30.0)
+ gaussian.normalization = af.TruncatedGaussianPrior(
+ mean=3.0, sigma=5.0, lower_limit=0.0
+ )
+ gaussian.sigma = af.TruncatedGaussianPrior(mean=10.0, sigma=10.0, lower_limit=0.0)
+
+ model = af.Collection(gaussian=gaussian)
+
+ model_list.append(model)
+
+"""
+__Analysis Factors__
+
+Now we have our `Analysis` classes and graphical model, we can compose our `AnalysisFactor`'s.
+
+However, unlike the previous tutorials, each `AnalysisFactor` is now assigned its own `search`. This is because the EP
+framework performs a model-fit to each node on the factor graph (e.g. each `AnalysisFactor`). Therefore, each node
+requires its own non-linear search, and in this tutorial we use `dynesty`. For complex graphs consisting of many
+nodes, one could easily use different searches for different nodes on the factor graph.
+
+Each `AnalysisFactor` is also given a `name`, corresponding to the name of the dataset it fits. These names are used
+to name the folders containing the results in the output directory.
+"""
+paths = af.DirectoryPaths(
+ name=path.join(
+ "howtofit", "chapter_graphical_models", "tutorial_5_expectation_propagation"
+ )
+)
+
+search = af.DynestyStatic(paths=paths, nlive=100, sample="rwalk")
+
+analysis_factor_list = []
+
+dataset_index = 0
+
+for model, analysis in zip(model_list, analysis_list):
+ dataset_name = f"dataset_{dataset_index}"
+ dataset_index += 1
+
+ analysis_factor = af.AnalysisFactor(
+ prior_model=model, analysis=analysis, optimiser=search, name=dataset_name
+ )
+
+ analysis_factor_list.append(analysis_factor)
+
+
+class LinearRegressionAnalysis(af.Analysis):
+ def log_likelihood_function(self, instance, xp=np):
+ return -1
+
+
+fwhm_list = [2 * np.sqrt(2 * np.log(2)) * model.gaussian.sigma for model in model_list]
+
+linear_regression_factor = af.AnalysisFactor(
+ prior_model=af.Collection(
+ m=af.GaussianPrior(mean=0.0, sigma=1.0),
+ c=af.GaussianPrior(mean=0.0, sigma=1.0),
+ *fwhm_list,
+ ),
+ analysis=LinearRegressionAnalysis(),
+ optimiser=search,
+ name="linear_regression",
+)
+
+"""
+__Factor Graph__
+
+We combine our `AnalysisFactors` into one, to compose the factor graph.
+"""
+factor_graph = af.FactorGraphModel(
+ *analysis_factor_list,
+ linear_regression_factor,
+)
+
+"""
+The factor graph model `info` attribute shows the model which we fit via expectaton propagation (note that we do
+not use `global_prior_model` below when performing the fit).
+"""
+print(factor_graph.global_prior_model.info)
+
+"""
+__Expectation Propagation__
+
+In the previous tutorials, we used the `global_prior_model` of the `factor_graph` to fit the global model. In this
+tutorial, we instead fit the `factor_graph` using the EP framework, which fits the graphical model composed in this
+tutorial as follows:
+
+1) Go to the first node on the factor graph (e.g. `analysis_factor_list[0]`) and fit its model to its dataset. This is
+simply a fit of the `Gaussian` model to the first 1D Gaussian dataset, the model-fit we are used to performing by now.
+
+2) Once the model-fit is complete, inspect the model for parameters that are shared with other nodes on the factor
+graph. In this example, the `centre` of the `Gaussian` fitted to the first dataset is global, and therefore connects
+to the other nodes on the factor graph (the `AnalysisFactor`'s) of the second and first `Gaussian` datasets.
+
+3) The EP framework now creates a 'message' that is to be passed to the connecting nodes on the factor graph. This
+message informs them of the results of the model-fit, so they can update their priors on the `Gaussian`'s centre
+accordingly and, more importantly, update their posterior inference and therefore estimate of the global centre.
+
+For example, the model fitted to the first Gaussian dataset includes the global centre. Therefore, after the model is
+fitted, the EP framework creates a 'message' informing the factor graph about its inference on that Gaussians's centre,
+thereby updating our overall inference on this shared parameter. This is termed 'message passing'.
+
+__Cyclic Fitting__
+
+After every `AnalysisFactor` has been fitted (e.g. after each fit to each of the 5 datasets in this example), we have a
+new estimate of the shared parameter `centre`. This updates our priors on the shared parameter `centre`, which needs
+to be reflected in each model-fit we perform on each `AnalysisFactor`.
+
+The EP framework therefore performs a second iteration of model-fits. It again cycles through each `AnalysisFactor`
+and refits the model, using updated priors on shared parameters like the `centre`. At the end of each fit, we again
+create messages that update our knowledge about other parameters on the graph.
+
+This process is repeated multiple times, until a convergence criteria is met whereby continued cycles are expected to
+produce the same estimate of the shared parameter `centre`.
+
+When we fit the factor graph a `name` is passed, which determines the folder all results of the factor graph are
+stored in.
+"""
+laplace = af.LaplaceOptimiser()
+
+factor_graph_result = factor_graph.optimise(
+ optimiser=laplace, paths=paths, ep_history=af.EPHistory(kl_tol=0.05), max_steps=5
+)
+
+"""
+__Result__
+
+An `info` attribute for the result of a factor graph fitted via EP does not exist yet, its on the to do list!
+
+The result can be seen in the `graph.result` file output to hard-disk.
+"""
+### print(factor_graph_result.info)##
+
+"""
+__Output__
+
+The results of the factor graph, using the EP framework and message passing, are contained in the folder
+`output/howtofit/chapter_graphical_models/tutorial_5_expectation_propagation`.
+
+The following folders and files are worth of note:
+
+ - `graph.info`: this provides an overall summary of the graphical model that is fitted, including every parameter,
+ how parameters are shared across `AnalysisFactor`'s and the priors associated to each individual parameter.
+
+ - The 3 folders titled `gaussian_x1_#__low_snr` correspond to the three `AnalysisFactor`'s and therefore signify
+ repeated non-linear searches that are performed to fit each dataset.
+
+ - Inside each of these folders are `optimization_#` folders, corresponding to each model-fit performed over cycles of
+ the EP fit. A careful inspection of the `model.info` files inside each folder reveals how the priors are updated
+ over each cycle, whereas the `model.results` file should indicate the improved estimate of model parameters over each
+ cycle.
+
+__Results__
+
+The `MeanField` object represent the posterior of the entire factor graph and is used to infer estimates of the
+values and error of each parameter in the graph.
+"""
+mean_field = factor_graph_result.updated_ep_mean_field.mean_field
+print(mean_field)
+print()
+
+"""
+The object has a `variables` property which lists every variable in the factor graph, which is essentially all of the
+free parameters on the graph.
+
+This includes the parameters specific to each data (E.g. each node on the graph) as well as the shared centre.
+"""
+print(mean_field.variables)
+print()
+
+# """
+# The variables above use the priors on each parameter as their key.
+#
+# Therefore to estimate mean-field quantities of the shared centre, we can simply use the `centre_shared_prior` defined
+# above.
+#
+# Each parameter estimate is given by the mean of its value in the `MeanField`. Below, we use the `centred_shared_prior`
+# as a key to the `MeanField.mean` dictionary to print the estimated value of the shared centre.
+# """
+# print(f"Centre Mean Parameter Estimate = {mean_field.mean[centre_shared_prior]}")
+# print()
+#
+# """
+# If we want the parameter estimate of another parameter in the model, we can use the `model_list` that we composed
+# above to pass a parameter prior to the mean field dictionary.
+# """
+# print(
+# f"Normalization Gaussian Dataset 0 Mean = {mean_field.mean[model_list[0].gaussian.normalization]}"
+# )
+#
+# """
+# The mean-field mean dictionary contains the estimate value of every parameter.
+# """
+# print(f"All Parameter Estimates = {mean_field.mean}")
+# print()
+#
+# """
+# The mean-field also contains a `variance` dictionary, which has the same keys as the `mean` dictionary above.
+#
+# This is the easier way to estimate the error on every parameter, for example that of the shared centre.
+# """
+# print(f"Centre Variance = {mean_field.variance[centre_shared_prior]}")
+# print()
+#
+# """
+# The standard deviation (or error at one sigma confidence interval) is given by the square root of the variance.
+# """
+# print(f"Centre 1 Sigma = {np.sqrt(mean_field.variance[centre_shared_prior])}")
+# print()
+#
+# """
+# The mean field object also contains a dictionary of the s.d./variance**0.5.
+# """
+# print(f"Centre SD/sqrt(variance) = {mean_field.scale[centre_shared_prior]}")
+# print()
diff --git a/scripts/howtofit/chapter_3_graphical_models/tutorial_optional_hierarchical_ep.py b/scripts/howtofit/chapter_3_graphical_models/tutorial_optional_hierarchical_ep.py
index 81cefd6d..1ddd8997 100644
--- a/scripts/howtofit/chapter_3_graphical_models/tutorial_optional_hierarchical_ep.py
+++ b/scripts/howtofit/chapter_3_graphical_models/tutorial_optional_hierarchical_ep.py
@@ -1,234 +1,230 @@
-"""
-Tutorial Optional: Hierarchical Expectation Propagation (EP)
-============================================================
-
-This optional tutorial gives an example of fitting a hierarchical model using EP.
-
-The API is a straightforward combination of tutorials 3 and 4.
-
-__Contents__
-
-This tutorial is split into the following sections:
-
-- **Example Source Code (`af.ex`)**: The example objects used in this tutorial.
-- **Dataset**: Load the hierarchical Gaussian datasets with variable centres.
-- **Analysis**: Create Analysis objects for each dataset.
-- **Model Individual Factors**: Set up individual Gaussian models with independent priors.
-- **Analysis Factors**: Compose Analysis Factors with individual searches for each dataset.
-- **Model**: Create a HierarchicalFactor with a parent Gaussian distribution.
-- **Factor Graph**: Compose the factor graph including the hierarchical factor.
-- **Model Fit**: Run the EP fit of the hierarchical model.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import numpy as np
-from os import path
-
-import autofit as af
-
-"""
-__Example Source Code (`af.ex`)__
-
-The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
-
- - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
- `visualize` functions.
-
- - `Gaussian`: a model component representing a 1D Gaussian profile.
-
- - `plot_profile_1d`: a function for plotting 1D profile datasets including their noise.
-
-These are functionally identical to the `Analysis`, `Gaussian` and `plot_profile_1d` objects and functions you have seen
-and used elsewhere throughout the workspace.
-
-__Dataset__
-
-For each dataset we now set up the correct path and load it.
-
-In this example, the three Gaussians have different centres, which are drawn from a parent Gaussian distribution
-whose mean and scatter we aim to estimate.
-"""
-total_datasets = 5
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(
- path.join("dataset", "example_1d", "gaussian_x1__hierarchical", "dataset_0")
-):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-dataset_name_list = []
-data_list = []
-noise_map_list = []
-
-for dataset_index in range(total_datasets):
- dataset_name = f"dataset_{dataset_index}"
-
- dataset_path = path.join(
- "dataset", "example_1d", "gaussian_x1__hierarchical", dataset_name
- )
-
- data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
- noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
- )
-
- dataset_name_list.append(dataset_name)
- data_list.append(data)
- noise_map_list.append(noise_map)
-
-"""
-By plotting the Gaussians we can just about make out that their centres are not all at pix 50, and are spreasd out
-around it (albeit its difficult to be sure, due to the low signal-to-noise of the data).
-"""
-for dataset_name, data in zip(dataset_name_list, data_list):
- af.ex.plot_profile_1d(
- xvalues=np.arange(data.shape[0]),
- profile_1d=data,
- title=dataset_name,
- ylabel="Data Values",
- color="k",
- )
-
-"""
-__Analysis__
-
-For each dataset we now create a corresponding `Analysis` class, like in the previous tutorial.
-"""
-analysis_list = []
-
-for data, noise_map in zip(data_list, noise_map_list):
- analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
- analysis_list.append(analysis)
-
-
-"""
-__Model Individual Factors__
-
-We first set up a model for each `Gaussian` which is individually fitted to each 1D dataset, which forms the
-factors on the factor graph we compose.
-
-This uses a nearly identical for loop to the previous tutorial, however a shared `centre` is no longer used and each
-`Gaussian` is given its own prior for the `centre`. We will see next how this `centre` is used to construct the
-hierachical model.
-"""
-
-model_list = []
-
-for model_index in range(len(data_list)):
- gaussian = af.Model(af.ex.Gaussian)
-
- # gaussian.centre = af.UniformPrior(lower_limit=0.0, upper_limit=1e2)
- # gaussian.normalization = af.UniformPrior(lower_limit=0.0, upper_limit=1e2)
- # gaussian.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=25.0)
-
- gaussian.centre = af.TruncatedGaussianPrior(
- mean=50.0, sigma=20.0, lower_limit=0.0, upper_limit=100.0
- )
- gaussian.normalization = af.TruncatedGaussianPrior(
- mean=3.0, sigma=5.0, lower_limit=0.0
- )
- gaussian.sigma = af.TruncatedGaussianPrior(mean=10.0, sigma=10.0, lower_limit=0.0)
-
- model_list.append(gaussian)
-
-"""
-__Analysis Factors__
-
-Now we have our `Analysis` classes and model components, we can compose our `AnalysisFactor`'s.
-
-The hierarchical model fit uses EP, therefore we again supply each `AnalysisFactor` its own `search` and `name`.
-"""
-dynesty = af.DynestyStatic(nlive=100, sample="rwalk")
-
-analysis_factor_list = []
-
-dataset_index = 0
-
-for model, analysis in zip(model_list, analysis_list):
- dataset_name = f"dataset_{dataset_index}"
- dataset_index += 1
-
- analysis_factor = af.AnalysisFactor(
- prior_model=model, analysis=analysis, optimiser=dynesty, name=dataset_name
- )
-
- analysis_factor_list.append(analysis_factor)
-
-
-"""
-__Model__
-
-We now compose the hierarchical model that we fit, using the individual Gaussian model components we created above.
-
-We first create a `HierarchicalFactor`, which represents the parent Gaussian distribution from which we will assume
-that the `centre` of each individual `Gaussian` dataset is drawn.
-
-For this parent `Gaussian`, we have to place priors on its `mean` and `sigma`, given that they are parameters in our
-model we are ultimately fitting for.
-"""
-hierarchical_factor = af.HierarchicalFactor(
- af.GaussianPrior,
- mean=af.TruncatedGaussianPrior(
- mean=50.0, sigma=10, lower_limit=0.0, upper_limit=100.0
- ),
- sigma=af.TruncatedGaussianPrior(
- mean=10.0, sigma=5.0, lower_limit=0.0, upper_limit=100.0
- ),
-)
-
-"""
-We now add each of the individual model `Gaussian`'s `centre` parameters to the `hierarchical_factor`.
-
-This composes the hierarchical model whereby the individual `centre` of every `Gaussian` in our dataset is now assumed
-to be drawn from a shared parent distribution. It is the `mean` and `sigma` of this distribution we are hoping to
-estimate.
-"""
-for model in model_list:
- hierarchical_factor.add_drawn_variable(model.centre)
-
-"""
-__Factor Graph__
-
-We now create the factor graph for this model, using the list of `AnalysisFactor`'s and the hierarchical factor.
-
-Note that in previous tutorials, when we created the `FactorGraphModel` we only passed the list of `AnalysisFactor`'s,
-which contained the necessary information on the model create the factor graph that was fitted. The `AnalysisFactor`'s
-were created before we composed the `HierachicalFactor`, which is why we need to pass it separate when composing the
-factor graph.
-"""
-factor_graph = af.FactorGraphModel(*analysis_factor_list, hierarchical_factor)
-
-"""
-__Model Fit__
-
-
-"""
-laplace = af.LaplaceOptimiser()
-
-# ep_result = factor_graph.optimise(
-# laplace,
-# paths=af.DirectoryPaths(
-# name=path.join(
-# "howtofit", "chapter_graphical_models", "tutorial_4_hierarchical"
-# )
-# ),
-# ep_history=af.EPHistory(kl_tol=1.0),
-# max_steps=5,
-# )
+"""
+Tutorial Optional: Hierarchical Expectation Propagation (EP)
+============================================================
+
+This optional tutorial gives an example of fitting a hierarchical model using EP.
+
+The API is a straightforward combination of tutorials 3 and 4.
+
+__Contents__
+
+This tutorial is split into the following sections:
+
+- **Example Source Code (`af.ex`)**: The example objects used in this tutorial.
+- **Dataset**: Load the hierarchical Gaussian datasets with variable centres.
+- **Analysis**: Create Analysis objects for each dataset.
+- **Model Individual Factors**: Set up individual Gaussian models with independent priors.
+- **Analysis Factors**: Compose Analysis Factors with individual searches for each dataset.
+- **Model**: Create a HierarchicalFactor with a parent Gaussian distribution.
+- **Factor Graph**: Compose the factor graph including the hierarchical factor.
+- **Model Fit**: Run the EP fit of the hierarchical model.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import numpy as np
+from os import path
+
+import autofit as af
+
+"""
+__Example Source Code (`af.ex`)__
+
+The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
+
+ - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
+ `visualize` functions.
+
+ - `Gaussian`: a model component representing a 1D Gaussian profile.
+
+ - `plot_profile_1d`: a function for plotting 1D profile datasets including their noise.
+
+These are functionally identical to the `Analysis`, `Gaussian` and `plot_profile_1d` objects and functions you have seen
+and used elsewhere throughout the workspace.
+
+__Dataset__
+
+For each dataset we now set up the correct path and load it.
+
+In this example, the three Gaussians have different centres, which are drawn from a parent Gaussian distribution
+whose mean and scatter we aim to estimate.
+"""
+total_datasets = 5
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(
+ path.join("dataset", "example_1d", "gaussian_x1__hierarchical", "dataset_0")
+):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+dataset_name_list = []
+data_list = []
+noise_map_list = []
+
+for dataset_index in range(total_datasets):
+ dataset_name = f"dataset_{dataset_index}"
+
+ dataset_path = path.join(
+ "dataset", "example_1d", "gaussian_x1__hierarchical", dataset_name
+ )
+
+ data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+ noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+ )
+
+ dataset_name_list.append(dataset_name)
+ data_list.append(data)
+ noise_map_list.append(noise_map)
+
+"""
+By plotting the Gaussians we can just about make out that their centres are not all at pix 50, and are spreasd out
+around it (albeit its difficult to be sure, due to the low signal-to-noise of the data).
+"""
+for dataset_name, data in zip(dataset_name_list, data_list):
+ af.ex.plot_profile_1d(
+ xvalues=np.arange(data.shape[0]),
+ profile_1d=data,
+ title=dataset_name,
+ ylabel="Data Values",
+ color="k",
+ )
+
+"""
+__Analysis__
+
+For each dataset we now create a corresponding `Analysis` class, like in the previous tutorial.
+"""
+analysis_list = []
+
+for data, noise_map in zip(data_list, noise_map_list):
+ analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+ analysis_list.append(analysis)
+
+
+"""
+__Model Individual Factors__
+
+We first set up a model for each `Gaussian` which is individually fitted to each 1D dataset, which forms the
+factors on the factor graph we compose.
+
+This uses a nearly identical for loop to the previous tutorial, however a shared `centre` is no longer used and each
+`Gaussian` is given its own prior for the `centre`. We will see next how this `centre` is used to construct the
+hierachical model.
+"""
+
+model_list = []
+
+for model_index in range(len(data_list)):
+ gaussian = af.Model(af.ex.Gaussian)
+
+ # gaussian.centre = af.UniformPrior(lower_limit=0.0, upper_limit=1e2)
+ # gaussian.normalization = af.UniformPrior(lower_limit=0.0, upper_limit=1e2)
+ # gaussian.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=25.0)
+
+ gaussian.centre = af.TruncatedGaussianPrior(
+ mean=50.0, sigma=20.0, lower_limit=0.0, upper_limit=100.0
+ )
+ gaussian.normalization = af.TruncatedGaussianPrior(
+ mean=3.0, sigma=5.0, lower_limit=0.0
+ )
+ gaussian.sigma = af.TruncatedGaussianPrior(mean=10.0, sigma=10.0, lower_limit=0.0)
+
+ model_list.append(gaussian)
+
+"""
+__Analysis Factors__
+
+Now we have our `Analysis` classes and model components, we can compose our `AnalysisFactor`'s.
+
+The hierarchical model fit uses EP, therefore we again supply each `AnalysisFactor` its own `search` and `name`.
+"""
+dynesty = af.DynestyStatic(nlive=100, sample="rwalk")
+
+analysis_factor_list = []
+
+dataset_index = 0
+
+for model, analysis in zip(model_list, analysis_list):
+ dataset_name = f"dataset_{dataset_index}"
+ dataset_index += 1
+
+ analysis_factor = af.AnalysisFactor(
+ prior_model=model, analysis=analysis, optimiser=dynesty, name=dataset_name
+ )
+
+ analysis_factor_list.append(analysis_factor)
+
+
+"""
+__Model__
+
+We now compose the hierarchical model that we fit, using the individual Gaussian model components we created above.
+
+We first create a `HierarchicalFactor`, which represents the parent Gaussian distribution from which we will assume
+that the `centre` of each individual `Gaussian` dataset is drawn.
+
+For this parent `Gaussian`, we have to place priors on its `mean` and `sigma`, given that they are parameters in our
+model we are ultimately fitting for.
+"""
+hierarchical_factor = af.HierarchicalFactor(
+ af.GaussianPrior,
+ mean=af.TruncatedGaussianPrior(
+ mean=50.0, sigma=10, lower_limit=0.0, upper_limit=100.0
+ ),
+ sigma=af.TruncatedGaussianPrior(
+ mean=10.0, sigma=5.0, lower_limit=0.0, upper_limit=100.0
+ ),
+)
+
+"""
+We now add each of the individual model `Gaussian`'s `centre` parameters to the `hierarchical_factor`.
+
+This composes the hierarchical model whereby the individual `centre` of every `Gaussian` in our dataset is now assumed
+to be drawn from a shared parent distribution. It is the `mean` and `sigma` of this distribution we are hoping to
+estimate.
+"""
+for model in model_list:
+ hierarchical_factor.add_drawn_variable(model.centre)
+
+"""
+__Factor Graph__
+
+We now create the factor graph for this model, using the list of `AnalysisFactor`'s and the hierarchical factor.
+
+Note that in previous tutorials, when we created the `FactorGraphModel` we only passed the list of `AnalysisFactor`'s,
+which contained the necessary information on the model create the factor graph that was fitted. The `AnalysisFactor`'s
+were created before we composed the `HierachicalFactor`, which is why we need to pass it separate when composing the
+factor graph.
+"""
+factor_graph = af.FactorGraphModel(*analysis_factor_list, hierarchical_factor)
+
+"""
+__Model Fit__
+
+
+"""
+laplace = af.LaplaceOptimiser()
+
+# ep_result = factor_graph.optimise(
+# laplace,
+# paths=af.DirectoryPaths(
+# name=path.join(
+# "howtofit", "chapter_graphical_models", "tutorial_4_hierarchical"
+# )
+# ),
+# ep_history=af.EPHistory(kl_tol=1.0),
+# max_steps=5,
+# )
diff --git a/scripts/howtofit/chapter_3_graphical_models/tutorial_optional_hierarchical_individual.py b/scripts/howtofit/chapter_3_graphical_models/tutorial_optional_hierarchical_individual.py
index 08ae17b1..e8961eee 100644
--- a/scripts/howtofit/chapter_3_graphical_models/tutorial_optional_hierarchical_individual.py
+++ b/scripts/howtofit/chapter_3_graphical_models/tutorial_optional_hierarchical_individual.py
@@ -1,344 +1,340 @@
-"""
-Tutorial Optional: Hierarchical Individual
-==========================================
-
-In tutorial 4, we fit a hierarchical model using a graphical model, whereby all datasets are fitted simultaneously
-and the hierarchical parameters are fitted for simultaneously with the model parameters of each 1D Gaussian in each
-dataset.
-
-This script illustrates how the hierarchical parameters can be estimated using a simpler approach, which fits
-each dataset one-by-one and estimates the hierarchical parameters afterwards by fitting the inferred `centres`
-with a Gaussian distribution.
-
-__Contents__
-
-This tutorial is split into the following sections:
-
-- **Example Source Code (`af.ex`)**: The example objects used in this tutorial.
-- **Dataset**: Load the hierarchical Gaussian datasets with variable centres.
-- **Analysis**: Create Analysis objects for each dataset.
-- **Model**: Define a simple Gaussian model with uniform priors.
-- **Model Fits (one-by-one)**: Fit each dataset individually using separate non-linear searches.
-- **Results**: Analyze and plot the results of each individual fit.
-- **Overall Gaussian Parent Distribution**: Fit a parent Gaussian distribution to the inferred centres.
-- **Model**: Set up a ParentGaussian model for the parent distribution fitting.
-- **Analysis + Search**: Create the analysis and search for the parent distribution fit.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import numpy as np
-from os import path
-
-import autofit as af
-
-"""
-__Example Source Code (`af.ex`)__
-
-The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
-
- - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
- `visualize` functions.
-
- - `Gaussian`: a model component representing a 1D Gaussian profile.
-
- - `plot_profile_1d`: a function for plotting 1D profile datasets including their noise.
-
-These are functionally identical to the `Analysis`, `Gaussian` and `plot_profile_1d` objects and functions you
-have seen and used elsewhere throughout the workspace.
-
-__Dataset__
-
-For each dataset we now set up the correct path and load it.
-
-We are loading a new Gaussian dataset, where the Gaussians have different centres which were drawn from a parent
-Gaussian distribution with a mean centre value of 50.0 and sigma of 10.0.
-"""
-total_datasets = 5
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(
- path.join("dataset", "example_1d", "gaussian_x1__hierarchical", "dataset_0")
-):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-dataset_name_list = []
-data_list = []
-noise_map_list = []
-
-for dataset_index in range(total_datasets):
- dataset_name = f"dataset_{dataset_index}"
-
- dataset_path = path.join(
- "dataset", "example_1d", "gaussian_x1__hierarchical", dataset_name
- )
-
- data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
- noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
- )
-
- dataset_name_list.append(dataset_name)
- data_list.append(data)
- noise_map_list.append(noise_map)
-
-"""
-By plotting the Gaussians we can just about make out that their centres are not all at pix 50, and are spread out
-around it (albeit its difficult to be sure, due to the low signal-to-noise of the data).
-"""
-for dataset_name, data in zip(dataset_name_list, data_list):
- af.ex.plot_profile_1d(
- xvalues=np.arange(data.shape[0]),
- profile_1d=data,
- title=dataset_name,
- ylabel="Data Values",
- color="k",
- )
-
-"""
-__Analysis__
-
-For each dataset we now create a corresponding `Analysis` class, like in the previous tutorial.
-"""
-analysis_list = []
-
-for data, noise_map in zip(data_list, noise_map_list):
- analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
- analysis_list.append(analysis)
-
-
-"""
-__Model__
-
-The model we fit to each dataset, which is a simple 1D Gaussian with all 3 parameters free.
-"""
-gaussian = af.Model(af.ex.Gaussian)
-
-gaussian.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-gaussian.normalization = af.UniformPrior(lower_limit=0.0, upper_limit=10.0)
-gaussian.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=50.0)
-
-model = af.Collection(gaussian=gaussian)
-
-
-"""
-__Model Fits (one-by-one)__
-
-For every dataset we now create an `Analysis` class using it and use `Dynesty` to fit it with a `Gaussian`.
-
-The `Result` is stored in the list `results`.
-"""
-result_list = []
-
-for dataset_name, analysis in zip(dataset_name_list, analysis_list):
- """
- Create the `DynestyStatic` non-linear search and use it to fit the data.
- """
- dynesty = af.DynestyStatic(
- name="tutorial_optional_hierarchical_individual",
- unique_tag=dataset_name,
- nlive=200,
- dlogz=1e-4,
- sample="rwalk",
- walks=10,
- )
-
- result_list.append(dynesty.fit(model=model, analysis=analysis))
-
-"""
-__Results__
-
-Checkout the output folder, you should see three new sets of results corresponding to our 3 `Gaussian` datasets.
-
-The `result_list` allows us to plot the median PDF value and 3.0 confidence intervals of the `centre` estimate from
-the model-fit to each dataset.
-"""
-samples_list = [result.samples for result in result_list]
-
-mp_instances = [samps.median_pdf() for samps in samples_list]
-ue3_instances = [samp.errors_at_upper_sigma(sigma=3.0) for samp in samples_list]
-le3_instances = [samp.errors_at_lower_sigma(sigma=3.0) for samp in samples_list]
-
-mp_centres = [instance.gaussian.centre for instance in mp_instances]
-ue3_centres = [instance.gaussian.centre for instance in ue3_instances]
-le3_centres = [instance.gaussian.centre for instance in le3_instances]
-
-print(f"Median PDF inferred centre values")
-print(mp_centres)
-print()
-
-"""
-__Overall Gaussian Parent Distribution__
-
-Fit the inferred `centre`'s from the fits performed above with a Gaussian distribution, in order to
-estimate the mean and scatter of the Gaussian from which the centres were drawn.
-
-We first extract the inferred median PDF centre values and their 1 sigma errors below, which will be the inputs
-to our fit for the parent Gaussian.
-"""
-ue1_instances = [samp.values_at_upper_sigma(sigma=1.0) for samp in samples_list]
-le1_instances = [samp.values_at_lower_sigma(sigma=1.0) for samp in samples_list]
-
-ue1_centres = [instance.gaussian.centre for instance in ue1_instances]
-le1_centres = [instance.gaussian.centre for instance in le1_instances]
-
-error_list = [ue1 - le1 for ue1, le1 in zip(ue1_centres, le1_centres)]
-
-"""
-The `Analysis` class below fits a Gaussian distribution to the inferred `centre` values from each of the fits above,
-where the inferred error values are used as the errors.
-"""
-
-
-class Analysis(af.Analysis):
- def __init__(self, data: np.ndarray, errors: np.ndarray):
- super().__init__()
-
- self.data = np.array(data)
- self.errors = np.array(errors)
-
- def log_likelihood_function(self, instance: af.ModelInstance) -> float:
- """
- Fits a set of 1D data points with a 1D Gaussian distribution, in order to determine from what Gaussian
- distribution the analysis classes `data` were drawn.
-
- In this example, this function determines from what parent Gaussian disrtribution the inferred centres
- of each 1D Gaussian were drawn.
- """
- log_likelihood_term_1 = np.sum(
- -np.divide(
- (self.data - instance.median) ** 2,
- 2 * (instance.scatter**2 + self.errors**2),
- )
- )
- log_likelihood_term_2 = -np.sum(
- 0.5 * np.log(instance.scatter**2 + self.errors**2)
- )
-
- return log_likelihood_term_1 + log_likelihood_term_2
-
-
-"""
-The `ParentGaussian` class is the model-component which used to fit the parent Gaussian to the inferred `centre` values.
-"""
-
-
-class ParentGaussian:
- def __init__(self, median: float = 0.0, scatter: float = 0.01):
- """
- A model component which represents a parent Gaussian distribution, which can be fitted to a 1D set of
- measurments with errors in order to determine the probabilty they were drawn from this Gaussian.
-
- Parameters
- ----------
- median
- The median value of the parent Gaussian distribution.
- scatter
- The scatter (E.g. the sigma value) of the Gaussian.
- """
-
- self.median = median
- self.scatter = scatter
-
- def probability_from_values(self, values: np.ndarray) -> float:
- """
- For a set of 1D values, determine the probability that they were random drawn from this parent Gaussian
- based on its `median` and `scatter` attributes.
-
- Parameters
- ----------
- values
- A set of 1D values from which we will determine the probability they were drawn from the parent Gaussian.
- """
- values = np.sort(np.array(values))
- transformed_values = np.subtract(values, self.median)
-
- return np.multiply(
- np.divide(1, self.scatter * np.sqrt(2.0 * np.pi)),
- np.exp(-0.5 * np.square(np.divide(transformed_values, self.scatter))),
- )
-
-
-"""
-__Model__
-
-The `ParentGaussian` is the model component we fit in order to determine the probability the inferred centres were
-drawn from the distribution.
-
-This will be fitted via a non-linear search and therefore is created as a model component using `af.Model()` as per
-usual in **PyAutoFit**.
-"""
-model = af.Model(ParentGaussian)
-
-model.median = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.scatter = af.UniformPrior(lower_limit=0.0, upper_limit=50.0)
-
-"""
-__Analysis + Search__
-
-We now create the Analysis class above which fits a parent 1D gaussian and create a dynesty search in order to fit
-it to the 1D inferred list of `centres`.
-"""
-analysis = Analysis(data=mp_centres, errors=error_list)
-search = af.DynestyStatic(nlive=100)
-
-result = search.fit(model=model, analysis=analysis)
-
-"""
-The results of this fit tell us the most probably values for the `median` and `scatter` of the 1D parent Gaussian fit.
-"""
-samples = result.samples
-
-median = samples.median_pdf().median
-
-u1_error = samples.values_at_upper_sigma(sigma=1.0).median
-l1_error = samples.values_at_lower_sigma(sigma=1.0).median
-
-u3_error = samples.values_at_upper_sigma(sigma=3.0).median
-l3_error = samples.values_at_lower_sigma(sigma=3.0).median
-
-print(
- f"Inferred value of the hierarchical median via simple fit to {total_datasets} datasets: \n "
-)
-print(f"{median} ({l1_error} {u1_error}) [1.0 sigma confidence intervals]")
-print(f"{median} ({l3_error} {u3_error}) [3.0 sigma confidence intervals]")
-print()
-
-scatter = samples.median_pdf().scatter
-
-u1_error = samples.values_at_upper_sigma(sigma=1.0).scatter
-l1_error = samples.values_at_lower_sigma(sigma=1.0).scatter
-
-u3_error = samples.values_at_upper_sigma(sigma=3.0).scatter
-l3_error = samples.values_at_lower_sigma(sigma=3.0).scatter
-
-print(
- f"Inferred value of the hierarchical scatter via simple fit to {total_datasets} datasets: \n "
-)
-print(f"{scatter} ({l1_error} {u1_error}) [1.0 sigma confidence intervals]")
-print(f"{scatter} ({l3_error} {u3_error}) [3.0 sigma confidence intervals]")
-print()
-
-"""
-We can compare these values to those inferred in `tutorial_4_hierarchical_model`, which fits all datasets and the
-hierarchical values of the parent Gaussian simultaneously.,
-
-The errors for the fit performed in this tutorial are much larger. This is because of how in a graphical model
-the "datasets talk to one another", which is described fully in that tutorials subsection "Benefits of Graphical Model".
-"""
+"""
+Tutorial Optional: Hierarchical Individual
+==========================================
+
+In tutorial 4, we fit a hierarchical model using a graphical model, whereby all datasets are fitted simultaneously
+and the hierarchical parameters are fitted for simultaneously with the model parameters of each 1D Gaussian in each
+dataset.
+
+This script illustrates how the hierarchical parameters can be estimated using a simpler approach, which fits
+each dataset one-by-one and estimates the hierarchical parameters afterwards by fitting the inferred `centres`
+with a Gaussian distribution.
+
+__Contents__
+
+This tutorial is split into the following sections:
+
+- **Example Source Code (`af.ex`)**: The example objects used in this tutorial.
+- **Dataset**: Load the hierarchical Gaussian datasets with variable centres.
+- **Analysis**: Create Analysis objects for each dataset.
+- **Model**: Define a simple Gaussian model with uniform priors.
+- **Model Fits (one-by-one)**: Fit each dataset individually using separate non-linear searches.
+- **Results**: Analyze and plot the results of each individual fit.
+- **Overall Gaussian Parent Distribution**: Fit a parent Gaussian distribution to the inferred centres.
+- **Model**: Set up a ParentGaussian model for the parent distribution fitting.
+- **Analysis + Search**: Create the analysis and search for the parent distribution fit.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import numpy as np
+from os import path
+
+import autofit as af
+
+"""
+__Example Source Code (`af.ex`)__
+
+The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
+
+ - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
+ `visualize` functions.
+
+ - `Gaussian`: a model component representing a 1D Gaussian profile.
+
+ - `plot_profile_1d`: a function for plotting 1D profile datasets including their noise.
+
+These are functionally identical to the `Analysis`, `Gaussian` and `plot_profile_1d` objects and functions you
+have seen and used elsewhere throughout the workspace.
+
+__Dataset__
+
+For each dataset we now set up the correct path and load it.
+
+We are loading a new Gaussian dataset, where the Gaussians have different centres which were drawn from a parent
+Gaussian distribution with a mean centre value of 50.0 and sigma of 10.0.
+"""
+total_datasets = 5
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(
+ path.join("dataset", "example_1d", "gaussian_x1__hierarchical", "dataset_0")
+):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+dataset_name_list = []
+data_list = []
+noise_map_list = []
+
+for dataset_index in range(total_datasets):
+ dataset_name = f"dataset_{dataset_index}"
+
+ dataset_path = path.join(
+ "dataset", "example_1d", "gaussian_x1__hierarchical", dataset_name
+ )
+
+ data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+ noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+ )
+
+ dataset_name_list.append(dataset_name)
+ data_list.append(data)
+ noise_map_list.append(noise_map)
+
+"""
+By plotting the Gaussians we can just about make out that their centres are not all at pix 50, and are spread out
+around it (albeit its difficult to be sure, due to the low signal-to-noise of the data).
+"""
+for dataset_name, data in zip(dataset_name_list, data_list):
+ af.ex.plot_profile_1d(
+ xvalues=np.arange(data.shape[0]),
+ profile_1d=data,
+ title=dataset_name,
+ ylabel="Data Values",
+ color="k",
+ )
+
+"""
+__Analysis__
+
+For each dataset we now create a corresponding `Analysis` class, like in the previous tutorial.
+"""
+analysis_list = []
+
+for data, noise_map in zip(data_list, noise_map_list):
+ analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+ analysis_list.append(analysis)
+
+
+"""
+__Model__
+
+The model we fit to each dataset, which is a simple 1D Gaussian with all 3 parameters free.
+"""
+gaussian = af.Model(af.ex.Gaussian)
+
+gaussian.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+gaussian.normalization = af.UniformPrior(lower_limit=0.0, upper_limit=10.0)
+gaussian.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=50.0)
+
+model = af.Collection(gaussian=gaussian)
+
+
+"""
+__Model Fits (one-by-one)__
+
+For every dataset we now create an `Analysis` class using it and use `Dynesty` to fit it with a `Gaussian`.
+
+The `Result` is stored in the list `results`.
+"""
+result_list = []
+
+for dataset_name, analysis in zip(dataset_name_list, analysis_list):
+ """
+ Create the `DynestyStatic` non-linear search and use it to fit the data.
+ """
+ dynesty = af.DynestyStatic(
+ name="tutorial_optional_hierarchical_individual",
+ unique_tag=dataset_name,
+ nlive=200,
+ dlogz=1e-4,
+ sample="rwalk",
+ walks=10,
+ )
+
+ result_list.append(dynesty.fit(model=model, analysis=analysis))
+
+"""
+__Results__
+
+Checkout the output folder, you should see three new sets of results corresponding to our 3 `Gaussian` datasets.
+
+The `result_list` allows us to plot the median PDF value and 3.0 confidence intervals of the `centre` estimate from
+the model-fit to each dataset.
+"""
+samples_list = [result.samples for result in result_list]
+
+mp_instances = [samps.median_pdf() for samps in samples_list]
+ue3_instances = [samp.errors_at_upper_sigma(sigma=3.0) for samp in samples_list]
+le3_instances = [samp.errors_at_lower_sigma(sigma=3.0) for samp in samples_list]
+
+mp_centres = [instance.gaussian.centre for instance in mp_instances]
+ue3_centres = [instance.gaussian.centre for instance in ue3_instances]
+le3_centres = [instance.gaussian.centre for instance in le3_instances]
+
+print(f"Median PDF inferred centre values")
+print(mp_centres)
+print()
+
+"""
+__Overall Gaussian Parent Distribution__
+
+Fit the inferred `centre`'s from the fits performed above with a Gaussian distribution, in order to
+estimate the mean and scatter of the Gaussian from which the centres were drawn.
+
+We first extract the inferred median PDF centre values and their 1 sigma errors below, which will be the inputs
+to our fit for the parent Gaussian.
+"""
+ue1_instances = [samp.values_at_upper_sigma(sigma=1.0) for samp in samples_list]
+le1_instances = [samp.values_at_lower_sigma(sigma=1.0) for samp in samples_list]
+
+ue1_centres = [instance.gaussian.centre for instance in ue1_instances]
+le1_centres = [instance.gaussian.centre for instance in le1_instances]
+
+error_list = [ue1 - le1 for ue1, le1 in zip(ue1_centres, le1_centres)]
+
+"""
+The `Analysis` class below fits a Gaussian distribution to the inferred `centre` values from each of the fits above,
+where the inferred error values are used as the errors.
+"""
+
+
+class Analysis(af.Analysis):
+ def __init__(self, data: np.ndarray, errors: np.ndarray):
+ super().__init__()
+
+ self.data = np.array(data)
+ self.errors = np.array(errors)
+
+ def log_likelihood_function(self, instance: af.ModelInstance) -> float:
+ """
+ Fits a set of 1D data points with a 1D Gaussian distribution, in order to determine from what Gaussian
+ distribution the analysis classes `data` were drawn.
+
+ In this example, this function determines from what parent Gaussian disrtribution the inferred centres
+ of each 1D Gaussian were drawn.
+ """
+ log_likelihood_term_1 = np.sum(
+ -np.divide(
+ (self.data - instance.median) ** 2,
+ 2 * (instance.scatter**2 + self.errors**2),
+ )
+ )
+ log_likelihood_term_2 = -np.sum(
+ 0.5 * np.log(instance.scatter**2 + self.errors**2)
+ )
+
+ return log_likelihood_term_1 + log_likelihood_term_2
+
+
+"""
+The `ParentGaussian` class is the model-component which used to fit the parent Gaussian to the inferred `centre` values.
+"""
+
+
+class ParentGaussian:
+ def __init__(self, median: float = 0.0, scatter: float = 0.01):
+ """
+ A model component which represents a parent Gaussian distribution, which can be fitted to a 1D set of
+ measurments with errors in order to determine the probabilty they were drawn from this Gaussian.
+
+ Parameters
+ ----------
+ median
+ The median value of the parent Gaussian distribution.
+ scatter
+ The scatter (E.g. the sigma value) of the Gaussian.
+ """
+
+ self.median = median
+ self.scatter = scatter
+
+ def probability_from_values(self, values: np.ndarray) -> float:
+ """
+ For a set of 1D values, determine the probability that they were random drawn from this parent Gaussian
+ based on its `median` and `scatter` attributes.
+
+ Parameters
+ ----------
+ values
+ A set of 1D values from which we will determine the probability they were drawn from the parent Gaussian.
+ """
+ values = np.sort(np.array(values))
+ transformed_values = np.subtract(values, self.median)
+
+ return np.multiply(
+ np.divide(1, self.scatter * np.sqrt(2.0 * np.pi)),
+ np.exp(-0.5 * np.square(np.divide(transformed_values, self.scatter))),
+ )
+
+
+"""
+__Model__
+
+The `ParentGaussian` is the model component we fit in order to determine the probability the inferred centres were
+drawn from the distribution.
+
+This will be fitted via a non-linear search and therefore is created as a model component using `af.Model()` as per
+usual in **PyAutoFit**.
+"""
+model = af.Model(ParentGaussian)
+
+model.median = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.scatter = af.UniformPrior(lower_limit=0.0, upper_limit=50.0)
+
+"""
+__Analysis + Search__
+
+We now create the Analysis class above which fits a parent 1D gaussian and create a dynesty search in order to fit
+it to the 1D inferred list of `centres`.
+"""
+analysis = Analysis(data=mp_centres, errors=error_list)
+search = af.DynestyStatic(nlive=100)
+
+result = search.fit(model=model, analysis=analysis)
+
+"""
+The results of this fit tell us the most probably values for the `median` and `scatter` of the 1D parent Gaussian fit.
+"""
+samples = result.samples
+
+median = samples.median_pdf().median
+
+u1_error = samples.values_at_upper_sigma(sigma=1.0).median
+l1_error = samples.values_at_lower_sigma(sigma=1.0).median
+
+u3_error = samples.values_at_upper_sigma(sigma=3.0).median
+l3_error = samples.values_at_lower_sigma(sigma=3.0).median
+
+print(
+ f"Inferred value of the hierarchical median via simple fit to {total_datasets} datasets: \n "
+)
+print(f"{median} ({l1_error} {u1_error}) [1.0 sigma confidence intervals]")
+print(f"{median} ({l3_error} {u3_error}) [3.0 sigma confidence intervals]")
+print()
+
+scatter = samples.median_pdf().scatter
+
+u1_error = samples.values_at_upper_sigma(sigma=1.0).scatter
+l1_error = samples.values_at_lower_sigma(sigma=1.0).scatter
+
+u3_error = samples.values_at_upper_sigma(sigma=3.0).scatter
+l3_error = samples.values_at_lower_sigma(sigma=3.0).scatter
+
+print(
+ f"Inferred value of the hierarchical scatter via simple fit to {total_datasets} datasets: \n "
+)
+print(f"{scatter} ({l1_error} {u1_error}) [1.0 sigma confidence intervals]")
+print(f"{scatter} ({l3_error} {u3_error}) [3.0 sigma confidence intervals]")
+print()
+
+"""
+We can compare these values to those inferred in `tutorial_4_hierarchical_model`, which fits all datasets and the
+hierarchical values of the parent Gaussian simultaneously.,
+
+The errors for the fit performed in this tutorial are much larger. This is because of how in a graphical model
+the "datasets talk to one another", which is described fully in that tutorials subsection "Benefits of Graphical Model".
+"""
diff --git a/scripts/overview/overview_1_the_basics.py b/scripts/overview/overview_1_the_basics.py
index df26b95b..33015739 100644
--- a/scripts/overview/overview_1_the_basics.py
+++ b/scripts/overview/overview_1_the_basics.py
@@ -1,883 +1,879 @@
-"""
-Overview: The Basics
---------------------
-
-**PyAutoFit** is a Python based probabilistic programming language for model fitting and Bayesian inference
-of large datasets.
-
-The basic **PyAutoFit** API allows us a user to quickly compose a probabilistic model and fit it to data via a
-log likelihood function, using a range of non-linear search algorithms (e.g. MCMC, nested sampling).
-
-This overview gives a run through of:
-
- - **Models**: Use Python classes to compose the model which is fitted to data.
- - **Instances**: Create instances of the model via its Python class.
- - **Analysis**: Define an ``Analysis`` class which includes the log likelihood function that fits the model to the data.
- - **Searches**: Choose an MCMC, nested sampling or maximum likelihood estimator non-linear search algorithm that fits the model to the data.
- - **Model Fit**: Fit the model to the data using the chosen non-linear search, with on-the-fly results and visualization.
- - **Results**: Use the results of the search to interpret and visualize the model fit.
- - **Samples**: Use the samples of the search to inspect the parameter samples and visualize the probability density function of the results.
- - **Multiple Datasets**: Dedicated support for simultaneously fitting multiple datasets, enabling scalable analysis of large datasets.
-
-This overviews provides a high level of the basic API, with more advanced functionality described in the following
-overviews and the **PyAutoFit** cookbooks.
-
-__Contents__
-
-This overview is split into the following sections:
-
-- **Example Use Case**: Introduce the 1D Gaussian profile fitting example used throughout this overview.
-- **Model**: Define a 1D Gaussian as a PyAutoFit model via a Python class.
-- **Instances**: Create model instances by mapping parameter vectors to Python class instances.
-- **Analysis**: Define an ``Analysis`` class with a ``log_likelihood_function`` for fitting the model to data.
-- **Non Linear Search**: Select and configure a non-linear search algorithm (Dynesty nested sampling).
-- **Model Fit**: Execute the non-linear search to fit the model to the data.
-- **Result**: Examine the result and maximum likelihood instance from the search.
-- **Samples**: Access parameter samples and posterior information to visualize results.
-- **Multiple Datasets**: Fit multiple datasets simultaneously using AnalysisFactor objects.
-- **Factor Graph**: Combine AnalysisFactors into a FactorGraphModel for global model fitting.
-- **Wrap Up**: Summary of the basic PyAutoFit functionality.
-- **Resources**: Links to cookbooks and documentation for advanced features.
-- **Extending Models**: Example of composing multi-component models (Gaussian + Exponential).
-
-To begin, lets import ``autofit`` (and ``numpy``) using the convention below:
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import autofit as af
-import autofit.plot as aplt
-
-import matplotlib.pyplot as plt
-import numpy as np
-from os import path
-
-"""
-__Example Use Case__
-
-To illustrate **PyAutoFit** we'll use the example modeling problem of fitting a 1D Gaussian profile to noisy data.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-"""
-We plot the data with error bars below, showing the noisy 1D signal.
-"""
-xvalues = range(data.shape[0])
-
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.title("Example Data")
-plt.xlabel("x values of data (pixels)")
-plt.ylabel("Signal Value")
-plt.show()
-plt.close()
-
-"""
-The 1D signal was generated using a 1D Gaussian profile of the form:
-
-\begin{equation*}
-g(x, I, \sigma) = \frac{N}{\sigma\sqrt{2\pi}} \exp{(-0.5 (x / \sigma)^2)}
-\end{equation*}
-
-Where:
-
- ``x``: The x-axis coordinate where the ``Gaussian`` is evaluated.
-
- ``N``: The overall normalization of the Gaussian.
-
- ``sigma``: Describes the size of the Gaussian.
-
-Our modeling task is to fit the data with a 1D Gaussian and recover its parameters (``x``, ``N``, ``sigma``).
-
-__Model__
-
-We therefore need to define a 1D Gaussian as a **PyAutoFit** model.
-
-We do this by writing it as the following Python class:
-"""
-
-
-class Gaussian:
- def __init__(
- self,
- centre=0.0, # <- PyAutoFit recognises these constructor arguments
- normalization=0.1, # <- are the Gaussian`s model parameters.
- sigma=0.01,
- ):
- """
- Represents a 1D `Gaussian` profile, which can be treated as a PyAutoFit
- model-component whose free parameters (centre, normalization and sigma)
- are fitted for by a non-linear search.
-
- Parameters
- ----------
- centre
- The x coordinate of the profile centre.
- normalization
- Overall normalization of the `Gaussian` profile.
- sigma
- The sigma value controlling the size of the Gaussian.
- """
- self.centre = centre
- self.normalization = normalization
- self.sigma = sigma
-
- def model_data_from(self, xvalues: np.ndarray) -> np.ndarray:
- """
- Returns the 1D Gaussian profile on a line of Cartesian x coordinates.
-
- The input xvalues are translated to a coordinate system centred on the
- Gaussian, by subtracting its centre.
-
- The output is referred to as the `model_data` to signify that it is
- a representation of the data from the model.
-
- Parameters
- ----------
- xvalues
- The x coordinates for which the Gaussian is evaluated.
- """
- transformed_xvalues = xvalues - self.centre
-
- return np.multiply(
- np.divide(self.normalization, self.sigma * np.sqrt(2.0 * np.pi)),
- np.exp(-0.5 * np.square(np.divide(transformed_xvalues, self.sigma))),
- )
-
- @property
- def fwhm(self) -> float:
- """
- The full-width half-maximum of the Gaussian profile.
-
- This is used to illustrate latent variables in **PyAutoFit**, which are values that can be inferred from
- the free parameters of the model which we are interested and may want to store the full samples information
- on (e.g. to create posteriors).
- """
- return 2 * np.sqrt(2 * np.log(2)) * self.sigma
-
-
-"""
-The **PyAutoFit** model above uses the following format:
-
-- The name of the class is the name of the model, in this case, "Gaussian".
-
-- The input arguments of the constructor (the ``__init__`` method) are the parameters of the model, in this case ``centre``, ``normalization`` and ``sigma``.
-
-- The default values of the input arguments define whether a parameter is a single-valued ``float`` or a multi-valued ``tuple``. In this case, all 3 input parameters are floats.
-
-- It includes functions associated with that model component, which are used when fitting the model to data.
-
-To compose a model using the `Gaussian` class above we use the `af.Model` object.
-"""
-model = af.Model(Gaussian)
-print("Model `Gaussian` object: \n")
-print(model)
-
-"""
-The model has a total of 3 parameters:
-"""
-print(model.total_free_parameters)
-
-"""
-All model information is given by printing its `info` attribute.
-
-This shows that each model parameter has an associated prior.
-
-[The `info` below may not display optimally on your computer screen, for example the whitespace between parameter
-names on the left and parameter priors on the right may lead them to appear across multiple lines. This is a
-common issue in Jupyter notebooks.
-
-The`info_whitespace_length` parameter in the file `config/general.yaml` in the [output] section can be changed to
-increase or decrease the amount of whitespace (The Jupyter notebook kernel will need to be reset for this change to
-appear in a notebook).]
-"""
-print(model.info)
-
-"""
-The priors can be manually altered as follows, noting that these updated priors will be used below when we fit the
-model to data.
-"""
-model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.normalization = af.UniformPrior(lower_limit=0.0, upper_limit=1e2)
-model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
-
-"""
-Printing the `model.info` displayed these updated priors.
-"""
-print(model.info)
-
-"""
-The example above uses the most basic PyAutoFit API to compose a simple model. The API is highly extensible and
-can scale to models with thousands of parameters, complex hierarchies and relationships between parameters.
-A complete overview is given in the `model cookbook `_.
-
-__Instances__
-
-Instances of a **PyAutoFit** model (created via `af.Model`) can be generated by mapping an input `vector` of parameter
-values to create an instance of the model's Python class.
-
-To define the input `vector` correctly, we need to know the order of parameters in the model. This information is
-contained in the model's `paths` attribute.
-"""
-print(model.paths)
-
-"""
-We input values for the three free parameters of our model in the order specified by the `paths`
-attribute (i.e., `centre=30.0`, `normalization=2.0`, and `sigma=3.0`):
-"""
-instance = model.instance_from_vector(vector=[30.0, 2.0, 3.0])
-
-"""
-This is an instance of the ``Gaussian`` class.
-"""
-print("Model Instance: \n")
-print(instance)
-
-"""
-It has the parameters of the `Gaussian` with the values input above.
-"""
-print("Instance Parameters \n")
-print("x = ", instance.centre)
-print("normalization = ", instance.normalization)
-print("sigma = ", instance.sigma)
-
-"""
-We can use functions associated with the class, specifically the `model_data_from` function, to
-create a realization of the `Gaussian` and plot it.
-"""
-xvalues = np.arange(0.0, 100.0, 1.0)
-
-model_data = instance.model_data_from(xvalues=xvalues)
-
-plt.plot(xvalues, model_data, color="r")
-plt.title("1D Gaussian Model Data.")
-plt.xlabel("x values of profile")
-plt.ylabel("Gaussian Value")
-plt.show()
-plt.clf()
-
-"""
-This "model mapping", whereby models map to an instances of their Python classes, is integral to the core **PyAutoFit**
-API for model composition and fitting.
-
-Mapping models to instance of their Python classes is an integral part of the core **PyAutoFit** API. It enables
-the advanced model composition and results management tools illustrated in the following overviews and cookbooks.
-
-__Analysis__
-
-We now tell **PyAutoFit** how to fit the model to the data.
-
-We define an `Analysis` class, which includes:
-
-- An `__init__` constructor that takes `data` and `noise_map` as inputs (this can be extended with additional elements
- necessary for fitting the model to the data).
-
-- A `log_likelihood_function` that defines how to fit an `instance` of the model to the data and return a log
- likelihood value.
-
-Read the comments and docstrings of the `Analysis` class in detail for a full description of how the analysis works.
-works.
-"""
-
-
-class Analysis(af.Analysis):
- def __init__(self, data: np.ndarray, noise_map: np.ndarray):
- """
- The `Analysis` class acts as an interface between the data and model in **PyAutoFit**.
-
- Its `log_likelihood_function` defines how the model is fitted to the data and it is called many times by
- the non-linear search fitting algorithm.
-
- In this example the `Analysis` `__init__` constructor only contains the `data` and `noise-map`, but it can be
- easily extended to include other quantities.
-
- Parameters
- ----------
- data
- A 1D numpy array containing the data (e.g. a noisy 1D signal) fitted in the workspace examples.
- noise_map
- A 1D numpy array containing the noise values of the data, used for computing the goodness of fit
- metric, the log likelihood.
- """
- super().__init__()
-
- self.data = data
- self.noise_map = noise_map
-
- def log_likelihood_function(self, instance) -> float:
- """
- Returns the log likelihood of a fit of a 1D Gaussian to the dataset.
-
- The data is fitted using an `instance` of the `Gaussian` class where its `model_data_from`
- is called in order to create a model data representation of the Gaussian that is fitted to the data.
- """
-
- """
- The `instance` that comes into this method is an instance of the `Gaussian` model above, which was created
- via `af.Model()`.
-
- The parameter values are chosen by the non-linear search, based on where it thinks the high likelihood regions
- of parameter space are.
-
- The lines of Python code are commented out below to prevent excessive print statements when we run the
- non-linear search, but feel free to uncomment them and run the search to see the parameters of every instance
- that it fits.
- """
-
- # print("Gaussian Instance:")
- # print("Centre = ", instance.centre)
- # print("Normalization = ", instance.normalization)
- # print("Sigma = ", instance.sigma)
-
- """
- Get the range of x-values the data is defined on, to evaluate the model of the Gaussian.
- """
- xvalues = np.arange(self.data.shape[0])
-
- """
- Use these xvalues to create model data of our Gaussian.
- """
- model_data = instance.model_data_from(xvalues=xvalues)
-
- """
- Fit the model gaussian line data to the observed data, computing the residuals, chi-squared and log likelihood.
- """
- residual_map = self.data - model_data
- chi_squared_map = (residual_map / self.noise_map) ** 2.0
- chi_squared = sum(chi_squared_map)
- noise_normalization = np.sum(np.log(2 * np.pi * self.noise_map**2.0))
- log_likelihood = -0.5 * (chi_squared + noise_normalization)
-
- return log_likelihood
-
-
-"""
-Create an instance of the `Analysis` class by passing the `data` and `noise_map`.
-"""
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-"""
-The `Analysis` class shown above is the simplest example possible. The API is highly extensible and can include
-model-specific output, visualization and latent variable calculations. A complete overview is given in the
-analysis cookbook `_.
-
-__Non Linear Search__
-
-We now have a model ready to fit the data and an analysis class that performs this fit.
-
-Next, we need to select a fitting algorithm, known as a "non-linear search," to fit the model to the data.
-
-**PyAutoFit** supports various non-linear searches, which can be broadly categorized into three types:
-MCMC (Markov Chain Monte Carlo), nested sampling, and maximum likelihood estimators.
-
-For this example, we will use the nested sampling algorithm called Dynesty.
-"""
-search = af.DynestyStatic(
- nlive=100, # Example how to customize the search settings
-)
-
-"""
-The default settings of the non-linear search are specified in the configuration files of **PyAutoFit**, just
-like the default priors of the model components above. The ensures the basic API of your code is concise and
-readable, but with the flexibility to customize the search to your specific model-fitting problem.
-
-PyAutoFit supports a wide range of non-linear searches, including detailed visualuzation, support for parallel
-processing, and GPU and gradient based methods using the library JAX (https://jax.readthedocs.io/en/latest/).
-A complete overview is given in the `searches cookbook `_.
-
-__Model Fit__
-
-We begin the non-linear search by calling its `fit` method.
-
-This will take a minute or so to run.
-"""
-print(
- """
- The non-linear search has begun running.
- This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
- """
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-print("The search has finished run - you may now continue the notebook.")
-
-"""
-__Result__
-
-The result object returned by the fit provides information on the results of the non-linear search.
-
-The `info` attribute shows the result in a readable format.
-
-[Above, we discussed that the `info_whitespace_length` parameter in the config files could b changed to make
-the `model.info` attribute display optimally on your computer. This attribute also controls the whitespace of the
-`result.info` attribute.]
-"""
-print(result.info)
-
-"""
-Results are returned as instances of the model, as we illustrated above in the model mapping section.
-
-For example, we can print the result's maximum likelihood instance.
-"""
-print(result.max_log_likelihood_instance)
-
-print("\n Model-fit Max Log-likelihood Parameter Estimates: \n")
-print("Centre = ", result.max_log_likelihood_instance.centre)
-print("Normalization = ", result.max_log_likelihood_instance.normalization)
-print("Sigma = ", result.max_log_likelihood_instance.sigma)
-
-"""
-A benefit of the result being an instance is that we can use any of its methods to inspect the results.
-
-Below, we use the maximum likelihood instance to compare the maximum likelihood `Gaussian` to the data.
-"""
-model_data = result.max_log_likelihood_instance.model_data_from(
- xvalues=np.arange(data.shape[0])
-)
-
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(xvalues, model_data, color="r")
-plt.title("Dynesty model fit to 1D Gaussian dataset.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-__Samples__
-
-The results object also contains a ``Samples`` object, which contains all information on the non-linear search.
-
-This includes parameter samples, log likelihood values, posterior information and results internal to the specific
-algorithm (e.g. the internal dynesty samples).
-
-Below we use the samples to plot the probability density function cornerplot of the results.
-"""
-aplt.corner_anesthetic(samples=result.samples)
-
-"""
-The `results cookbook `_ also provides
-a run through of the samples object API.
-
-__Multiple Datasets__
-
-Many model-fitting problems require multiple datasets to be fitted simultaneously in order to provide the best
-constraints on the model.
-
-In **PyAutoFit**, all you have to do to fit multiple datasets is combine them with the model via `AnalysisFactor`
-objects.
-"""
-# For illustration purposes, we'll input the same data and noise-map as the example, but for a realistic example
-# you would input different datasets and noise-maps to each analysis.
-
-analysis_0 = Analysis(data=data, noise_map=noise_map)
-analysis_1 = Analysis(data=data, noise_map=noise_map)
-
-analysis_list = [analysis_0, analysis_1]
-
-analysis_factor_list = []
-
-for analysis in analysis_list:
-
- # The model can be customized here so that different model parameters are tied to each analysis.
- model_analysis = model.copy()
-
- analysis_factor = af.AnalysisFactor(prior_model=model_analysis, analysis=analysis)
-
- analysis_factor_list.append(analysis_factor)
-
-"""
-__Factor Graph__
-
-All `AnalysisFactor` objects are combined into a `FactorGraphModel`, which represents a global model fit to
-multiple datasets using a graphical model structure.
-
-The key outcomes of this setup are:
-
- - The individual log likelihoods from each `Analysis` object are summed to form the total log likelihood
- evaluated during the model-fitting process.
-
- - Results from all datasets are output to a unified directory, with subdirectories for visualizations
- from each analysis object, as defined by their `visualize` methods.
-
-This is a basic use of **PyAutoFit**'s graphical modeling capabilities, which support advanced hierarchical
-and probabilistic modeling for large, multi-dataset analyses.
-"""
-factor_graph = af.FactorGraphModel(*analysis_factor_list)
-
-"""
-To inspect the model, we print `factor_graph.global_prior_model.info`.
-"""
-print(factor_graph.global_prior_model.info)
-
-"""
-To fit multiple datasets, we pass the `FactorGraphModel` to a non-linear search.
-
-Unlike single-dataset fitting, we now pass the `factor_graph.global_prior_model` as the model and
-the `factor_graph` itself as the analysis object.
-
-This structure enables simultaneous fitting of multiple datasets in a consistent and scalable way.
-"""
-search = af.DynestyStatic(
- nlive=100,
-)
-
-result_list = search.fit(model=factor_graph.global_prior_model, analysis=factor_graph)
-
-"""
-The `multiple datasets cookbook `_ also
-provides a run through of the samples object API.
-
-__Wrap Up__
-
-This overview covers the basic functionality of **PyAutoFit** using a simple model, dataset, and model-fitting problem,
-demonstrating the fundamental aspects of its API.
-
-By now, you should have a clear understanding of how to define and compose your own models, fit them to data using
-a non-linear search, and interpret the results.
-
-The **PyAutoFit** API introduced here is highly extensible and customizable, making it adaptable to a wide range
-of model-fitting problems.
-
-The next overview will delve into setting up a scientific workflow with **PyAutoFit**, utilizing its API to
-optimize model-fitting efficiency and scalability for large datasets. This approach ensures that detailed scientific
-interpretation of the results remains feasible and insightful.
-
-__Resources__
-
-The `autofit_workspace: `_ repository on GitHub provides numerous
-examples demonstrating more complex model-fitting tasks.
-
-This includes cookbooks, which provide a concise reference guide to the **PyAutoFit** API for advanced model-fitting:
-
-- [Model Cookbook](https://pyautofit.readthedocs.io/en/latest/cookbooks/model.html): Learn how to compose complex models using multiple Python classes, lists, dictionaries, NumPy arrays and customize their parameterization.
-
-- [Analysis Cookbook](https://pyautofit.readthedocs.io/en/latest/cookbooks/search.html): Customize the analysis with model-specific output and visualization to gain deeper insights into your model fits.
-
-- [Searches Cookbook](https://pyautofit.readthedocs.io/en/latest/cookbooks/analysis.html): Choose from a variety of non-linear searches and customize their behavior. This includes options like outputting results to hard disk and parallelizing the search process.
-
-- [Results Cookbook](https://pyautofit.readthedocs.io/en/latest/cookbooks/result.html): Explore the various results available from a fit, such as parameter estimates, error estimates, model comparison metrics, and customizable visualizations.
-
-- [Configs Cookbook](https://pyautofit.readthedocs.io/en/latest/cookbooks/configs.html): Customize default settings using configuration files. This allows you to set priors, search settings, visualization preferences, and more.
-
-- [Multiple Dataset Cookbook](https://pyautofit.readthedocs.io/en/latest/cookbooks/multiple_datasets.html): Learn how to fit multiple datasets simultaneously by combining their analysis classes so that their log likelihoods are summed.
-
-These cookbooks provide detailed guides and examples to help you leverage the **PyAutoFit** API effectively for a wide range of model-fitting tasks.
-
-__Extending Models__
-
-The main overview is now complete, howeveer below we provide an example of how to compose and fit a model
-consisting of multiple components, which is a common requirement in many model-fitting problems.
-
-The model composition API is designed to make composing complex models, consisting of multiple components with many
-free parameters, straightforward and scalable.
-
-To illustrate this, we will extend our model to include a second component, representing a symmetric 1D Exponential
-profile, and fit it to data generated with both profiles.
-
-Lets begin by loading and plotting this data.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1__exponential_x1")
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-xvalues = range(data.shape[0])
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.title("Example Data With Multiple Components")
-plt.xlabel("x values of data (pixels)")
-plt.ylabel("Signal Value")
-plt.show()
-plt.close()
-
-"""
-We define a Python class for the `Exponential` model component, exactly as we did for the `Gaussian` above.
-"""
-
-
-class Exponential:
- def __init__(
- self,
- centre=30.0, # <- **PyAutoFit** recognises these constructor arguments
- normalization=1.0, # <- are the Exponentials`s model parameters.
- rate=0.01,
- ):
- """
- Represents a symmetric 1D Exponential profile.
-
- Parameters
- ----------
- centre
- The x coordinate of the profile centre.
- normalization
- Overall normalization of the profile.
- ratw
- The decay rate controlling has fast the Exponential declines.
- """
- self.centre = centre
- self.normalization = normalization
- self.rate = rate
-
- def model_data_from(self, xvalues: np.ndarray):
- """
- Returns the symmetric 1D Exponential on an input list of Cartesian x coordinates.
-
- The input xvalues are translated to a coordinate system centred on the Gaussian, via its `centre`.
-
- The output is referred to as the `model_data` to signify that it is a representation of the data from the
- model.
-
- Parameters
- ----------
- xvalues
- The x coordinates in the original reference frame of the data.
- """
- transformed_xvalues = np.subtract(xvalues, self.centre)
- return self.normalization * np.multiply(
- self.rate, np.exp(-1.0 * self.rate * abs(transformed_xvalues))
- )
-
-
-"""
-We can easily compose a model consisting of 1 `Gaussian` object and 1 `Exponential` object using the `af.Collection`
-object:
-"""
-model = af.Collection(gaussian=af.Model(Gaussian), exponential=af.Model(Exponential))
-
-"""
-A `Collection` behaves analogous to a `Model`, but it contains a multiple model components.
-
-We can see this by printing its `paths` attribute, where paths to all 6 free parameters via both model components
-are shown.
-
-The paths have the entries `.gaussian.` and `.exponential.`, which correspond to the names we input into
-the `af.Collection` above.
-"""
-print(model.paths)
-
-"""
-We can use the paths to customize the priors of each parameter.
-"""
-model.gaussian.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.gaussian.normalization = af.UniformPrior(lower_limit=0.0, upper_limit=1e2)
-model.gaussian.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
-model.exponential.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.exponential.normalization = af.UniformPrior(lower_limit=0.0, upper_limit=1e2)
-model.exponential.rate = af.UniformPrior(lower_limit=0.0, upper_limit=10.0)
-
-"""
-All of the information about the model created via the collection can be printed at once using its `info` attribute:
-"""
-print(model.info)
-
-"""
-A model instance can again be created by mapping an input `vector`, which now has 6 entries.
-"""
-instance = model.instance_from_vector(vector=[0.1, 0.2, 0.3, 0.4, 0.5, 0.01])
-
-"""
-This `instance` contains each of the model components we defined above.
-
-The argument names input into the `Collection` define the attribute names of the `instance`:
-"""
-print("Instance Parameters \n")
-print("x (Gaussian) = ", instance.gaussian.centre)
-print("normalization (Gaussian) = ", instance.gaussian.normalization)
-print("sigma (Gaussian) = ", instance.gaussian.sigma)
-print("x (Exponential) = ", instance.exponential.centre)
-print("normalization (Exponential) = ", instance.exponential.normalization)
-print("sigma (Exponential) = ", instance.exponential.rate)
-
-"""
-The `Analysis` class above assumed the `instance` contained only a single model-component.
-
-We update its `log_likelihood_function` to use both model components in the `instance` to fit the data.
-"""
-
-
-class Analysis(af.Analysis):
- def __init__(self, data: np.ndarray, noise_map: np.ndarray):
- """
- The `Analysis` class acts as an interface between the data and model in **PyAutoFit**.
-
- Its `log_likelihood_function` defines how the model is fitted to the data and it is called many times by
- the non-linear search fitting algorithm.
-
- In this example the `Analysis` `__init__` constructor only contains the `data` and `noise-map`, but it can be
- easily extended to include other quantities.
-
- Parameters
- ----------
- data
- A 1D numpy array containing the data (e.g. a noisy 1D signal) fitted in the workspace examples.
- noise_map
- A 1D numpy array containing the noise values of the data, used for computing the goodness of fit
- metric, the log likelihood.
- """
- super().__init__()
-
- self.data = data
- self.noise_map = noise_map
-
- def log_likelihood_function(self, instance) -> float:
- """
- Returns the log likelihood of a fit of a 1D Gaussian to the dataset.
-
- The data is fitted using an `instance` of multiple 1D profiles (e.g. a `Gaussian`, `Exponential`) where
- their `model_data_from` methods are called and sumed in order to create a model data
- representation that is fitted to the data.
- """
-
- """
- The `instance` that comes into this method is an instance of the `Gaussian` and `Exponential` models above,
- which were created via `af.Collection()`.
-
- It contains instances of every class we instantiated it with, where each instance is named following the names
- given to the Collection, which in this example is a `Gaussian` (with name `gaussian) and Exponential (with
- name `exponential`).
-
- The parameter values are again chosen by the non-linear search, based on where it thinks the high likelihood
- regions of parameter space are. The lines of Python code are commented out below to prevent excessive print
- statements.
- """
-
- # print("Gaussian Instance:")
- # print("Centre = ", instance.gaussian.centre)
- # print("Normalization = ", instance.gaussian.normalization)
- # print("Sigma = ", instance.gaussian.sigma)
-
- # print("Exponential Instance:")
- # print("Centre = ", instance.exponential.centre)
- # print("Normalization = ", instance.exponential.normalization)
- # print("Rate = ", instance.exponential.rate)
-
- """
- Get the range of x-values the data is defined on, to evaluate the model of the Gaussian.
- """
- xvalues = np.arange(self.data.shape[0])
-
- """
- Internally, the `instance` variable is a list of all model components pass to the `Collection` above.
-
- we can therefore iterate over them and use their `model_data_from` methods to create the
- summed overall model data.
- """
- model_data = sum(
- [profile_1d.model_data_from(xvalues=xvalues) for profile_1d in instance]
- )
-
- """
- Fit the model gaussian line data to the observed data, computing the residuals, chi-squared and log likelihood.
- """
- residual_map = self.data - model_data
- chi_squared_map = (residual_map / self.noise_map) ** 2.0
- chi_squared = sum(chi_squared_map)
- noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
- log_likelihood = -0.5 * (chi_squared + noise_normalization)
-
- return log_likelihood
-
-
-"""
-We can now fit this model to the data using the same API we did before.
-"""
-analysis = Analysis(data=data, noise_map=noise_map)
-
-search = af.DynestyStatic(
- nlive=100,
- sample="rwalk",
- number_of_cores=1,
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-"""
-The `info` attribute shows the result in a readable format, showing that all 6 free parameters were fitted for.
-"""
-print(result.info)
-
-"""
-We can again use the max log likelihood instance to visualize the model data of the best fit model compared to the
-data.
-"""
-instance = result.max_log_likelihood_instance
-
-model_gaussian = instance.gaussian.model_data_from(xvalues=np.arange(data.shape[0]))
-model_exponential = instance.exponential.model_data_from(
- xvalues=np.arange(data.shape[0])
-)
-model_data = model_gaussian + model_exponential
-
-plt.errorbar(
- x=xvalues,
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(range(data.shape[0]), model_data, color="r")
-plt.plot(range(data.shape[0]), model_gaussian, "--")
-plt.plot(range(data.shape[0]), model_exponential, "--")
-plt.title("Dynesty model fit to 1D Gaussian + Exponential dataset.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-Finish.
-"""
+"""
+Overview: The Basics
+--------------------
+
+**PyAutoFit** is a Python based probabilistic programming language for model fitting and Bayesian inference
+of large datasets.
+
+The basic **PyAutoFit** API allows us a user to quickly compose a probabilistic model and fit it to data via a
+log likelihood function, using a range of non-linear search algorithms (e.g. MCMC, nested sampling).
+
+This overview gives a run through of:
+
+ - **Models**: Use Python classes to compose the model which is fitted to data.
+ - **Instances**: Create instances of the model via its Python class.
+ - **Analysis**: Define an ``Analysis`` class which includes the log likelihood function that fits the model to the data.
+ - **Searches**: Choose an MCMC, nested sampling or maximum likelihood estimator non-linear search algorithm that fits the model to the data.
+ - **Model Fit**: Fit the model to the data using the chosen non-linear search, with on-the-fly results and visualization.
+ - **Results**: Use the results of the search to interpret and visualize the model fit.
+ - **Samples**: Use the samples of the search to inspect the parameter samples and visualize the probability density function of the results.
+ - **Multiple Datasets**: Dedicated support for simultaneously fitting multiple datasets, enabling scalable analysis of large datasets.
+
+This overviews provides a high level of the basic API, with more advanced functionality described in the following
+overviews and the **PyAutoFit** cookbooks.
+
+__Contents__
+
+This overview is split into the following sections:
+
+- **Example Use Case**: Introduce the 1D Gaussian profile fitting example used throughout this overview.
+- **Model**: Define a 1D Gaussian as a PyAutoFit model via a Python class.
+- **Instances**: Create model instances by mapping parameter vectors to Python class instances.
+- **Analysis**: Define an ``Analysis`` class with a ``log_likelihood_function`` for fitting the model to data.
+- **Non Linear Search**: Select and configure a non-linear search algorithm (Dynesty nested sampling).
+- **Model Fit**: Execute the non-linear search to fit the model to the data.
+- **Result**: Examine the result and maximum likelihood instance from the search.
+- **Samples**: Access parameter samples and posterior information to visualize results.
+- **Multiple Datasets**: Fit multiple datasets simultaneously using AnalysisFactor objects.
+- **Factor Graph**: Combine AnalysisFactors into a FactorGraphModel for global model fitting.
+- **Wrap Up**: Summary of the basic PyAutoFit functionality.
+- **Resources**: Links to cookbooks and documentation for advanced features.
+- **Extending Models**: Example of composing multi-component models (Gaussian + Exponential).
+
+To begin, lets import ``autofit`` (and ``numpy``) using the convention below:
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import autofit as af
+import autofit.plot as aplt
+
+import matplotlib.pyplot as plt
+import numpy as np
+from os import path
+
+"""
+__Example Use Case__
+
+To illustrate **PyAutoFit** we'll use the example modeling problem of fitting a 1D Gaussian profile to noisy data.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+"""
+We plot the data with error bars below, showing the noisy 1D signal.
+"""
+xvalues = range(data.shape[0])
+
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.title("Example Data")
+plt.xlabel("x values of data (pixels)")
+plt.ylabel("Signal Value")
+plt.show()
+plt.close()
+
+"""
+The 1D signal was generated using a 1D Gaussian profile of the form:
+
+\begin{equation*}
+g(x, I, \sigma) = \frac{N}{\sigma\sqrt{2\pi}} \exp{(-0.5 (x / \sigma)^2)}
+\end{equation*}
+
+Where:
+
+ ``x``: The x-axis coordinate where the ``Gaussian`` is evaluated.
+
+ ``N``: The overall normalization of the Gaussian.
+
+ ``sigma``: Describes the size of the Gaussian.
+
+Our modeling task is to fit the data with a 1D Gaussian and recover its parameters (``x``, ``N``, ``sigma``).
+
+__Model__
+
+We therefore need to define a 1D Gaussian as a **PyAutoFit** model.
+
+We do this by writing it as the following Python class:
+"""
+
+
+class Gaussian:
+ def __init__(
+ self,
+ centre=0.0, # <- PyAutoFit recognises these constructor arguments
+ normalization=0.1, # <- are the Gaussian`s model parameters.
+ sigma=0.01,
+ ):
+ """
+ Represents a 1D `Gaussian` profile, which can be treated as a PyAutoFit
+ model-component whose free parameters (centre, normalization and sigma)
+ are fitted for by a non-linear search.
+
+ Parameters
+ ----------
+ centre
+ The x coordinate of the profile centre.
+ normalization
+ Overall normalization of the `Gaussian` profile.
+ sigma
+ The sigma value controlling the size of the Gaussian.
+ """
+ self.centre = centre
+ self.normalization = normalization
+ self.sigma = sigma
+
+ def model_data_from(self, xvalues: np.ndarray) -> np.ndarray:
+ """
+ Returns the 1D Gaussian profile on a line of Cartesian x coordinates.
+
+ The input xvalues are translated to a coordinate system centred on the
+ Gaussian, by subtracting its centre.
+
+ The output is referred to as the `model_data` to signify that it is
+ a representation of the data from the model.
+
+ Parameters
+ ----------
+ xvalues
+ The x coordinates for which the Gaussian is evaluated.
+ """
+ transformed_xvalues = xvalues - self.centre
+
+ return np.multiply(
+ np.divide(self.normalization, self.sigma * np.sqrt(2.0 * np.pi)),
+ np.exp(-0.5 * np.square(np.divide(transformed_xvalues, self.sigma))),
+ )
+
+ @property
+ def fwhm(self) -> float:
+ """
+ The full-width half-maximum of the Gaussian profile.
+
+ This is used to illustrate latent variables in **PyAutoFit**, which are values that can be inferred from
+ the free parameters of the model which we are interested and may want to store the full samples information
+ on (e.g. to create posteriors).
+ """
+ return 2 * np.sqrt(2 * np.log(2)) * self.sigma
+
+
+"""
+The **PyAutoFit** model above uses the following format:
+
+- The name of the class is the name of the model, in this case, "Gaussian".
+
+- The input arguments of the constructor (the ``__init__`` method) are the parameters of the model, in this case ``centre``, ``normalization`` and ``sigma``.
+
+- The default values of the input arguments define whether a parameter is a single-valued ``float`` or a multi-valued ``tuple``. In this case, all 3 input parameters are floats.
+
+- It includes functions associated with that model component, which are used when fitting the model to data.
+
+To compose a model using the `Gaussian` class above we use the `af.Model` object.
+"""
+model = af.Model(Gaussian)
+print("Model `Gaussian` object: \n")
+print(model)
+
+"""
+The model has a total of 3 parameters:
+"""
+print(model.total_free_parameters)
+
+"""
+All model information is given by printing its `info` attribute.
+
+This shows that each model parameter has an associated prior.
+
+[The `info` below may not display optimally on your computer screen, for example the whitespace between parameter
+names on the left and parameter priors on the right may lead them to appear across multiple lines. This is a
+common issue in Jupyter notebooks.
+
+The`info_whitespace_length` parameter in the file `config/general.yaml` in the [output] section can be changed to
+increase or decrease the amount of whitespace (The Jupyter notebook kernel will need to be reset for this change to
+appear in a notebook).]
+"""
+print(model.info)
+
+"""
+The priors can be manually altered as follows, noting that these updated priors will be used below when we fit the
+model to data.
+"""
+model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.normalization = af.UniformPrior(lower_limit=0.0, upper_limit=1e2)
+model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
+
+"""
+Printing the `model.info` displayed these updated priors.
+"""
+print(model.info)
+
+"""
+The example above uses the most basic PyAutoFit API to compose a simple model. The API is highly extensible and
+can scale to models with thousands of parameters, complex hierarchies and relationships between parameters.
+A complete overview is given in the `model cookbook `_.
+
+__Instances__
+
+Instances of a **PyAutoFit** model (created via `af.Model`) can be generated by mapping an input `vector` of parameter
+values to create an instance of the model's Python class.
+
+To define the input `vector` correctly, we need to know the order of parameters in the model. This information is
+contained in the model's `paths` attribute.
+"""
+print(model.paths)
+
+"""
+We input values for the three free parameters of our model in the order specified by the `paths`
+attribute (i.e., `centre=30.0`, `normalization=2.0`, and `sigma=3.0`):
+"""
+instance = model.instance_from_vector(vector=[30.0, 2.0, 3.0])
+
+"""
+This is an instance of the ``Gaussian`` class.
+"""
+print("Model Instance: \n")
+print(instance)
+
+"""
+It has the parameters of the `Gaussian` with the values input above.
+"""
+print("Instance Parameters \n")
+print("x = ", instance.centre)
+print("normalization = ", instance.normalization)
+print("sigma = ", instance.sigma)
+
+"""
+We can use functions associated with the class, specifically the `model_data_from` function, to
+create a realization of the `Gaussian` and plot it.
+"""
+xvalues = np.arange(0.0, 100.0, 1.0)
+
+model_data = instance.model_data_from(xvalues=xvalues)
+
+plt.plot(xvalues, model_data, color="r")
+plt.title("1D Gaussian Model Data.")
+plt.xlabel("x values of profile")
+plt.ylabel("Gaussian Value")
+plt.show()
+plt.clf()
+
+"""
+This "model mapping", whereby models map to an instances of their Python classes, is integral to the core **PyAutoFit**
+API for model composition and fitting.
+
+Mapping models to instance of their Python classes is an integral part of the core **PyAutoFit** API. It enables
+the advanced model composition and results management tools illustrated in the following overviews and cookbooks.
+
+__Analysis__
+
+We now tell **PyAutoFit** how to fit the model to the data.
+
+We define an `Analysis` class, which includes:
+
+- An `__init__` constructor that takes `data` and `noise_map` as inputs (this can be extended with additional elements
+ necessary for fitting the model to the data).
+
+- A `log_likelihood_function` that defines how to fit an `instance` of the model to the data and return a log
+ likelihood value.
+
+Read the comments and docstrings of the `Analysis` class in detail for a full description of how the analysis works.
+works.
+"""
+
+
+class Analysis(af.Analysis):
+ def __init__(self, data: np.ndarray, noise_map: np.ndarray):
+ """
+ The `Analysis` class acts as an interface between the data and model in **PyAutoFit**.
+
+ Its `log_likelihood_function` defines how the model is fitted to the data and it is called many times by
+ the non-linear search fitting algorithm.
+
+ In this example the `Analysis` `__init__` constructor only contains the `data` and `noise-map`, but it can be
+ easily extended to include other quantities.
+
+ Parameters
+ ----------
+ data
+ A 1D numpy array containing the data (e.g. a noisy 1D signal) fitted in the workspace examples.
+ noise_map
+ A 1D numpy array containing the noise values of the data, used for computing the goodness of fit
+ metric, the log likelihood.
+ """
+ super().__init__()
+
+ self.data = data
+ self.noise_map = noise_map
+
+ def log_likelihood_function(self, instance) -> float:
+ """
+ Returns the log likelihood of a fit of a 1D Gaussian to the dataset.
+
+ The data is fitted using an `instance` of the `Gaussian` class where its `model_data_from`
+ is called in order to create a model data representation of the Gaussian that is fitted to the data.
+ """
+
+ """
+ The `instance` that comes into this method is an instance of the `Gaussian` model above, which was created
+ via `af.Model()`.
+
+ The parameter values are chosen by the non-linear search, based on where it thinks the high likelihood regions
+ of parameter space are.
+
+ The lines of Python code are commented out below to prevent excessive print statements when we run the
+ non-linear search, but feel free to uncomment them and run the search to see the parameters of every instance
+ that it fits.
+ """
+
+ # print("Gaussian Instance:")
+ # print("Centre = ", instance.centre)
+ # print("Normalization = ", instance.normalization)
+ # print("Sigma = ", instance.sigma)
+
+ """
+ Get the range of x-values the data is defined on, to evaluate the model of the Gaussian.
+ """
+ xvalues = np.arange(self.data.shape[0])
+
+ """
+ Use these xvalues to create model data of our Gaussian.
+ """
+ model_data = instance.model_data_from(xvalues=xvalues)
+
+ """
+ Fit the model gaussian line data to the observed data, computing the residuals, chi-squared and log likelihood.
+ """
+ residual_map = self.data - model_data
+ chi_squared_map = (residual_map / self.noise_map) ** 2.0
+ chi_squared = sum(chi_squared_map)
+ noise_normalization = np.sum(np.log(2 * np.pi * self.noise_map**2.0))
+ log_likelihood = -0.5 * (chi_squared + noise_normalization)
+
+ return log_likelihood
+
+
+"""
+Create an instance of the `Analysis` class by passing the `data` and `noise_map`.
+"""
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+"""
+The `Analysis` class shown above is the simplest example possible. The API is highly extensible and can include
+model-specific output, visualization and latent variable calculations. A complete overview is given in the
+analysis cookbook `_.
+
+__Non Linear Search__
+
+We now have a model ready to fit the data and an analysis class that performs this fit.
+
+Next, we need to select a fitting algorithm, known as a "non-linear search," to fit the model to the data.
+
+**PyAutoFit** supports various non-linear searches, which can be broadly categorized into three types:
+MCMC (Markov Chain Monte Carlo), nested sampling, and maximum likelihood estimators.
+
+For this example, we will use the nested sampling algorithm called Dynesty.
+"""
+search = af.DynestyStatic(
+ nlive=100, # Example how to customize the search settings
+)
+
+"""
+The default settings of the non-linear search are specified in the configuration files of **PyAutoFit**, just
+like the default priors of the model components above. The ensures the basic API of your code is concise and
+readable, but with the flexibility to customize the search to your specific model-fitting problem.
+
+PyAutoFit supports a wide range of non-linear searches, including detailed visualuzation, support for parallel
+processing, and GPU and gradient based methods using the library JAX (https://jax.readthedocs.io/en/latest/).
+A complete overview is given in the `searches cookbook `_.
+
+__Model Fit__
+
+We begin the non-linear search by calling its `fit` method.
+
+This will take a minute or so to run.
+"""
+print(
+ """
+ The non-linear search has begun running.
+ This Jupyter notebook cell with progress once the search has completed - this could take a few minutes!
+ """
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+print("The search has finished run - you may now continue the notebook.")
+
+"""
+__Result__
+
+The result object returned by the fit provides information on the results of the non-linear search.
+
+The `info` attribute shows the result in a readable format.
+
+[Above, we discussed that the `info_whitespace_length` parameter in the config files could b changed to make
+the `model.info` attribute display optimally on your computer. This attribute also controls the whitespace of the
+`result.info` attribute.]
+"""
+print(result.info)
+
+"""
+Results are returned as instances of the model, as we illustrated above in the model mapping section.
+
+For example, we can print the result's maximum likelihood instance.
+"""
+print(result.max_log_likelihood_instance)
+
+print("\n Model-fit Max Log-likelihood Parameter Estimates: \n")
+print("Centre = ", result.max_log_likelihood_instance.centre)
+print("Normalization = ", result.max_log_likelihood_instance.normalization)
+print("Sigma = ", result.max_log_likelihood_instance.sigma)
+
+"""
+A benefit of the result being an instance is that we can use any of its methods to inspect the results.
+
+Below, we use the maximum likelihood instance to compare the maximum likelihood `Gaussian` to the data.
+"""
+model_data = result.max_log_likelihood_instance.model_data_from(
+ xvalues=np.arange(data.shape[0])
+)
+
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(xvalues, model_data, color="r")
+plt.title("Dynesty model fit to 1D Gaussian dataset.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+__Samples__
+
+The results object also contains a ``Samples`` object, which contains all information on the non-linear search.
+
+This includes parameter samples, log likelihood values, posterior information and results internal to the specific
+algorithm (e.g. the internal dynesty samples).
+
+Below we use the samples to plot the probability density function cornerplot of the results.
+"""
+aplt.corner_anesthetic(samples=result.samples)
+
+"""
+The `results cookbook `_ also provides
+a run through of the samples object API.
+
+__Multiple Datasets__
+
+Many model-fitting problems require multiple datasets to be fitted simultaneously in order to provide the best
+constraints on the model.
+
+In **PyAutoFit**, all you have to do to fit multiple datasets is combine them with the model via `AnalysisFactor`
+objects.
+"""
+# For illustration purposes, we'll input the same data and noise-map as the example, but for a realistic example
+# you would input different datasets and noise-maps to each analysis.
+
+analysis_0 = Analysis(data=data, noise_map=noise_map)
+analysis_1 = Analysis(data=data, noise_map=noise_map)
+
+analysis_list = [analysis_0, analysis_1]
+
+analysis_factor_list = []
+
+for analysis in analysis_list:
+
+ # The model can be customized here so that different model parameters are tied to each analysis.
+ model_analysis = model.copy()
+
+ analysis_factor = af.AnalysisFactor(prior_model=model_analysis, analysis=analysis)
+
+ analysis_factor_list.append(analysis_factor)
+
+"""
+__Factor Graph__
+
+All `AnalysisFactor` objects are combined into a `FactorGraphModel`, which represents a global model fit to
+multiple datasets using a graphical model structure.
+
+The key outcomes of this setup are:
+
+ - The individual log likelihoods from each `Analysis` object are summed to form the total log likelihood
+ evaluated during the model-fitting process.
+
+ - Results from all datasets are output to a unified directory, with subdirectories for visualizations
+ from each analysis object, as defined by their `visualize` methods.
+
+This is a basic use of **PyAutoFit**'s graphical modeling capabilities, which support advanced hierarchical
+and probabilistic modeling for large, multi-dataset analyses.
+"""
+factor_graph = af.FactorGraphModel(*analysis_factor_list)
+
+"""
+To inspect the model, we print `factor_graph.global_prior_model.info`.
+"""
+print(factor_graph.global_prior_model.info)
+
+"""
+To fit multiple datasets, we pass the `FactorGraphModel` to a non-linear search.
+
+Unlike single-dataset fitting, we now pass the `factor_graph.global_prior_model` as the model and
+the `factor_graph` itself as the analysis object.
+
+This structure enables simultaneous fitting of multiple datasets in a consistent and scalable way.
+"""
+search = af.DynestyStatic(
+ nlive=100,
+)
+
+result_list = search.fit(model=factor_graph.global_prior_model, analysis=factor_graph)
+
+"""
+The `multiple datasets cookbook `_ also
+provides a run through of the samples object API.
+
+__Wrap Up__
+
+This overview covers the basic functionality of **PyAutoFit** using a simple model, dataset, and model-fitting problem,
+demonstrating the fundamental aspects of its API.
+
+By now, you should have a clear understanding of how to define and compose your own models, fit them to data using
+a non-linear search, and interpret the results.
+
+The **PyAutoFit** API introduced here is highly extensible and customizable, making it adaptable to a wide range
+of model-fitting problems.
+
+The next overview will delve into setting up a scientific workflow with **PyAutoFit**, utilizing its API to
+optimize model-fitting efficiency and scalability for large datasets. This approach ensures that detailed scientific
+interpretation of the results remains feasible and insightful.
+
+__Resources__
+
+The `autofit_workspace: `_ repository on GitHub provides numerous
+examples demonstrating more complex model-fitting tasks.
+
+This includes cookbooks, which provide a concise reference guide to the **PyAutoFit** API for advanced model-fitting:
+
+- [Model Cookbook](https://pyautofit.readthedocs.io/en/latest/cookbooks/model.html): Learn how to compose complex models using multiple Python classes, lists, dictionaries, NumPy arrays and customize their parameterization.
+
+- [Analysis Cookbook](https://pyautofit.readthedocs.io/en/latest/cookbooks/search.html): Customize the analysis with model-specific output and visualization to gain deeper insights into your model fits.
+
+- [Searches Cookbook](https://pyautofit.readthedocs.io/en/latest/cookbooks/analysis.html): Choose from a variety of non-linear searches and customize their behavior. This includes options like outputting results to hard disk and parallelizing the search process.
+
+- [Results Cookbook](https://pyautofit.readthedocs.io/en/latest/cookbooks/result.html): Explore the various results available from a fit, such as parameter estimates, error estimates, model comparison metrics, and customizable visualizations.
+
+- [Configs Cookbook](https://pyautofit.readthedocs.io/en/latest/cookbooks/configs.html): Customize default settings using configuration files. This allows you to set priors, search settings, visualization preferences, and more.
+
+- [Multiple Dataset Cookbook](https://pyautofit.readthedocs.io/en/latest/cookbooks/multiple_datasets.html): Learn how to fit multiple datasets simultaneously by combining their analysis classes so that their log likelihoods are summed.
+
+These cookbooks provide detailed guides and examples to help you leverage the **PyAutoFit** API effectively for a wide range of model-fitting tasks.
+
+__Extending Models__
+
+The main overview is now complete, howeveer below we provide an example of how to compose and fit a model
+consisting of multiple components, which is a common requirement in many model-fitting problems.
+
+The model composition API is designed to make composing complex models, consisting of multiple components with many
+free parameters, straightforward and scalable.
+
+To illustrate this, we will extend our model to include a second component, representing a symmetric 1D Exponential
+profile, and fit it to data generated with both profiles.
+
+Lets begin by loading and plotting this data.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1__exponential_x1")
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+xvalues = range(data.shape[0])
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.title("Example Data With Multiple Components")
+plt.xlabel("x values of data (pixels)")
+plt.ylabel("Signal Value")
+plt.show()
+plt.close()
+
+"""
+We define a Python class for the `Exponential` model component, exactly as we did for the `Gaussian` above.
+"""
+
+
+class Exponential:
+ def __init__(
+ self,
+ centre=30.0, # <- **PyAutoFit** recognises these constructor arguments
+ normalization=1.0, # <- are the Exponentials`s model parameters.
+ rate=0.01,
+ ):
+ """
+ Represents a symmetric 1D Exponential profile.
+
+ Parameters
+ ----------
+ centre
+ The x coordinate of the profile centre.
+ normalization
+ Overall normalization of the profile.
+ ratw
+ The decay rate controlling has fast the Exponential declines.
+ """
+ self.centre = centre
+ self.normalization = normalization
+ self.rate = rate
+
+ def model_data_from(self, xvalues: np.ndarray):
+ """
+ Returns the symmetric 1D Exponential on an input list of Cartesian x coordinates.
+
+ The input xvalues are translated to a coordinate system centred on the Gaussian, via its `centre`.
+
+ The output is referred to as the `model_data` to signify that it is a representation of the data from the
+ model.
+
+ Parameters
+ ----------
+ xvalues
+ The x coordinates in the original reference frame of the data.
+ """
+ transformed_xvalues = np.subtract(xvalues, self.centre)
+ return self.normalization * np.multiply(
+ self.rate, np.exp(-1.0 * self.rate * abs(transformed_xvalues))
+ )
+
+
+"""
+We can easily compose a model consisting of 1 `Gaussian` object and 1 `Exponential` object using the `af.Collection`
+object:
+"""
+model = af.Collection(gaussian=af.Model(Gaussian), exponential=af.Model(Exponential))
+
+"""
+A `Collection` behaves analogous to a `Model`, but it contains a multiple model components.
+
+We can see this by printing its `paths` attribute, where paths to all 6 free parameters via both model components
+are shown.
+
+The paths have the entries `.gaussian.` and `.exponential.`, which correspond to the names we input into
+the `af.Collection` above.
+"""
+print(model.paths)
+
+"""
+We can use the paths to customize the priors of each parameter.
+"""
+model.gaussian.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.gaussian.normalization = af.UniformPrior(lower_limit=0.0, upper_limit=1e2)
+model.gaussian.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
+model.exponential.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.exponential.normalization = af.UniformPrior(lower_limit=0.0, upper_limit=1e2)
+model.exponential.rate = af.UniformPrior(lower_limit=0.0, upper_limit=10.0)
+
+"""
+All of the information about the model created via the collection can be printed at once using its `info` attribute:
+"""
+print(model.info)
+
+"""
+A model instance can again be created by mapping an input `vector`, which now has 6 entries.
+"""
+instance = model.instance_from_vector(vector=[0.1, 0.2, 0.3, 0.4, 0.5, 0.01])
+
+"""
+This `instance` contains each of the model components we defined above.
+
+The argument names input into the `Collection` define the attribute names of the `instance`:
+"""
+print("Instance Parameters \n")
+print("x (Gaussian) = ", instance.gaussian.centre)
+print("normalization (Gaussian) = ", instance.gaussian.normalization)
+print("sigma (Gaussian) = ", instance.gaussian.sigma)
+print("x (Exponential) = ", instance.exponential.centre)
+print("normalization (Exponential) = ", instance.exponential.normalization)
+print("sigma (Exponential) = ", instance.exponential.rate)
+
+"""
+The `Analysis` class above assumed the `instance` contained only a single model-component.
+
+We update its `log_likelihood_function` to use both model components in the `instance` to fit the data.
+"""
+
+
+class Analysis(af.Analysis):
+ def __init__(self, data: np.ndarray, noise_map: np.ndarray):
+ """
+ The `Analysis` class acts as an interface between the data and model in **PyAutoFit**.
+
+ Its `log_likelihood_function` defines how the model is fitted to the data and it is called many times by
+ the non-linear search fitting algorithm.
+
+ In this example the `Analysis` `__init__` constructor only contains the `data` and `noise-map`, but it can be
+ easily extended to include other quantities.
+
+ Parameters
+ ----------
+ data
+ A 1D numpy array containing the data (e.g. a noisy 1D signal) fitted in the workspace examples.
+ noise_map
+ A 1D numpy array containing the noise values of the data, used for computing the goodness of fit
+ metric, the log likelihood.
+ """
+ super().__init__()
+
+ self.data = data
+ self.noise_map = noise_map
+
+ def log_likelihood_function(self, instance) -> float:
+ """
+ Returns the log likelihood of a fit of a 1D Gaussian to the dataset.
+
+ The data is fitted using an `instance` of multiple 1D profiles (e.g. a `Gaussian`, `Exponential`) where
+ their `model_data_from` methods are called and sumed in order to create a model data
+ representation that is fitted to the data.
+ """
+
+ """
+ The `instance` that comes into this method is an instance of the `Gaussian` and `Exponential` models above,
+ which were created via `af.Collection()`.
+
+ It contains instances of every class we instantiated it with, where each instance is named following the names
+ given to the Collection, which in this example is a `Gaussian` (with name `gaussian) and Exponential (with
+ name `exponential`).
+
+ The parameter values are again chosen by the non-linear search, based on where it thinks the high likelihood
+ regions of parameter space are. The lines of Python code are commented out below to prevent excessive print
+ statements.
+ """
+
+ # print("Gaussian Instance:")
+ # print("Centre = ", instance.gaussian.centre)
+ # print("Normalization = ", instance.gaussian.normalization)
+ # print("Sigma = ", instance.gaussian.sigma)
+
+ # print("Exponential Instance:")
+ # print("Centre = ", instance.exponential.centre)
+ # print("Normalization = ", instance.exponential.normalization)
+ # print("Rate = ", instance.exponential.rate)
+
+ """
+ Get the range of x-values the data is defined on, to evaluate the model of the Gaussian.
+ """
+ xvalues = np.arange(self.data.shape[0])
+
+ """
+ Internally, the `instance` variable is a list of all model components pass to the `Collection` above.
+
+ we can therefore iterate over them and use their `model_data_from` methods to create the
+ summed overall model data.
+ """
+ model_data = sum(
+ [profile_1d.model_data_from(xvalues=xvalues) for profile_1d in instance]
+ )
+
+ """
+ Fit the model gaussian line data to the observed data, computing the residuals, chi-squared and log likelihood.
+ """
+ residual_map = self.data - model_data
+ chi_squared_map = (residual_map / self.noise_map) ** 2.0
+ chi_squared = sum(chi_squared_map)
+ noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
+ log_likelihood = -0.5 * (chi_squared + noise_normalization)
+
+ return log_likelihood
+
+
+"""
+We can now fit this model to the data using the same API we did before.
+"""
+analysis = Analysis(data=data, noise_map=noise_map)
+
+search = af.DynestyStatic(
+ nlive=100,
+ sample="rwalk",
+ number_of_cores=1,
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+"""
+The `info` attribute shows the result in a readable format, showing that all 6 free parameters were fitted for.
+"""
+print(result.info)
+
+"""
+We can again use the max log likelihood instance to visualize the model data of the best fit model compared to the
+data.
+"""
+instance = result.max_log_likelihood_instance
+
+model_gaussian = instance.gaussian.model_data_from(xvalues=np.arange(data.shape[0]))
+model_exponential = instance.exponential.model_data_from(
+ xvalues=np.arange(data.shape[0])
+)
+model_data = model_gaussian + model_exponential
+
+plt.errorbar(
+ x=xvalues,
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(range(data.shape[0]), model_data, color="r")
+plt.plot(range(data.shape[0]), model_gaussian, "--")
+plt.plot(range(data.shape[0]), model_exponential, "--")
+plt.title("Dynesty model fit to 1D Gaussian + Exponential dataset.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+Finish.
+"""
diff --git a/scripts/overview/overview_2_scientific_workflow.py b/scripts/overview/overview_2_scientific_workflow.py
index cf0f0d45..5e0f37b1 100644
--- a/scripts/overview/overview_2_scientific_workflow.py
+++ b/scripts/overview/overview_2_scientific_workflow.py
@@ -1,704 +1,700 @@
-"""
-Overview: Scientific Workflow
-=============================
-
-A scientific workflow comprises the tasks you perform to conduct a scientific study. This includes fitting models to
-datasets, interpreting the results, and gaining insights into your scientific problem.
-
-Different problems require different scientific workflows, depending on factors such as model complexity, dataset size,
-and computational run times. For example, some problems involve fitting a single dataset with many models to gain
-scientific insights, while others involve fitting thousands of datasets with a single model for large-scale studies.
-
-The **PyAutoFit** API is flexible, customizable, and extensible, enabling users to develop scientific workflows
-tailored to their specific problems.
-
-This overview covers the key features of **PyAutoFit** that support the development of effective scientific workflows:
-
-- **On The Fly**: Display results immediately (e.g., in Jupyter notebooks) to provide instant feedback for adapting your workflow.
-- **Hard Disk Output**: Output results to hard disk with high customization, allowing quick and detailed inspection of fits to many datasets.
-- **Visualization**: Generate model-specific visualizations to create custom plots that streamline result inspection.
-- **Loading Results**: Load results from the hard disk to inspect and interpret the outcomes of a model fit.
-- **Result Customization**: Customize the returned results to simplify scientific interpretation.
-- **Model Composition**: Extensible model composition makes it easy to fit many models with different parameterizations and assumptions.
-- **Searches**: Support for various non-linear searches (e.g., nested sampling, MCMC), including gradient based fitting using JAX, to find the right method for your problem.
-- **Configs**: Configuration files that set default model, fitting, and visualization behaviors, streamlining model fitting.
-- **Database**: Store results in a relational SQLite3 database, enabling efficient management of large modeling results.
-- **Scaling Up**: Guidance on scaling up your scientific workflow from small to large datasets.
-
-__Contents__
-
-This overview is split into the following sections:
-
-- **Data**: Load the 1D Gaussian data from disk to illustrate the scientific workflow.
-- **On The Fly**: Display intermediate results during model fitting for instant feedback.
-- **Hard Disk Output**: Enable persistent saving of search results with customizable output structure.
-- **Visualization**: Generate model-specific visualizations saved to disk during fitting.
-- **Loading Results**: Use the Aggregator API to load and inspect results from hard disk.
-- **Result Customization**: Extend the Result class with custom properties specific to the model-fitting problem.
-- **Model Composition**: Construct diverse models with parameter assignments and complex hierarchies.
-- **Searches**: Select and customize non-linear search methods appropriate for the problem.
-- **Configs**: Use configuration files to define default model priors and search settings.
-- **Database**: Store and query results in a SQLite3 relational database.
-- **Scaling Up**: Guidance on expanding workflows from small to large datasets.
-- **Wrap Up**: Summary of scientific workflow features in PyAutoFit.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import numpy as np
-from typing import Optional
-from os import path
-
-import autofit as af
-
-"""
-__Data__
-
-To illustrate a few aspects of the scientific workflow, we'll fit a 1D Gaussian profile to data, which
-we load from hard-disk.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-
-"""
-__On The Fly__
-
-The on-the-fly feature described below is not implemented yet, we are working on it currently.
-The best way to get on-the-fly output is to output to hard-disk, which is described in the next section.
-This feature is fully implemented and provides on-the-fly output of results to hard-disk.
-
-When a model fit is running, information about the fit is displayed at user-specified intervals.
-
-The frequency of this on-the-fly output is controlled by a search's `iterations_per_full_update` parameter, which
-specifies how often this information is output. The example code below outputs on-the-fly information every 1000
-iterations:
-"""
-search = af.DynestyStatic(iterations_per_full_update=1000)
-
-"""
-In a Jupyter notebook, the default behavior is for this information to appear in the cell being run and to include:
-
-- Text displaying the maximum likelihood model inferred so far and related information.
-- A visual showing how the search has sampled parameter space so far, providing intuition on how the search is
-performing.
-
-Here is an image of how this looks:
-
-
-
-The most valuable on-the-fly output is often specific to the model and dataset you are fitting. For instance, it
-might be a ``matplotlib`` subplot showing the maximum likelihood model's fit to the dataset, complete with residuals
-and other diagnostic information.
-
-The on-the-fly output can be fully customized by extending the ``on_the_fly_output`` method of the ``Analysis``
-class being used to fit the model.
-
-The example below shows how this is done for the simple case of fitting a 1D Gaussian profile:
-"""
-
-
-class Analysis(af.Analysis):
- def __init__(self, data: np.ndarray, noise_map: np.ndarray):
- """
- Example Analysis class illustrating how to customize the on-the-fly output of a model-fit.
- """
- super().__init__()
-
- self.data = data
- self.noise_map = noise_map
-
- def on_the_fly_output(self, instance):
- """
- During a model-fit, the `on_the_fly_output` method is called throughout the non-linear search.
-
- The `instance` passed into the method is maximum log likelihood solution obtained by the model-fit so far and it can be
- used to provide on-the-fly output showing how the model-fit is going.
- """
- xvalues = np.arange(self.data.shape[0])
-
- model_data = instance.model_data_from(xvalues=xvalues)
-
- """
- The visualizer now outputs images of the best-fit results to hard-disk (checkout `visualizer.py`).
- """
- import matplotlib.pyplot as plt
-
- plt.errorbar(
- x=xvalues,
- y=self.data,
- yerr=self.noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
- )
- plt.plot(xvalues, model_data, color="r")
- plt.title("Maximum Likelihood Fit")
- plt.xlabel("x value of profile")
- plt.ylabel("Profile Normalization")
- plt.show() # By using `plt.show()` the plot will be displayed in the Jupyter notebook.
-
-
-"""
-Here's how the visuals appear in a Jupyter Notebook:
-
-
-
-In the early stages of setting up a scientific workflow, on-the-fly output is invaluable. It provides immediate
-feedback on how your model fitting is performing, which is often crucial at the beginning of a project when things
-might not be going well. It also encourages you to prioritize visualizing your fit and diagnosing whether the process
-is working correctly.
-
-We highly recommend users starting a new model-fitting problem begin by setting up on-the-fly output!
-
-__Hard Disk Output__
-
-By default, a non-linear search does not save its results to the hard disk; the results can only be inspected in
-a Jupyter Notebook or Python script via the returned `result`.
-
-However, you can enable the output of non-linear search results to the hard disk by specifying
-the `name` and/or `path_prefix` attributes. These attributes determine how files are named and where results
-are saved on your hard disk.
-
-Benefits of saving results to the hard disk include:
-
-- More efficient inspection of results for multiple datasets compared to using a Jupyter Notebook.
-- Results are saved on-the-fly, allowing you to check the progress of a fit midway.
-- Additional information about a fit, such as visualizations, can be saved (see below).
-- Unfinished runs can be resumed from where they left off if they are terminated.
-- On high-performance supercomputers, results often need to be saved in this manner.
-
-Here's how to enable the output of results to the hard disk:
-"""
-search = af.Emcee(path_prefix=path.join("folder_0", "folder_1"), name="my_search_name")
-
-"""
-The screenshot below shows the output folder where all output is enabled:
-
-.. image:: https://raw.githubusercontent.com/Jammy2211/PyAutoFit/main/docs/overview/image/output_example.png
- :width: 400
- :alt: Alternative text
-
-Let's break down the output folder generated by **PyAutoFit**:
-
-- **Unique Identifier**: Results are saved in a folder named with a unique identifier composed of random characters.
- This identifier is automatically generated based on the specific model fit. For scientific workflows involving
- numerous model fits, this ensures that each fit is uniquely identified without requiring manual updates to output paths.
-
-- **Info Files**: These files contain valuable information about the fit. For instance, `model.info` provides the
- complete model composition used in the fit, while `search.summary` details how long the search has been running
- and other relevant search-specific information.
-
-- **Files Folder**: Within the output folder, the `files` directory contains detailed information saved as `.json`
- files. For example, `model.json` stores the model configuration used in the fit. This enables researchers to
- revisit the results later and review how the fit was performed.
-
-**PyAutoFit** offers extensive tools for customizing hard-disk output. This includes using configuration files to
-control what information is saved, which helps manage disk space utilization. Additionally, specific `.json` files
-tailored to different models can be utilized for more detailed output.
-
-For many scientific workflows, having detailed output for each fit is crucial for thorough inspection and accurate
-interpretation of results. However, in scenarios where the volume of output data might overwhelm users or impede
-scientific study, this feature can be easily disabled by omitting the `name` or `path prefix` when initiating the search.
-
-__Visualization__
-
-When search hard-disk output is enabled in **PyAutoFit**, the visualization of model fits can also be saved directly
-to disk. This capability is crucial for many scientific workflows as it allows for quick and effective assessment of
-fit quality.
-
-To accomplish this, you can customize the `Visualizer` object of an `Analysis` class with a custom `Visualizer` class.
-This custom class is responsible for generating and saving visual representations of the model fits. By leveraging
-this approach, scientists can efficiently visualize and analyze the outcomes of model fitting processes.
-"""
-
-
-class Visualizer(af.Visualizer):
- @staticmethod
- def visualize_before_fit(
- analysis, paths: af.DirectoryPaths, model: af.AbstractPriorModel
- ):
- """
- Before a model-fit, the `visualize_before_fit` method is called to perform visualization.
-
- The function receives as input an instance of the `Analysis` class which is being used to perform the fit,
- which is used to perform the visualization (e.g. it contains the data and noise map which are plotted).
-
- This can output visualization of quantities which do not change during the model-fit, for example the
- data and noise-map.
-
- The `paths` object contains the path to the folder where the visualization should be output, which is determined
- by the non-linear search `name` and other inputs.
- """
-
- import matplotlib.pyplot as plt
-
- xvalues = np.arange(analysis.data.shape[0])
-
- plt.errorbar(
- x=xvalues,
- y=analysis.data,
- yerr=analysis.noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
- )
- plt.title("Maximum Likelihood Fit")
- plt.xlabel("x value of profile")
- plt.ylabel("Profile Normalization")
- plt.savefig(path.join(paths.image_path, f"data.png"))
- plt.clf()
-
- @staticmethod
- def visualize(analysis, paths: af.DirectoryPaths, instance, during_analysis):
- """
- During a model-fit, the `visualize` method is called throughout the non-linear search.
-
- The function receives as input an instance of the `Analysis` class which is being used to perform the fit,
- which is used to perform the visualization (e.g. it generates the model data which is plotted).
-
- The `instance` passed into the visualize method is maximum log likelihood solution obtained by the model-fit
- so far and it can be used to provide on-the-fly images showing how the model-fit is going.
-
- The `paths` object contains the path to the folder where the visualization should be output, which is determined
- by the non-linear search `name` and other inputs.
- """
- xvalues = np.arange(analysis.data.shape[0])
-
- model_data = instance.model_data_from(xvalues=xvalues)
- residual_map = analysis.data - model_data
-
- """
- The visualizer now outputs images of the best-fit results to hard-disk (checkout `visualizer.py`).
- """
- import matplotlib.pyplot as plt
-
- plt.errorbar(
- x=xvalues,
- y=analysis.data,
- yerr=analysis.noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
- )
- plt.plot(xvalues, model_data, color="r")
- plt.title("Maximum Likelihood Fit")
- plt.xlabel("x value of profile")
- plt.ylabel("Profile Normalization")
- plt.savefig(path.join(paths.image_path, f"model_fit.png"))
- plt.clf()
-
- plt.errorbar(
- x=xvalues,
- y=residual_map,
- yerr=analysis.noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
- )
- plt.title("Residuals of Maximum Likelihood Fit")
- plt.xlabel("x value of profile")
- plt.ylabel("Residual")
- plt.savefig(path.join(paths.image_path, f"model_fit.png"))
- plt.clf()
-
-
-"""
-The ``Analysis`` class is defined following the same API as before, but now with its `Visualizer` class attribute
-overwritten with the ``Visualizer`` class above.
-"""
-
-
-class Analysis(af.Analysis):
- """
- This over-write means the `Visualizer` class is used for visualization throughout the model-fit.
-
- This `VisualizerExample` object is in the `autofit.example.visualize` module and is used to customize the
- plots output during the model-fit.
-
- It has been extended with visualize methods that output visuals specific to the fitting of `1D` data.
- """
-
- Visualizer = Visualizer
-
- def __init__(self, data, noise_map):
- """
- An Analysis class which illustrates visualization.
- """
- super().__init__()
-
- self.data = data
- self.noise_map = noise_map
-
- def log_likelihood_function(self, instance):
- """
- The `log_likelihood_function` is identical to the example above
- """
- xvalues = np.arange(self.data.shape[0])
-
- model_data = instance.model_data_from(xvalues=xvalues)
- residual_map = self.data - model_data
- chi_squared_map = (residual_map / self.noise_map) ** 2.0
- chi_squared = sum(chi_squared_map)
- noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
- log_likelihood = -0.5 * (chi_squared + noise_normalization)
-
- return log_likelihood
-
-
-"""
-Visualization of the results of the non-linear search, for example the "Probability Density
-Function", are also automatically output during the model-fit on the fly.
-
-We now perform a quick fit, outputting the results to the hard disk and visualizing the model-fit,
-so you can see how the results are output and the visualizations produced.
-"""
-analysis = Analysis(data=data, noise_map=noise_map)
-
-model = af.Model(af.ex.Gaussian)
-
-search = af.DynestyStatic(
- path_prefix=path.join("result_folder"), name="overview_2_scientific_workflow"
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-"""
-__Loading Results__
-
-In your scientific workflow, you'll likely conduct numerous model fits, each generating outputs stored in individual
-folders on your hard disk.
-
-To efficiently work with these results in Python scripts or Jupyter notebooks, **PyAutoFit** provides
-the `aggregator` API. This tool simplifies the process of loading results from hard disk into Python variables.
-By pointing the aggregator at the folder containing your results, it automatically loads all relevant information
-from each model fit.
-
-This capability streamlines the workflow by enabling easy manipulation and inspection of model-fit results directly
-within your Python environment. It's particularly useful for managing and analyzing large-scale studies where
-handling multiple model fits and their associated outputs is essential.
-"""
-from autofit.aggregator.aggregator import Aggregator
-
-agg = Aggregator.from_directory(
- directory=path.join("result_folder"),
-)
-
-"""
-The ``values`` method is used to specify the information that is loaded from the hard-disk, for example the
-``samples`` of the model-fit.
-
-The for loop below iterates over all results in the folder passed to the aggregator above.
-"""
-for samples in agg.values("samples"):
- print(samples.parameter_lists[0])
-
-"""
-Result loading uses Python generators to ensure that memory use is minimized, meaning that even when loading
-thousands of results from hard-disk the memory use of your machine is not exceeded.
-
-The `result cookbook `_ gives a full run-through of
-the tools that allow results to be loaded and inspected.
-
-__Result Customization__
-
-An effective scientific workflow ensures that this object contains all information a user needs to quickly inspect
-the quality of a model-fit and undertake scientific interpretation.
-
-The result can be can be customized to include additional information about the model-fit that is specific to your
-model-fitting problem.
-
-For example, for fitting 1D profiles, the ``Result`` could include the maximum log likelihood model 1D data,
-which would enable the following code to be used after the model-fit:
-
-print(result.max_log_likelihood_model_data_1d)
-
-To do this we use the custom result API, where we first define a custom ``Result`` class which includes the
-property ``max_log_likelihood_model_data_1d``:
-"""
-
-
-class ResultExample(af.Result):
- @property
- def max_log_likelihood_model_data_1d(self) -> np.ndarray:
- """
- Returns the maximum log likelihood model's 1D model data.
-
- This is an example of how we can pass the `Analysis` class a custom `Result` object and extend this result
- object with new properties that are specific to the model-fit we are performing.
- """
- xvalues = np.arange(self.analysis.data.shape[0])
-
- return self.instance.model_data_from(xvalues=xvalues)
-
-
-"""
-The custom result has access to the analysis class, meaning that we can use any of its methods or properties to
-compute custom result properties.
-
-To make it so that the ``ResultExample`` object above is returned by the search we overwrite the ``Result`` class attribute
-of the ``Analysis`` and define a ``make_result`` object describing what we want it to contain:
-"""
-
-
-class Analysis(af.Analysis):
- """
- This overwrite means the `ResultExample` class is returned after the model-fit.
- """
-
- Result = ResultExample
-
- def __init__(self, data, noise_map):
- """
- An Analysis class which illustrates custom results.
- """
- super().__init__()
-
- self.data = data
- self.noise_map = noise_map
-
- def log_likelihood_function(self, instance):
- """
- The `log_likelihood_function` is identical to the example above
- """
- xvalues = np.arange(self.data.shape[0])
-
- model_data = instance.model_data_from(xvalues=xvalues)
- residual_map = self.data - model_data
- chi_squared_map = (residual_map / self.noise_map) ** 2.0
- chi_squared = sum(chi_squared_map)
- noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
- log_likelihood = -0.5 * (chi_squared + noise_normalization)
-
- return log_likelihood
-
- def make_result(
- self,
- samples_summary: af.SamplesSummary,
- paths: af.AbstractPaths,
- samples: Optional[af.SamplesPDF] = None,
- search_internal: Optional[object] = None,
- analysis: Optional[object] = None,
- ) -> Result:
- """
- Returns the `Result` of the non-linear search after it is completed.
-
- The result type is defined as a class variable in the `Analysis` class (see top of code under the python code
- `class Analysis(af.Analysis)`.
-
- The result can be manually overwritten by a user to return a user-defined result object, which can be extended
- with additional methods and attribute specific to the model-fit.
-
- This example class does example this, whereby the analysis result has been overwritten with the `ResultExample`
- class, which contains a property `max_log_likelihood_model_data_1d` that returns the model data of the
- best-fit model. This API means you can customize your result object to include whatever attributes you want
- and therefore make a result object specific to your model-fit and model-fitting problem.
-
- The `Result` object you return can be customized to include:
-
- - The samples summary, which contains the maximum log likelihood instance and median PDF model.
-
- - The paths of the search, which are used for loading the samples and search internal below when a search
- is resumed.
-
- - The samples of the non-linear search (e.g. MCMC chains) also stored in `samples.csv`.
-
- - The non-linear search used for the fit in its internal representation, which is used for resuming a search
- and making bespoke visualization using the search's internal results.
-
- - The analysis used to fit the model (default disabled to save memory, but option may be useful for certain
- projects).
-
- Parameters
- ----------
- samples_summary
- The summary of the samples of the non-linear search, which include the maximum log likelihood instance and
- median PDF model.
- paths
- An object describing the paths for saving data (e.g. hard-disk directories or entries in sqlite database).
- samples
- The samples of the non-linear search, for example the chains of an MCMC run.
- search_internal
- The internal representation of the non-linear search used to perform the model-fit.
- analysis
- The analysis used to fit the model.
-
- Returns
- -------
- Result
- The result of the non-linear search, which is defined as a class variable in the `Analysis` class.
- """
- return self.Result(
- samples_summary=samples_summary,
- paths=paths,
- samples=samples,
- search_internal=search_internal,
- analysis=self,
- )
-
-
-"""
-By repeating the model-fit above, the `Result` object returned by the search will be an instance of the `ResultExample`
-class, which includes the property `max_log_likelihood_model_data_1d`.
-"""
-analysis = Analysis(data=data, noise_map=noise_map)
-
-model = af.Model(af.ex.Gaussian)
-
-search = af.DynestyStatic(
- path_prefix=path.join("output", "result_folder"),
- name="overview_2_scientific_workflow",
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-print(result.max_log_likelihood_model_data_1d)
-
-"""
-Result customization has full support for **latent variables**, which are parameters that are not sampled by the non-linear
-search but are computed from the sampled parameters.
-
-They are often integral to assessing and interpreting the results of a model-fit, as they present information
-on the model in a different way to the sampled parameters.
-
-The `result cookbook `_ gives a full run-through of
-all the different ways the result can be customized.
-
-__Model Composition__
-
-In many scientific workflows, there's often a need to construct and fit a variety of different models. This
-could range from making minor adjustments to a model's parameters to handling complex models with thousands of parameters and multiple components.
-
-For simpler scenarios, adjustments might include:
-
-- **Parameter Assignment**: Setting specific values for certain parameters or linking parameters together so they share the same value.
-- **Parameter Assertions**: Imposing constraints on model parameters, such as requiring one parameter to be greater than another.
-- **Model Arithmetic**: Defining relationships between parameters using arithmetic operations, such as defining a
- linear relationship like `y = mx + c`, where `m` and `c` are model parameters.
-
-In more intricate cases, models might involve numerous parameters and complex compositions of multiple model components.
-
-**PyAutoFit** offers a sophisticated model composition API designed to handle these complexities. It provides
-tools for constructing elaborate models using lists of Python classes, NumPy arrays and hierarchical structures of Python classes.
-
-For a detailed exploration of these capabilities, you can refer to
-the `model cookbook `_, which provides comprehensive
-guidance on using the model composition API. This resource covers everything from basic parameter assignments to
-constructing complex models with hierarchical structures.
-
-__Searches__
-
-Different model-fitting problems often require different approaches to fitting the model effectively.
-
-The choice of the most suitable search method depends on several factors:
-
-- **Model Dimensions**: How many parameters constitute the model and its non-linear parameter space?
-- **Model Complexity**: Different models exhibit varying degrees of parameter degeneracy, which necessitates different
- non-linear search techniques.
-- **Run Times**: How efficiently can the likelihood function be evaluated and the model-fit performed?
-- **Gradients**: If your likelihood function is differentiable, leveraging JAX and using a search that exploits
- gradient information can be advantageous.
-
-**PyAutoFit** provides support for a wide range of non-linear searches, ensuring that users can select the method
-best suited to their specific problem.
-
-During the initial stages of setting up your scientific workflow, it's beneficial to experiment with different
-searches. This process helps identify which methods reliably infer maximum likelihood fits to the data and assess
-their efficiency in terms of computational time.
-
-For a comprehensive exploration of available search methods and customization options, refer to
-the `search cookbook `_. This resource covers
-detailed guides on all non-linear searches supported by PyAutoFit and provides insights into how to tailor them to your
-needs.
-
-There are currently no documentation guiding reads on what search might be appropriate for their problem and how to
-profile and experiment with different methods. Writing such documentation is on the to do list and will appear
-in the future. However, you can make progress now simply using visuals output by PyAutoFit and the ``search.summary` file.
-
-__Configs__
-
-As you refine your scientific workflow, you'll often find yourself repeatedly setting up models with identical priors
-and using the same non-linear search configurations. This repetition can result in lengthy Python scripts with
-redundant inputs.
-
-To streamline this process, configuration files can be utilized to define default values. This approach eliminates
-the need to specify identical prior inputs and search settings in every script, leading to more concise and
-readable Python code. Moreover, it reduces the cognitive load associated with performing model-fitting tasks.
-
-For a comprehensive guide on setting up and utilizing configuration files effectively, refer
-to the `configs cookbook `_. This resource provides
-detailed instructions on configuring and optimizing your PyAutoFit workflow through the use of configuration files.
-
-__Database__
-
-By default, model-fitting results are written to folders on hard-disk, which is straightforward for navigating and
-manual inspection. However, this approach becomes impractical for large datasets or extensive scientific workflows,
-where manually checking each result can be time-consuming.
-
-To address this challenge, all results can be stored in an sqlite3 relational database. This enables loading results
-directly into Jupyter notebooks or Python scripts for inspection, analysis, and interpretation. The database
-supports advanced querying capabilities, allowing users to retrieve specific model-fits based on criteria such
-as the fitted model or dataset.
-
-For a comprehensive guide on using the database functionality within PyAutoFit, refer to
-the `database cookbook `. This resource
-provides detailed instructions on leveraging the database to manage and analyze model-fitting results efficiently.
-
-__Scaling Up__
-
-Regardless of your final scientific objective, it's crucial to consider scalability in your scientific workflow and
-ensure it remains flexible to accommodate varying scales of complexity.
-
-Initially, scientific studies often begin with a small number of datasets (e.g., tens of datasets). During this phase,
-researchers iteratively refine their models and gain insights through trial and error. This involves fitting numerous
-models to datasets and manually inspecting results to evaluate model performance. A flexible workflow is essential
-here, allowing rapid iteration and outputting results in a format that facilitates quick inspection and interpretation.
-
-As the study progresses, researchers may scale up to larger datasets (e.g., thousands of datasets). Manual inspection
-of individual results becomes impractical, necessitating a more automated approach to model fitting and interpretation.
-Additionally, analyses may transition to high-performance computing environments, requiring output formats suitable for
-these setups.
-
-**PyAutoFit** is designed to enable the development of effective scientific workflows for both small and large datasets.
-
-__Wrap Up__
-
-This overview has provided a comprehensive guide to the key features of **PyAutoFit** that support the development of
-effective scientific workflows. By leveraging these tools, researchers can tailor their workflows to specific problems,
-streamline model fitting, and gain valuable insights into their scientific studies.
-
-The final aspect of core functionality, described in the next overview, is the wide variety of statistical
-inference methods available in **PyAutoFit**. These methods include graphical models, hierarchical models,
-Bayesian model comparison and many more.
-"""
+"""
+Overview: Scientific Workflow
+=============================
+
+A scientific workflow comprises the tasks you perform to conduct a scientific study. This includes fitting models to
+datasets, interpreting the results, and gaining insights into your scientific problem.
+
+Different problems require different scientific workflows, depending on factors such as model complexity, dataset size,
+and computational run times. For example, some problems involve fitting a single dataset with many models to gain
+scientific insights, while others involve fitting thousands of datasets with a single model for large-scale studies.
+
+The **PyAutoFit** API is flexible, customizable, and extensible, enabling users to develop scientific workflows
+tailored to their specific problems.
+
+This overview covers the key features of **PyAutoFit** that support the development of effective scientific workflows:
+
+- **On The Fly**: Display results immediately (e.g., in Jupyter notebooks) to provide instant feedback for adapting your workflow.
+- **Hard Disk Output**: Output results to hard disk with high customization, allowing quick and detailed inspection of fits to many datasets.
+- **Visualization**: Generate model-specific visualizations to create custom plots that streamline result inspection.
+- **Loading Results**: Load results from the hard disk to inspect and interpret the outcomes of a model fit.
+- **Result Customization**: Customize the returned results to simplify scientific interpretation.
+- **Model Composition**: Extensible model composition makes it easy to fit many models with different parameterizations and assumptions.
+- **Searches**: Support for various non-linear searches (e.g., nested sampling, MCMC), including gradient based fitting using JAX, to find the right method for your problem.
+- **Configs**: Configuration files that set default model, fitting, and visualization behaviors, streamlining model fitting.
+- **Database**: Store results in a relational SQLite3 database, enabling efficient management of large modeling results.
+- **Scaling Up**: Guidance on scaling up your scientific workflow from small to large datasets.
+
+__Contents__
+
+This overview is split into the following sections:
+
+- **Data**: Load the 1D Gaussian data from disk to illustrate the scientific workflow.
+- **On The Fly**: Display intermediate results during model fitting for instant feedback.
+- **Hard Disk Output**: Enable persistent saving of search results with customizable output structure.
+- **Visualization**: Generate model-specific visualizations saved to disk during fitting.
+- **Loading Results**: Use the Aggregator API to load and inspect results from hard disk.
+- **Result Customization**: Extend the Result class with custom properties specific to the model-fitting problem.
+- **Model Composition**: Construct diverse models with parameter assignments and complex hierarchies.
+- **Searches**: Select and customize non-linear search methods appropriate for the problem.
+- **Configs**: Use configuration files to define default model priors and search settings.
+- **Database**: Store and query results in a SQLite3 relational database.
+- **Scaling Up**: Guidance on expanding workflows from small to large datasets.
+- **Wrap Up**: Summary of scientific workflow features in PyAutoFit.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import numpy as np
+from typing import Optional
+from os import path
+
+import autofit as af
+
+"""
+__Data__
+
+To illustrate a few aspects of the scientific workflow, we'll fit a 1D Gaussian profile to data, which
+we load from hard-disk.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+
+"""
+__On The Fly__
+
+The on-the-fly feature described below is not implemented yet, we are working on it currently.
+The best way to get on-the-fly output is to output to hard-disk, which is described in the next section.
+This feature is fully implemented and provides on-the-fly output of results to hard-disk.
+
+When a model fit is running, information about the fit is displayed at user-specified intervals.
+
+The frequency of this on-the-fly output is controlled by a search's `iterations_per_full_update` parameter, which
+specifies how often this information is output. The example code below outputs on-the-fly information every 1000
+iterations:
+"""
+search = af.DynestyStatic(iterations_per_full_update=1000)
+
+"""
+In a Jupyter notebook, the default behavior is for this information to appear in the cell being run and to include:
+
+- Text displaying the maximum likelihood model inferred so far and related information.
+- A visual showing how the search has sampled parameter space so far, providing intuition on how the search is
+performing.
+
+Here is an image of how this looks:
+
+
+
+The most valuable on-the-fly output is often specific to the model and dataset you are fitting. For instance, it
+might be a ``matplotlib`` subplot showing the maximum likelihood model's fit to the dataset, complete with residuals
+and other diagnostic information.
+
+The on-the-fly output can be fully customized by extending the ``on_the_fly_output`` method of the ``Analysis``
+class being used to fit the model.
+
+The example below shows how this is done for the simple case of fitting a 1D Gaussian profile:
+"""
+
+
+class Analysis(af.Analysis):
+ def __init__(self, data: np.ndarray, noise_map: np.ndarray):
+ """
+ Example Analysis class illustrating how to customize the on-the-fly output of a model-fit.
+ """
+ super().__init__()
+
+ self.data = data
+ self.noise_map = noise_map
+
+ def on_the_fly_output(self, instance):
+ """
+ During a model-fit, the `on_the_fly_output` method is called throughout the non-linear search.
+
+ The `instance` passed into the method is maximum log likelihood solution obtained by the model-fit so far and it can be
+ used to provide on-the-fly output showing how the model-fit is going.
+ """
+ xvalues = np.arange(self.data.shape[0])
+
+ model_data = instance.model_data_from(xvalues=xvalues)
+
+ """
+ The visualizer now outputs images of the best-fit results to hard-disk (checkout `visualizer.py`).
+ """
+ import matplotlib.pyplot as plt
+
+ plt.errorbar(
+ x=xvalues,
+ y=self.data,
+ yerr=self.noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+ )
+ plt.plot(xvalues, model_data, color="r")
+ plt.title("Maximum Likelihood Fit")
+ plt.xlabel("x value of profile")
+ plt.ylabel("Profile Normalization")
+ plt.show() # By using `plt.show()` the plot will be displayed in the Jupyter notebook.
+
+
+"""
+Here's how the visuals appear in a Jupyter Notebook:
+
+
+
+In the early stages of setting up a scientific workflow, on-the-fly output is invaluable. It provides immediate
+feedback on how your model fitting is performing, which is often crucial at the beginning of a project when things
+might not be going well. It also encourages you to prioritize visualizing your fit and diagnosing whether the process
+is working correctly.
+
+We highly recommend users starting a new model-fitting problem begin by setting up on-the-fly output!
+
+__Hard Disk Output__
+
+By default, a non-linear search does not save its results to the hard disk; the results can only be inspected in
+a Jupyter Notebook or Python script via the returned `result`.
+
+However, you can enable the output of non-linear search results to the hard disk by specifying
+the `name` and/or `path_prefix` attributes. These attributes determine how files are named and where results
+are saved on your hard disk.
+
+Benefits of saving results to the hard disk include:
+
+- More efficient inspection of results for multiple datasets compared to using a Jupyter Notebook.
+- Results are saved on-the-fly, allowing you to check the progress of a fit midway.
+- Additional information about a fit, such as visualizations, can be saved (see below).
+- Unfinished runs can be resumed from where they left off if they are terminated.
+- On high-performance supercomputers, results often need to be saved in this manner.
+
+Here's how to enable the output of results to the hard disk:
+"""
+search = af.Emcee(path_prefix=path.join("folder_0", "folder_1"), name="my_search_name")
+
+"""
+The screenshot below shows the output folder where all output is enabled:
+
+.. image:: https://raw.githubusercontent.com/Jammy2211/PyAutoFit/main/docs/overview/image/output_example.png
+ :width: 400
+ :alt: Alternative text
+
+Let's break down the output folder generated by **PyAutoFit**:
+
+- **Unique Identifier**: Results are saved in a folder named with a unique identifier composed of random characters.
+ This identifier is automatically generated based on the specific model fit. For scientific workflows involving
+ numerous model fits, this ensures that each fit is uniquely identified without requiring manual updates to output paths.
+
+- **Info Files**: These files contain valuable information about the fit. For instance, `model.info` provides the
+ complete model composition used in the fit, while `search.summary` details how long the search has been running
+ and other relevant search-specific information.
+
+- **Files Folder**: Within the output folder, the `files` directory contains detailed information saved as `.json`
+ files. For example, `model.json` stores the model configuration used in the fit. This enables researchers to
+ revisit the results later and review how the fit was performed.
+
+**PyAutoFit** offers extensive tools for customizing hard-disk output. This includes using configuration files to
+control what information is saved, which helps manage disk space utilization. Additionally, specific `.json` files
+tailored to different models can be utilized for more detailed output.
+
+For many scientific workflows, having detailed output for each fit is crucial for thorough inspection and accurate
+interpretation of results. However, in scenarios where the volume of output data might overwhelm users or impede
+scientific study, this feature can be easily disabled by omitting the `name` or `path prefix` when initiating the search.
+
+__Visualization__
+
+When search hard-disk output is enabled in **PyAutoFit**, the visualization of model fits can also be saved directly
+to disk. This capability is crucial for many scientific workflows as it allows for quick and effective assessment of
+fit quality.
+
+To accomplish this, you can customize the `Visualizer` object of an `Analysis` class with a custom `Visualizer` class.
+This custom class is responsible for generating and saving visual representations of the model fits. By leveraging
+this approach, scientists can efficiently visualize and analyze the outcomes of model fitting processes.
+"""
+
+
+class Visualizer(af.Visualizer):
+ @staticmethod
+ def visualize_before_fit(
+ analysis, paths: af.DirectoryPaths, model: af.AbstractPriorModel
+ ):
+ """
+ Before a model-fit, the `visualize_before_fit` method is called to perform visualization.
+
+ The function receives as input an instance of the `Analysis` class which is being used to perform the fit,
+ which is used to perform the visualization (e.g. it contains the data and noise map which are plotted).
+
+ This can output visualization of quantities which do not change during the model-fit, for example the
+ data and noise-map.
+
+ The `paths` object contains the path to the folder where the visualization should be output, which is determined
+ by the non-linear search `name` and other inputs.
+ """
+
+ import matplotlib.pyplot as plt
+
+ xvalues = np.arange(analysis.data.shape[0])
+
+ plt.errorbar(
+ x=xvalues,
+ y=analysis.data,
+ yerr=analysis.noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+ )
+ plt.title("Maximum Likelihood Fit")
+ plt.xlabel("x value of profile")
+ plt.ylabel("Profile Normalization")
+ plt.savefig(path.join(paths.image_path, f"data.png"))
+ plt.clf()
+
+ @staticmethod
+ def visualize(analysis, paths: af.DirectoryPaths, instance, during_analysis):
+ """
+ During a model-fit, the `visualize` method is called throughout the non-linear search.
+
+ The function receives as input an instance of the `Analysis` class which is being used to perform the fit,
+ which is used to perform the visualization (e.g. it generates the model data which is plotted).
+
+ The `instance` passed into the visualize method is maximum log likelihood solution obtained by the model-fit
+ so far and it can be used to provide on-the-fly images showing how the model-fit is going.
+
+ The `paths` object contains the path to the folder where the visualization should be output, which is determined
+ by the non-linear search `name` and other inputs.
+ """
+ xvalues = np.arange(analysis.data.shape[0])
+
+ model_data = instance.model_data_from(xvalues=xvalues)
+ residual_map = analysis.data - model_data
+
+ """
+ The visualizer now outputs images of the best-fit results to hard-disk (checkout `visualizer.py`).
+ """
+ import matplotlib.pyplot as plt
+
+ plt.errorbar(
+ x=xvalues,
+ y=analysis.data,
+ yerr=analysis.noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+ )
+ plt.plot(xvalues, model_data, color="r")
+ plt.title("Maximum Likelihood Fit")
+ plt.xlabel("x value of profile")
+ plt.ylabel("Profile Normalization")
+ plt.savefig(path.join(paths.image_path, f"model_fit.png"))
+ plt.clf()
+
+ plt.errorbar(
+ x=xvalues,
+ y=residual_map,
+ yerr=analysis.noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+ )
+ plt.title("Residuals of Maximum Likelihood Fit")
+ plt.xlabel("x value of profile")
+ plt.ylabel("Residual")
+ plt.savefig(path.join(paths.image_path, f"model_fit.png"))
+ plt.clf()
+
+
+"""
+The ``Analysis`` class is defined following the same API as before, but now with its `Visualizer` class attribute
+overwritten with the ``Visualizer`` class above.
+"""
+
+
+class Analysis(af.Analysis):
+ """
+ This over-write means the `Visualizer` class is used for visualization throughout the model-fit.
+
+ This `VisualizerExample` object is in the `autofit.example.visualize` module and is used to customize the
+ plots output during the model-fit.
+
+ It has been extended with visualize methods that output visuals specific to the fitting of `1D` data.
+ """
+
+ Visualizer = Visualizer
+
+ def __init__(self, data, noise_map):
+ """
+ An Analysis class which illustrates visualization.
+ """
+ super().__init__()
+
+ self.data = data
+ self.noise_map = noise_map
+
+ def log_likelihood_function(self, instance):
+ """
+ The `log_likelihood_function` is identical to the example above
+ """
+ xvalues = np.arange(self.data.shape[0])
+
+ model_data = instance.model_data_from(xvalues=xvalues)
+ residual_map = self.data - model_data
+ chi_squared_map = (residual_map / self.noise_map) ** 2.0
+ chi_squared = sum(chi_squared_map)
+ noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
+ log_likelihood = -0.5 * (chi_squared + noise_normalization)
+
+ return log_likelihood
+
+
+"""
+Visualization of the results of the non-linear search, for example the "Probability Density
+Function", are also automatically output during the model-fit on the fly.
+
+We now perform a quick fit, outputting the results to the hard disk and visualizing the model-fit,
+so you can see how the results are output and the visualizations produced.
+"""
+analysis = Analysis(data=data, noise_map=noise_map)
+
+model = af.Model(af.ex.Gaussian)
+
+search = af.DynestyStatic(
+ path_prefix=path.join("result_folder"), name="overview_2_scientific_workflow"
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+"""
+__Loading Results__
+
+In your scientific workflow, you'll likely conduct numerous model fits, each generating outputs stored in individual
+folders on your hard disk.
+
+To efficiently work with these results in Python scripts or Jupyter notebooks, **PyAutoFit** provides
+the `aggregator` API. This tool simplifies the process of loading results from hard disk into Python variables.
+By pointing the aggregator at the folder containing your results, it automatically loads all relevant information
+from each model fit.
+
+This capability streamlines the workflow by enabling easy manipulation and inspection of model-fit results directly
+within your Python environment. It's particularly useful for managing and analyzing large-scale studies where
+handling multiple model fits and their associated outputs is essential.
+"""
+from autofit.aggregator.aggregator import Aggregator
+
+agg = Aggregator.from_directory(
+ directory=path.join("result_folder"),
+)
+
+"""
+The ``values`` method is used to specify the information that is loaded from the hard-disk, for example the
+``samples`` of the model-fit.
+
+The for loop below iterates over all results in the folder passed to the aggregator above.
+"""
+for samples in agg.values("samples"):
+ print(samples.parameter_lists[0])
+
+"""
+Result loading uses Python generators to ensure that memory use is minimized, meaning that even when loading
+thousands of results from hard-disk the memory use of your machine is not exceeded.
+
+The `result cookbook `_ gives a full run-through of
+the tools that allow results to be loaded and inspected.
+
+__Result Customization__
+
+An effective scientific workflow ensures that this object contains all information a user needs to quickly inspect
+the quality of a model-fit and undertake scientific interpretation.
+
+The result can be can be customized to include additional information about the model-fit that is specific to your
+model-fitting problem.
+
+For example, for fitting 1D profiles, the ``Result`` could include the maximum log likelihood model 1D data,
+which would enable the following code to be used after the model-fit:
+
+print(result.max_log_likelihood_model_data_1d)
+
+To do this we use the custom result API, where we first define a custom ``Result`` class which includes the
+property ``max_log_likelihood_model_data_1d``:
+"""
+
+
+class ResultExample(af.Result):
+ @property
+ def max_log_likelihood_model_data_1d(self) -> np.ndarray:
+ """
+ Returns the maximum log likelihood model's 1D model data.
+
+ This is an example of how we can pass the `Analysis` class a custom `Result` object and extend this result
+ object with new properties that are specific to the model-fit we are performing.
+ """
+ xvalues = np.arange(self.analysis.data.shape[0])
+
+ return self.instance.model_data_from(xvalues=xvalues)
+
+
+"""
+The custom result has access to the analysis class, meaning that we can use any of its methods or properties to
+compute custom result properties.
+
+To make it so that the ``ResultExample`` object above is returned by the search we overwrite the ``Result`` class attribute
+of the ``Analysis`` and define a ``make_result`` object describing what we want it to contain:
+"""
+
+
+class Analysis(af.Analysis):
+ """
+ This overwrite means the `ResultExample` class is returned after the model-fit.
+ """
+
+ Result = ResultExample
+
+ def __init__(self, data, noise_map):
+ """
+ An Analysis class which illustrates custom results.
+ """
+ super().__init__()
+
+ self.data = data
+ self.noise_map = noise_map
+
+ def log_likelihood_function(self, instance):
+ """
+ The `log_likelihood_function` is identical to the example above
+ """
+ xvalues = np.arange(self.data.shape[0])
+
+ model_data = instance.model_data_from(xvalues=xvalues)
+ residual_map = self.data - model_data
+ chi_squared_map = (residual_map / self.noise_map) ** 2.0
+ chi_squared = sum(chi_squared_map)
+ noise_normalization = np.sum(np.log(2 * np.pi * noise_map**2.0))
+ log_likelihood = -0.5 * (chi_squared + noise_normalization)
+
+ return log_likelihood
+
+ def make_result(
+ self,
+ samples_summary: af.SamplesSummary,
+ paths: af.AbstractPaths,
+ samples: Optional[af.SamplesPDF] = None,
+ search_internal: Optional[object] = None,
+ analysis: Optional[object] = None,
+ ) -> Result:
+ """
+ Returns the `Result` of the non-linear search after it is completed.
+
+ The result type is defined as a class variable in the `Analysis` class (see top of code under the python code
+ `class Analysis(af.Analysis)`.
+
+ The result can be manually overwritten by a user to return a user-defined result object, which can be extended
+ with additional methods and attribute specific to the model-fit.
+
+ This example class does example this, whereby the analysis result has been overwritten with the `ResultExample`
+ class, which contains a property `max_log_likelihood_model_data_1d` that returns the model data of the
+ best-fit model. This API means you can customize your result object to include whatever attributes you want
+ and therefore make a result object specific to your model-fit and model-fitting problem.
+
+ The `Result` object you return can be customized to include:
+
+ - The samples summary, which contains the maximum log likelihood instance and median PDF model.
+
+ - The paths of the search, which are used for loading the samples and search internal below when a search
+ is resumed.
+
+ - The samples of the non-linear search (e.g. MCMC chains) also stored in `samples.csv`.
+
+ - The non-linear search used for the fit in its internal representation, which is used for resuming a search
+ and making bespoke visualization using the search's internal results.
+
+ - The analysis used to fit the model (default disabled to save memory, but option may be useful for certain
+ projects).
+
+ Parameters
+ ----------
+ samples_summary
+ The summary of the samples of the non-linear search, which include the maximum log likelihood instance and
+ median PDF model.
+ paths
+ An object describing the paths for saving data (e.g. hard-disk directories or entries in sqlite database).
+ samples
+ The samples of the non-linear search, for example the chains of an MCMC run.
+ search_internal
+ The internal representation of the non-linear search used to perform the model-fit.
+ analysis
+ The analysis used to fit the model.
+
+ Returns
+ -------
+ Result
+ The result of the non-linear search, which is defined as a class variable in the `Analysis` class.
+ """
+ return self.Result(
+ samples_summary=samples_summary,
+ paths=paths,
+ samples=samples,
+ search_internal=search_internal,
+ analysis=self,
+ )
+
+
+"""
+By repeating the model-fit above, the `Result` object returned by the search will be an instance of the `ResultExample`
+class, which includes the property `max_log_likelihood_model_data_1d`.
+"""
+analysis = Analysis(data=data, noise_map=noise_map)
+
+model = af.Model(af.ex.Gaussian)
+
+search = af.DynestyStatic(
+ path_prefix=path.join("output", "result_folder"),
+ name="overview_2_scientific_workflow",
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+print(result.max_log_likelihood_model_data_1d)
+
+"""
+Result customization has full support for **latent variables**, which are parameters that are not sampled by the non-linear
+search but are computed from the sampled parameters.
+
+They are often integral to assessing and interpreting the results of a model-fit, as they present information
+on the model in a different way to the sampled parameters.
+
+The `result cookbook `_ gives a full run-through of
+all the different ways the result can be customized.
+
+__Model Composition__
+
+In many scientific workflows, there's often a need to construct and fit a variety of different models. This
+could range from making minor adjustments to a model's parameters to handling complex models with thousands of parameters and multiple components.
+
+For simpler scenarios, adjustments might include:
+
+- **Parameter Assignment**: Setting specific values for certain parameters or linking parameters together so they share the same value.
+- **Parameter Assertions**: Imposing constraints on model parameters, such as requiring one parameter to be greater than another.
+- **Model Arithmetic**: Defining relationships between parameters using arithmetic operations, such as defining a
+ linear relationship like `y = mx + c`, where `m` and `c` are model parameters.
+
+In more intricate cases, models might involve numerous parameters and complex compositions of multiple model components.
+
+**PyAutoFit** offers a sophisticated model composition API designed to handle these complexities. It provides
+tools for constructing elaborate models using lists of Python classes, NumPy arrays and hierarchical structures of Python classes.
+
+For a detailed exploration of these capabilities, you can refer to
+the `model cookbook `_, which provides comprehensive
+guidance on using the model composition API. This resource covers everything from basic parameter assignments to
+constructing complex models with hierarchical structures.
+
+__Searches__
+
+Different model-fitting problems often require different approaches to fitting the model effectively.
+
+The choice of the most suitable search method depends on several factors:
+
+- **Model Dimensions**: How many parameters constitute the model and its non-linear parameter space?
+- **Model Complexity**: Different models exhibit varying degrees of parameter degeneracy, which necessitates different
+ non-linear search techniques.
+- **Run Times**: How efficiently can the likelihood function be evaluated and the model-fit performed?
+- **Gradients**: If your likelihood function is differentiable, leveraging JAX and using a search that exploits
+ gradient information can be advantageous.
+
+**PyAutoFit** provides support for a wide range of non-linear searches, ensuring that users can select the method
+best suited to their specific problem.
+
+During the initial stages of setting up your scientific workflow, it's beneficial to experiment with different
+searches. This process helps identify which methods reliably infer maximum likelihood fits to the data and assess
+their efficiency in terms of computational time.
+
+For a comprehensive exploration of available search methods and customization options, refer to
+the `search cookbook `_. This resource covers
+detailed guides on all non-linear searches supported by PyAutoFit and provides insights into how to tailor them to your
+needs.
+
+There are currently no documentation guiding reads on what search might be appropriate for their problem and how to
+profile and experiment with different methods. Writing such documentation is on the to do list and will appear
+in the future. However, you can make progress now simply using visuals output by PyAutoFit and the ``search.summary` file.
+
+__Configs__
+
+As you refine your scientific workflow, you'll often find yourself repeatedly setting up models with identical priors
+and using the same non-linear search configurations. This repetition can result in lengthy Python scripts with
+redundant inputs.
+
+To streamline this process, configuration files can be utilized to define default values. This approach eliminates
+the need to specify identical prior inputs and search settings in every script, leading to more concise and
+readable Python code. Moreover, it reduces the cognitive load associated with performing model-fitting tasks.
+
+For a comprehensive guide on setting up and utilizing configuration files effectively, refer
+to the `configs cookbook `_. This resource provides
+detailed instructions on configuring and optimizing your PyAutoFit workflow through the use of configuration files.
+
+__Database__
+
+By default, model-fitting results are written to folders on hard-disk, which is straightforward for navigating and
+manual inspection. However, this approach becomes impractical for large datasets or extensive scientific workflows,
+where manually checking each result can be time-consuming.
+
+To address this challenge, all results can be stored in an sqlite3 relational database. This enables loading results
+directly into Jupyter notebooks or Python scripts for inspection, analysis, and interpretation. The database
+supports advanced querying capabilities, allowing users to retrieve specific model-fits based on criteria such
+as the fitted model or dataset.
+
+For a comprehensive guide on using the database functionality within PyAutoFit, refer to
+the `database cookbook `. This resource
+provides detailed instructions on leveraging the database to manage and analyze model-fitting results efficiently.
+
+__Scaling Up__
+
+Regardless of your final scientific objective, it's crucial to consider scalability in your scientific workflow and
+ensure it remains flexible to accommodate varying scales of complexity.
+
+Initially, scientific studies often begin with a small number of datasets (e.g., tens of datasets). During this phase,
+researchers iteratively refine their models and gain insights through trial and error. This involves fitting numerous
+models to datasets and manually inspecting results to evaluate model performance. A flexible workflow is essential
+here, allowing rapid iteration and outputting results in a format that facilitates quick inspection and interpretation.
+
+As the study progresses, researchers may scale up to larger datasets (e.g., thousands of datasets). Manual inspection
+of individual results becomes impractical, necessitating a more automated approach to model fitting and interpretation.
+Additionally, analyses may transition to high-performance computing environments, requiring output formats suitable for
+these setups.
+
+**PyAutoFit** is designed to enable the development of effective scientific workflows for both small and large datasets.
+
+__Wrap Up__
+
+This overview has provided a comprehensive guide to the key features of **PyAutoFit** that support the development of
+effective scientific workflows. By leveraging these tools, researchers can tailor their workflows to specific problems,
+streamline model fitting, and gain valuable insights into their scientific studies.
+
+The final aspect of core functionality, described in the next overview, is the wide variety of statistical
+inference methods available in **PyAutoFit**. These methods include graphical models, hierarchical models,
+Bayesian model comparison and many more.
+"""
diff --git a/scripts/plot/DynestyPlotter.py b/scripts/plot/DynestyPlotter.py
index 4b884e52..dbd2eb4e 100644
--- a/scripts/plot/DynestyPlotter.py
+++ b/scripts/plot/DynestyPlotter.py
@@ -1,344 +1,340 @@
-"""
-Plots: DynestyPlotter
-=====================
-
-This example illustrates how to plot visualization summarizing the results of a dynesty non-linear search using
-the `autofit.plot` module-level functions.
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Notation**: How parameter labels and superscripts are customized for plots.
-- **Plotting**: Using the plot functions to visualize Dynesty search results.
-- **Search Specific Visualization**: Accessing the native Dynesty sampler for custom visualizations.
-- **Plots**: Producing Dynesty-specific diagnostic plots.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import matplotlib.pyplot as plt
-from os import path
-
-import autofit as af
-import autofit.plot as aplt
-
-"""
-First, lets create a result via dynesty by repeating the simple model-fit that is performed in
-the `overview/simple/fit.py` example.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-model = af.Model(af.ex.Gaussian)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-search = af.DynestyStatic(path_prefix="plot", name="NestPlotter")
-
-result = search.fit(model=model, analysis=analysis)
-
-samples = result.samples
-
-"""
-__Notation__
-
-Plot are labeled with short hand parameter names (e.g. the `centre` parameters are plotted using an `x`).
-
-The mappings of every parameter to its shorthand symbol for plots is specified in the `config/notation.yaml` file
-and can be customized.
-
-Each label also has a superscript corresponding to the model component the parameter originates from. For example,
-Gaussians are given the superscript `g`. This can also be customized in the `config/notation.yaml` file.
-
-__Plotting__
-
-We now use the `autofit.plot` module-level functions to visualize the results.
-
-The dynesty readthedocs describes fully all of the methods used below
-
- - https://dynesty.readthedocs.io/en/latest/quickstart.html
- - https://dynesty.readthedocs.io/en/latest/api.html#module-dynesty.plotting
-
-In all the examples below, we use the `kwargs` of this function to pass in any of the input parameters that are
-described in the API docs.
-
-Dynesty plotters use `_kwargs` dictionaries to pass visualization settings to matplotlib lib. For example, below,
-we:
-
- - Set the fontsize of the x and y labels by passing `label_kwargs={"fontsize": 16}`.
- - Set the fontsize of the title by passing `title_kwargs={"fontsize": "10"}`.
-
-There are other `_kwargs` inputs we pass as None, you should check out the Dynesty docs if you need to customize your
-figure.
-"""
-"""
-The `corner_anesthetic` function produces a triangle of 1D and 2D PDF's of every parameter using the library `anesthetic`.
-"""
-aplt.corner_anesthetic(samples=samples)
-
-"""
-The `corner_cornerpy` function produces a triangle of 1D and 2D PDF's of every parameter using the library `corner.py`.
-"""
-aplt.corner_cornerpy(
- samples=samples,
- dims=None,
- span=None,
- quantiles=[0.025, 0.5, 0.975],
- color="black",
- smooth=0.02,
- quantiles_2d=None,
- hist_kwargs=None,
- hist2d_kwargs=None,
- label_kwargs={"fontsize": 16},
- show_titles=True,
- title_fmt=".2f",
- title_kwargs={"fontsize": "10"},
- truths=None,
- truth_color="red",
- truth_kwargs=None,
- max_n_ticks=5,
- top_ticks=False,
- use_math_text=False,
- verbose=False,
-)
-
-"""
-__Search Specific Visualization__
-
-The internal sampler can be used to plot the results of the non-linear search.
-
-We do this using the `search_internal` attribute which contains the sampler in its native form.
-
-The first time you run a search, the `search_internal` attribute will be available because it is passed ot the
-result via memory.
-
-If you rerun the fit on a completed result, it will not be available in memory, and therefore
-will be loaded from the `files/search_internal` folder. The `search_internal` entry of the `output.yaml` must be true
-for this to be possible.
-"""
-search_internal = result.search_internal
-
-"""
-__Plots__
-
-All plots use dynesty's inbuilt plotting library and the model.
-"""
-from dynesty import plotting as dyplot
-
-model = result.model
-
-"""
-The boundplot plots the bounding distribution used to propose either (1) live points at a given iteration or (2) a
-specific dead point during the course of a run, projected onto the two dimensions specified by `dims`.
-"""
-dyplot.boundplot(
- results=search_internal.results,
- labels=model.parameter_labels_with_superscripts_latex,
- dims=(2, 2),
- it=-1, # The iteration number to make the plot.
- idx=None,
- prior_transform=None,
- periodic=None,
- reflective=None,
- ndraws=5000,
- color="gray",
- plot_kwargs=None,
- label_kwargs={"fontsize": 16},
- max_n_ticks=5,
- use_math_text=False,
- show_live=False,
- live_color="darkviolet",
- live_kwargs=None,
- span=None,
- fig=None,
-)
-
-plt.show()
-plt.close()
-
-"""
-The cornerbound plots the bounding distribution used to propose either (1) live points at a given iteration or (2) a
-specific dead point during the course of a run, projected onto all pairs of dimensions.
-"""
-dyplot.cornerbound(
- results=search_internal.results,
- labels=model.parameter_labels_with_superscripts_latex,
- it=-1, # The iteration number to make the plot.
- idx=None,
- dims=None,
- prior_transform=None,
- periodic=None,
- reflective=None,
- ndraws=5000,
- color="gray",
- plot_kwargs=None,
- label_kwargs={"fontsize": 16},
- max_n_ticks=5,
- use_math_text=False,
- show_live=False,
- live_color="darkviolet",
- live_kwargs=None,
- span=None,
- fig=None,
-)
-
-plt.show()
-plt.close()
-
-"""
-The cornerplot plots a corner plot of the 1-D and 2-D marginalized posteriors.
-"""
-
-try:
- dyplot.cornerplot(
- results=search_internal.results,
- labels=model.parameter_labels_with_superscripts_latex,
- dims=None,
- span=None,
- quantiles=[0.025, 0.5, 0.975],
- color="black",
- smooth=0.02,
- quantiles_2d=None,
- hist_kwargs=None,
- hist2d_kwargs=None,
- label_kwargs={"fontsize": 16},
- show_titles=True,
- title_fmt=".2f",
- title_kwargs={"fontsize": "10"},
- truths=None,
- truth_color="red",
- truth_kwargs=None,
- max_n_ticks=5,
- top_ticks=False,
- use_math_text=False,
- verbose=False,
- )
-
- plt.show()
- plt.close()
-
-except ValueError:
- pass
-
-"""
-The cornerpoints plots a (sub-)corner plot of (weighted) samples.
-"""
-dyplot.cornerpoints(
- results=search_internal.results,
- labels=model.parameter_labels_with_superscripts_latex,
- dims=None,
- thin=1,
- span=None,
- cmap="plasma",
- color=None,
- kde=True,
- nkde=1000,
- plot_kwargs=None,
- label_kwargs={"fontsize": 16},
- truths=None,
- truth_color="red",
- truth_kwargs=None,
- max_n_ticks=5,
- use_math_text=False,
- fig=None,
-)
-
-plt.show()
-plt.close()
-
-
-"""
-The runplot plots live points, ln(likelihood), ln(weight), and ln(evidence) as a function of ln(prior volume).
-"""
-dyplot.runplot(
- results=search_internal.results,
- span=None,
- logplot=False,
- kde=True,
- nkde=1000,
- color="blue",
- plot_kwargs=None,
- label_kwargs={"fontsize": 16},
- lnz_error=True,
- lnz_truth=None,
- truth_color="red",
- truth_kwargs=None,
- max_x_ticks=8,
- max_y_ticks=3,
- use_math_text=True,
- mark_final_live=True,
- fig=None,
-)
-
-plt.show()
-plt.close()
-
-
-"""
-The traceplot plots traces and marginalized posteriors for each parameter.
-"""
-try:
- dyplot.traceplot(
- results=search_internal.results,
- span=None,
- quantiles=[0.025, 0.5, 0.975],
- smooth=0.02,
- thin=1,
- dims=None,
- post_color="blue",
- post_kwargs=None,
- kde=True,
- nkde=1000,
- trace_cmap="plasma",
- trace_color=None,
- trace_kwargs=None,
- connect=False,
- connect_highlight=10,
- connect_color="red",
- connect_kwargs=None,
- max_n_ticks=5,
- use_math_text=False,
- label_kwargs={"fontsize": 16},
- show_titles=True,
- title_fmt=".2f",
- title_kwargs={"fontsize": "10"},
- truths=None,
- truth_color="red",
- truth_kwargs=None,
- verbose=False,
- fig=None,
- )
-
- plt.show()
- plt.close()
-
-except ValueError:
- pass
-
-"""
-Finish.
-"""
+"""
+Plots: DynestyPlotter
+=====================
+
+This example illustrates how to plot visualization summarizing the results of a dynesty non-linear search using
+the `autofit.plot` module-level functions.
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Notation**: How parameter labels and superscripts are customized for plots.
+- **Plotting**: Using the plot functions to visualize Dynesty search results.
+- **Search Specific Visualization**: Accessing the native Dynesty sampler for custom visualizations.
+- **Plots**: Producing Dynesty-specific diagnostic plots.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import matplotlib.pyplot as plt
+from os import path
+
+import autofit as af
+import autofit.plot as aplt
+
+"""
+First, lets create a result via dynesty by repeating the simple model-fit that is performed in
+the `overview/simple/fit.py` example.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+model = af.Model(af.ex.Gaussian)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+search = af.DynestyStatic(path_prefix="plot", name="NestPlotter")
+
+result = search.fit(model=model, analysis=analysis)
+
+samples = result.samples
+
+"""
+__Notation__
+
+Plot are labeled with short hand parameter names (e.g. the `centre` parameters are plotted using an `x`).
+
+The mappings of every parameter to its shorthand symbol for plots is specified in the `config/notation.yaml` file
+and can be customized.
+
+Each label also has a superscript corresponding to the model component the parameter originates from. For example,
+Gaussians are given the superscript `g`. This can also be customized in the `config/notation.yaml` file.
+
+__Plotting__
+
+We now use the `autofit.plot` module-level functions to visualize the results.
+
+The dynesty readthedocs describes fully all of the methods used below
+
+ - https://dynesty.readthedocs.io/en/latest/quickstart.html
+ - https://dynesty.readthedocs.io/en/latest/api.html#module-dynesty.plotting
+
+In all the examples below, we use the `kwargs` of this function to pass in any of the input parameters that are
+described in the API docs.
+
+Dynesty plotters use `_kwargs` dictionaries to pass visualization settings to matplotlib lib. For example, below,
+we:
+
+ - Set the fontsize of the x and y labels by passing `label_kwargs={"fontsize": 16}`.
+ - Set the fontsize of the title by passing `title_kwargs={"fontsize": "10"}`.
+
+There are other `_kwargs` inputs we pass as None, you should check out the Dynesty docs if you need to customize your
+figure.
+"""
+"""
+The `corner_anesthetic` function produces a triangle of 1D and 2D PDF's of every parameter using the library `anesthetic`.
+"""
+aplt.corner_anesthetic(samples=samples)
+
+"""
+The `corner_cornerpy` function produces a triangle of 1D and 2D PDF's of every parameter using the library `corner.py`.
+"""
+aplt.corner_cornerpy(
+ samples=samples,
+ dims=None,
+ span=None,
+ quantiles=[0.025, 0.5, 0.975],
+ color="black",
+ smooth=0.02,
+ quantiles_2d=None,
+ hist_kwargs=None,
+ hist2d_kwargs=None,
+ label_kwargs={"fontsize": 16},
+ show_titles=True,
+ title_fmt=".2f",
+ title_kwargs={"fontsize": "10"},
+ truths=None,
+ truth_color="red",
+ truth_kwargs=None,
+ max_n_ticks=5,
+ top_ticks=False,
+ use_math_text=False,
+ verbose=False,
+)
+
+"""
+__Search Specific Visualization__
+
+The internal sampler can be used to plot the results of the non-linear search.
+
+We do this using the `search_internal` attribute which contains the sampler in its native form.
+
+The first time you run a search, the `search_internal` attribute will be available because it is passed ot the
+result via memory.
+
+If you rerun the fit on a completed result, it will not be available in memory, and therefore
+will be loaded from the `files/search_internal` folder. The `search_internal` entry of the `output.yaml` must be true
+for this to be possible.
+"""
+search_internal = result.search_internal
+
+"""
+__Plots__
+
+All plots use dynesty's inbuilt plotting library and the model.
+"""
+from dynesty import plotting as dyplot
+
+model = result.model
+
+"""
+The boundplot plots the bounding distribution used to propose either (1) live points at a given iteration or (2) a
+specific dead point during the course of a run, projected onto the two dimensions specified by `dims`.
+"""
+dyplot.boundplot(
+ results=search_internal.results,
+ labels=model.parameter_labels_with_superscripts_latex,
+ dims=(2, 2),
+ it=-1, # The iteration number to make the plot.
+ idx=None,
+ prior_transform=None,
+ periodic=None,
+ reflective=None,
+ ndraws=5000,
+ color="gray",
+ plot_kwargs=None,
+ label_kwargs={"fontsize": 16},
+ max_n_ticks=5,
+ use_math_text=False,
+ show_live=False,
+ live_color="darkviolet",
+ live_kwargs=None,
+ span=None,
+ fig=None,
+)
+
+plt.show()
+plt.close()
+
+"""
+The cornerbound plots the bounding distribution used to propose either (1) live points at a given iteration or (2) a
+specific dead point during the course of a run, projected onto all pairs of dimensions.
+"""
+dyplot.cornerbound(
+ results=search_internal.results,
+ labels=model.parameter_labels_with_superscripts_latex,
+ it=-1, # The iteration number to make the plot.
+ idx=None,
+ dims=None,
+ prior_transform=None,
+ periodic=None,
+ reflective=None,
+ ndraws=5000,
+ color="gray",
+ plot_kwargs=None,
+ label_kwargs={"fontsize": 16},
+ max_n_ticks=5,
+ use_math_text=False,
+ show_live=False,
+ live_color="darkviolet",
+ live_kwargs=None,
+ span=None,
+ fig=None,
+)
+
+plt.show()
+plt.close()
+
+"""
+The cornerplot plots a corner plot of the 1-D and 2-D marginalized posteriors.
+"""
+
+try:
+ dyplot.cornerplot(
+ results=search_internal.results,
+ labels=model.parameter_labels_with_superscripts_latex,
+ dims=None,
+ span=None,
+ quantiles=[0.025, 0.5, 0.975],
+ color="black",
+ smooth=0.02,
+ quantiles_2d=None,
+ hist_kwargs=None,
+ hist2d_kwargs=None,
+ label_kwargs={"fontsize": 16},
+ show_titles=True,
+ title_fmt=".2f",
+ title_kwargs={"fontsize": "10"},
+ truths=None,
+ truth_color="red",
+ truth_kwargs=None,
+ max_n_ticks=5,
+ top_ticks=False,
+ use_math_text=False,
+ verbose=False,
+ )
+
+ plt.show()
+ plt.close()
+
+except ValueError:
+ pass
+
+"""
+The cornerpoints plots a (sub-)corner plot of (weighted) samples.
+"""
+dyplot.cornerpoints(
+ results=search_internal.results,
+ labels=model.parameter_labels_with_superscripts_latex,
+ dims=None,
+ thin=1,
+ span=None,
+ cmap="plasma",
+ color=None,
+ kde=True,
+ nkde=1000,
+ plot_kwargs=None,
+ label_kwargs={"fontsize": 16},
+ truths=None,
+ truth_color="red",
+ truth_kwargs=None,
+ max_n_ticks=5,
+ use_math_text=False,
+ fig=None,
+)
+
+plt.show()
+plt.close()
+
+
+"""
+The runplot plots live points, ln(likelihood), ln(weight), and ln(evidence) as a function of ln(prior volume).
+"""
+dyplot.runplot(
+ results=search_internal.results,
+ span=None,
+ logplot=False,
+ kde=True,
+ nkde=1000,
+ color="blue",
+ plot_kwargs=None,
+ label_kwargs={"fontsize": 16},
+ lnz_error=True,
+ lnz_truth=None,
+ truth_color="red",
+ truth_kwargs=None,
+ max_x_ticks=8,
+ max_y_ticks=3,
+ use_math_text=True,
+ mark_final_live=True,
+ fig=None,
+)
+
+plt.show()
+plt.close()
+
+
+"""
+The traceplot plots traces and marginalized posteriors for each parameter.
+"""
+try:
+ dyplot.traceplot(
+ results=search_internal.results,
+ span=None,
+ quantiles=[0.025, 0.5, 0.975],
+ smooth=0.02,
+ thin=1,
+ dims=None,
+ post_color="blue",
+ post_kwargs=None,
+ kde=True,
+ nkde=1000,
+ trace_cmap="plasma",
+ trace_color=None,
+ trace_kwargs=None,
+ connect=False,
+ connect_highlight=10,
+ connect_color="red",
+ connect_kwargs=None,
+ max_n_ticks=5,
+ use_math_text=False,
+ label_kwargs={"fontsize": 16},
+ show_titles=True,
+ title_fmt=".2f",
+ title_kwargs={"fontsize": "10"},
+ truths=None,
+ truth_color="red",
+ truth_kwargs=None,
+ verbose=False,
+ fig=None,
+ )
+
+ plt.show()
+ plt.close()
+
+except ValueError:
+ pass
+
+"""
+Finish.
+"""
diff --git a/scripts/plot/EmceePlotter.py b/scripts/plot/EmceePlotter.py
index 16997752..1034aad0 100644
--- a/scripts/plot/EmceePlotter.py
+++ b/scripts/plot/EmceePlotter.py
@@ -1,199 +1,195 @@
-"""
-Plots: EmceePlotter
-===================
-
-This example illustrates how to plot visualization summarizing the results of a emcee non-linear search using
-the `autofit.plot` module-level functions.
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Notation**: How parameter labels and superscripts are customized for plots.
-- **Plotting**: Using the plot functions to visualize Emcee search results.
-- **Search Specific Visualization**: Accessing the native Emcee sampler for custom visualizations.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import matplotlib.pyplot as plt
-from os import path
-
-import autofit as af
-import autofit.plot as aplt
-
-"""
-First, lets create a result via emcee by repeating the simple model-fit that is performed in
-the `overview/simple/fit.py` example.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-model = af.Model(af.ex.Gaussian)
-
-model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.normalization = af.UniformPrior(lower_limit=1e-2, upper_limit=1e2)
-model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-search = af.Emcee(
- path_prefix=path.join("plot"), name="MCMCPlotter", nwalkers=100, nsteps=500
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-"""
-__Notation__
-
-Plot are labeled with short hand parameter names (e.g. the `centre` parameters are plotted using an `x`).
-
-The mappings of every parameter to its shorthand symbol for plots is specified in the `config/notation.yaml` file
-and can be customized.
-
-Each label also has a superscript corresponding to the model component the parameter originates from. For example,
-Gaussians are given the superscript `g`. This can also be customized in the `config/notation.yaml` file.
-
-__Plotting__
-
-We now use the `autofit.plot` module-level functions to visualize the results.
-
-The emcee readthedocs describes fully all of the methods used below
-
- - https://emcee.readthedocs.io/en/stable/user/sampler/
-
-The `aplt.corner_cornerpy` function wraps the library `corner.py` to make corner plots of the PDF:
-
-- https://corner.readthedocs.io/en/latest/index.html
-
-In all the examples below, we use the `kwargs` of this function to pass in any of the input parameters that are
-described in the API docs.
-"""
-samples = result.samples
-
-"""
-The `corner_cornerpy` function produces a triangle of 1D and 2D PDF's of every parameter using the library `corner.py`.
-"""
-aplt.corner_cornerpy(
- samples=samples,
- bins=20,
- range=None,
- color="k",
- hist_bin_factor=1,
- smooth=None,
- smooth1d=None,
- label_kwargs=None,
- titles=None,
- show_titles=False,
- title_fmt=".2f",
- title_kwargs=None,
- truths=None,
- truth_color="#4682b4",
- scale_hist=False,
- quantiles=None,
- verbose=False,
- fig=None,
- max_n_ticks=5,
- top_ticks=False,
- use_math_text=False,
- reverse=False,
- labelpad=0.0,
- hist_kwargs=None,
- group="posterior",
- var_names=None,
- filter_vars=None,
- coords=None,
- divergences=False,
- divergences_kwargs=None,
- labeller=None,
-)
-
-"""
-__Search Specific Visualization__
-
-The internal sampler can be used to plot the results of the non-linear search.
-
-We do this using the `search_internal` attribute which contains the sampler in its native form.
-
-The first time you run a search, the `search_internal` attribute will be available because it is passed ot the
-result via memory.
-
-If you rerun the fit on a completed result, it will not be available in memory, and therefore
-will be loaded from the `files/search_internal` folder. The `search_internal` entry of the `output.yaml` must be true
-for this to be possible.
-"""
-search_internal = result.search_internal
-
-"""
-The method below shows a 2D projection of the walker trajectories.
-"""
-fig, axes = plt.subplots(result.model.prior_count, figsize=(10, 7))
-
-for i in range(result.model.prior_count):
- for walker_index in range(search_internal.get_log_prob().shape[1]):
- ax = axes[i]
- ax.plot(
- search_internal.get_chain()[:, walker_index, i],
- search_internal.get_log_prob()[:, walker_index],
- alpha=0.3,
- )
-
- ax.set_ylabel("Log Likelihood")
- ax.set_xlabel(result.model.parameter_labels_with_superscripts_latex[i])
-
-plt.show()
-
-"""
-This method shows the likelihood as a series of steps.
-"""
-
-fig, axes = plt.subplots(1, figsize=(10, 7))
-
-for walker_index in range(search_internal.get_log_prob().shape[1]):
- axes.plot(search_internal.get_log_prob()[:, walker_index], alpha=0.3)
-
-axes.set_ylabel("Log Likelihood")
-axes.set_xlabel("step number")
-
-plt.show()
-
-"""
-This method shows the parameter values of every walker at every step.
-"""
-fig, axes = plt.subplots(result.samples.model.prior_count, figsize=(10, 7), sharex=True)
-
-for i in range(result.samples.model.prior_count):
- ax = axes[i]
- ax.plot(search_internal.get_chain()[:, :, i], alpha=0.3)
- ax.set_ylabel(result.model.parameter_labels_with_superscripts_latex[i])
-
-axes[-1].set_xlabel("step number")
-
-plt.show()
-
-"""
-Finish.
-"""
+"""
+Plots: EmceePlotter
+===================
+
+This example illustrates how to plot visualization summarizing the results of a emcee non-linear search using
+the `autofit.plot` module-level functions.
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Notation**: How parameter labels and superscripts are customized for plots.
+- **Plotting**: Using the plot functions to visualize Emcee search results.
+- **Search Specific Visualization**: Accessing the native Emcee sampler for custom visualizations.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import matplotlib.pyplot as plt
+from os import path
+
+import autofit as af
+import autofit.plot as aplt
+
+"""
+First, lets create a result via emcee by repeating the simple model-fit that is performed in
+the `overview/simple/fit.py` example.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+model = af.Model(af.ex.Gaussian)
+
+model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.normalization = af.UniformPrior(lower_limit=1e-2, upper_limit=1e2)
+model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+search = af.Emcee(
+ path_prefix=path.join("plot"), name="MCMCPlotter", nwalkers=100, nsteps=500
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+"""
+__Notation__
+
+Plot are labeled with short hand parameter names (e.g. the `centre` parameters are plotted using an `x`).
+
+The mappings of every parameter to its shorthand symbol for plots is specified in the `config/notation.yaml` file
+and can be customized.
+
+Each label also has a superscript corresponding to the model component the parameter originates from. For example,
+Gaussians are given the superscript `g`. This can also be customized in the `config/notation.yaml` file.
+
+__Plotting__
+
+We now use the `autofit.plot` module-level functions to visualize the results.
+
+The emcee readthedocs describes fully all of the methods used below
+
+ - https://emcee.readthedocs.io/en/stable/user/sampler/
+
+The `aplt.corner_cornerpy` function wraps the library `corner.py` to make corner plots of the PDF:
+
+- https://corner.readthedocs.io/en/latest/index.html
+
+In all the examples below, we use the `kwargs` of this function to pass in any of the input parameters that are
+described in the API docs.
+"""
+samples = result.samples
+
+"""
+The `corner_cornerpy` function produces a triangle of 1D and 2D PDF's of every parameter using the library `corner.py`.
+"""
+aplt.corner_cornerpy(
+ samples=samples,
+ bins=20,
+ range=None,
+ color="k",
+ hist_bin_factor=1,
+ smooth=None,
+ smooth1d=None,
+ label_kwargs=None,
+ titles=None,
+ show_titles=False,
+ title_fmt=".2f",
+ title_kwargs=None,
+ truths=None,
+ truth_color="#4682b4",
+ scale_hist=False,
+ quantiles=None,
+ verbose=False,
+ fig=None,
+ max_n_ticks=5,
+ top_ticks=False,
+ use_math_text=False,
+ reverse=False,
+ labelpad=0.0,
+ hist_kwargs=None,
+ group="posterior",
+ var_names=None,
+ filter_vars=None,
+ coords=None,
+ divergences=False,
+ divergences_kwargs=None,
+ labeller=None,
+)
+
+"""
+__Search Specific Visualization__
+
+The internal sampler can be used to plot the results of the non-linear search.
+
+We do this using the `search_internal` attribute which contains the sampler in its native form.
+
+The first time you run a search, the `search_internal` attribute will be available because it is passed ot the
+result via memory.
+
+If you rerun the fit on a completed result, it will not be available in memory, and therefore
+will be loaded from the `files/search_internal` folder. The `search_internal` entry of the `output.yaml` must be true
+for this to be possible.
+"""
+search_internal = result.search_internal
+
+"""
+The method below shows a 2D projection of the walker trajectories.
+"""
+fig, axes = plt.subplots(result.model.prior_count, figsize=(10, 7))
+
+for i in range(result.model.prior_count):
+ for walker_index in range(search_internal.get_log_prob().shape[1]):
+ ax = axes[i]
+ ax.plot(
+ search_internal.get_chain()[:, walker_index, i],
+ search_internal.get_log_prob()[:, walker_index],
+ alpha=0.3,
+ )
+
+ ax.set_ylabel("Log Likelihood")
+ ax.set_xlabel(result.model.parameter_labels_with_superscripts_latex[i])
+
+plt.show()
+
+"""
+This method shows the likelihood as a series of steps.
+"""
+
+fig, axes = plt.subplots(1, figsize=(10, 7))
+
+for walker_index in range(search_internal.get_log_prob().shape[1]):
+ axes.plot(search_internal.get_log_prob()[:, walker_index], alpha=0.3)
+
+axes.set_ylabel("Log Likelihood")
+axes.set_xlabel("step number")
+
+plt.show()
+
+"""
+This method shows the parameter values of every walker at every step.
+"""
+fig, axes = plt.subplots(result.samples.model.prior_count, figsize=(10, 7), sharex=True)
+
+for i in range(result.samples.model.prior_count):
+ ax = axes[i]
+ ax.plot(search_internal.get_chain()[:, :, i], alpha=0.3)
+ ax.set_ylabel(result.model.parameter_labels_with_superscripts_latex[i])
+
+axes[-1].set_xlabel("step number")
+
+plt.show()
+
+"""
+Finish.
+"""
diff --git a/scripts/plot/GetDist.py b/scripts/plot/GetDist.py
index 6a99493c..cddd8d8d 100644
--- a/scripts/plot/GetDist.py
+++ b/scripts/plot/GetDist.py
@@ -1,261 +1,257 @@
-"""
-Plots: GetDist
-==============
-
-This example illustrates how to plot visualization summarizing the results of model-fit using any non-linear search
-using GetDist:
-
- - https://getdist.readthedocs.io/en/latest/
-
-GetDist is an optional library which creates 1D and 2D plots of probability distribution functions (PDF)s. Its
-visualization tools has more than the in-built visualization tools of many non-linear searches (e.g. dynesty /
-emcee) and can often produce better looking plots.
-
-GetDist was developed for the analysis of Cosmological datasets.
-
-Installation
-------------
-
-Because GetDist is an optional library, you will likely have to install it manually via the command:
-
-`pip install getdist`
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Model Fit**: Create a Dynesty result for visualization with GetDist.
-- **Param Names**: Generate the GetDist parameter names file.
-- **GetDist MCSamples**: Create a GetDist MCSamples object from PyAutoFit samples.
-- **Parameter Names**: Document the parameter naming conventions.
-- **GetDist Plotter**: Create a GetDist plotter object.
-- **GetDist Subplots**: Create triangle plots and other multi-parameter plots.
-- **GetDist Single Plots**: Create individual 1D, 2D, and 3D PDF plots.
-- **Output**: Save figures to disk.
-- **GetDist Other Plots**: Reference additional plot options available in GetDist.
-- **Plotting Multiple Samples**: Demonstrate plotting results from multiple searches.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-
-import numpy as np
-import matplotlib.pyplot as plt
-from os import path
-
-from getdist import MCSamples
-from getdist import plots
-
-import autofit as af
-
-"""
-__Model Fit__
-
-First, lets create a result so we have samples to plot, repeating the simple model-fit that is performed in
-the `overview/simple/fit.py` example.
-
-We'll use dynesty in this example, but any MCMC / nested sampling non-linear search which produces samples of
-the posterior could be used.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-model = af.Model(af.ex.Gaussian)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-search = af.DynestyStatic(path_prefix="plot", name="GetDist")
-
-result = search.fit(model=model, analysis=analysis)
-
-samples = result.samples
-
-"""
-__Param Names__
-
-GetDist uses a `model.paramnames` file to load the name of every parameter in the model-fit and pair it with the
-latex symbol used to represent it in plots.
-
-This file is not created by **PyAutoLens** by default, but can be output by the `search.paths` object as shown below.
-"""
-search.paths._save_parameter_names_file(model=model)
-search.paths.zip_remove()
-search.paths._zip()
-
-
-"""
-__GetDist MCSamples__
-
-GetDist uses an `MCSamples` object to store the samples of a non-linear search.
-
-We create this object via a conversion from **PyAutoFit** `Samples`, as well as using the `names`
-and `labels` of parameters in the `Samples` object.
-
-The input `sampler="nested"` is input because we used a nested sampling, `dynesty`. For MCMC this should be
-replaced with "mcmc".
-"""
-
-gd_samples = MCSamples(
- samples=np.asarray(samples.parameter_lists),
- loglikes=np.asarray(samples.log_likelihood_list),
- weights=np.asarray(samples.weight_list),
- names=samples.model.model_component_and_parameter_names,
- labels=samples.model.parameter_labels_with_superscripts,
- sampler="nested",
-)
-
-"""
-__Parameter Names__
-
-Note that in order to customize the figure, we will use the `samples.model.parameter_names` list.
-"""
-print(samples.model.model_component_and_parameter_names)
-
-"""
-__GetDist Plotter__
-
-To make plots we use a GetDist plotter object, which can be customized to change the appearance of the plots.
-"""
-gd_plotter = plots.get_subplot_plotter(width_inch=12)
-
-"""
-__GetDist Subplots__
-
-Using the plotter we can make different plots, for example a triangle plot showing the 1D and 2D PDFs of every
-parameter.
-"""
-gd_plotter.triangle_plot(roots=gd_samples, filled=True)
-
-plt.show()
-plt.close()
-
-"""
-A triangle plot with specific parameters can be plotted by using the `params` input, whereby we specify the specific
-parameter names to plot.
-"""
-gd_plotter.triangle_plot(roots=gd_samples, filled=True, params=["centre", "sigma"])
-
-plt.show()
-plt.close()
-
-"""
-Rectangle plots can be used to show specific 2D combinations of parameters.
-"""
-gd_plotter.rectangle_plot(
- roots=gd_samples, yparams=["centre"], xparams=["normalization", "sigma"]
-)
-
-plt.show()
-plt.close()
-
-"""
-__GetDist Single Plots__
-
-We can make plots of specific 1D or 2D PDFs, using the single plotter object.
-"""
-gd_plotter = plots.get_single_plotter()
-
-gd_plotter.plot_1d(roots=gd_samples, param="centre")
-
-plt.show()
-plt.close()
-
-gd_plotter = plots.get_single_plotter()
-
-gd_plotter.plot_2d(roots=gd_samples, param1="centre", param2="sigma")
-
-plt.show()
-plt.close()
-
-"""
-We can also make a 3D plot, where the 2D PDF is plotted colored by the value of a third parameter.
-"""
-gd_plotter = plots.get_single_plotter()
-
-gd_plotter.plot_3d(roots=gd_samples, params=["centre", "sigma", "normalization"])
-
-plt.show()
-plt.close()
-
-"""
-__Output__
-
-A figure can be output using standard matplotlib functionality.
-"""
-
-gd_plotter = plots.get_single_plotter()
-
-gd_plotter.plot_3d(roots=gd_samples, params=["centre", "sigma", "normalization"])
-
-output_path = path.join("output")
-
-plt.savefig(path.join(output_path, "getdist.png"))
-plt.close()
-
-"""
-__GetDist Other Plots__
-
-There are many more ways to visualize PDFs possible with GetDist, checkout the official documentation for them all!
-
- - https://getdist.readthedocs.io/en/latest/
- - https://getdist.readthedocs.io/en/latest/plots.html
-
-__Plotting Multiple Samples__
-
-Finally, we can plot the results of multiple different non-linear searches on the same plot, using all
-of the functions above.
-
-Lets quickly make a second set of `dynesty` results and plot them on the same figure above with the results
-of the first search.
-"""
-
-search = af.DynestyStatic(path_prefix="plot", name="GetDist_2")
-
-result_extra = search.fit(model=model, analysis=analysis)
-
-samples_extra = result_extra.samples
-
-gd_samples_extra = MCSamples(
- samples=np.asarray(samples_extra.parameter_lists),
- loglikes=np.asarray(samples_extra.log_likelihood_list),
- weights=np.asarray(samples_extra.weight_list),
- names=samples_extra.model.parameter_names,
- labels=samples_extra.model.parameter_labels_with_superscripts_latex,
- sampler="nested",
-)
-
-gd_plotter = plots.get_subplot_plotter(width_inch=12)
-
-gd_plotter.triangle_plot(roots=[gd_samples, gd_samples_extra], filled=True)
-
-plt.show()
-plt.close()
-
-"""
-Note that the models do not need to be the same to make the plots above.
-
-GetDist will clever use the `names` of the parameters to combine the parameters into customizeable PDF plots.
-"""
+"""
+Plots: GetDist
+==============
+
+This example illustrates how to plot visualization summarizing the results of model-fit using any non-linear search
+using GetDist:
+
+ - https://getdist.readthedocs.io/en/latest/
+
+GetDist is an optional library which creates 1D and 2D plots of probability distribution functions (PDF)s. Its
+visualization tools has more than the in-built visualization tools of many non-linear searches (e.g. dynesty /
+emcee) and can often produce better looking plots.
+
+GetDist was developed for the analysis of Cosmological datasets.
+
+Installation
+------------
+
+Because GetDist is an optional library, you will likely have to install it manually via the command:
+
+`pip install getdist`
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Model Fit**: Create a Dynesty result for visualization with GetDist.
+- **Param Names**: Generate the GetDist parameter names file.
+- **GetDist MCSamples**: Create a GetDist MCSamples object from PyAutoFit samples.
+- **Parameter Names**: Document the parameter naming conventions.
+- **GetDist Plotter**: Create a GetDist plotter object.
+- **GetDist Subplots**: Create triangle plots and other multi-parameter plots.
+- **GetDist Single Plots**: Create individual 1D, 2D, and 3D PDF plots.
+- **Output**: Save figures to disk.
+- **GetDist Other Plots**: Reference additional plot options available in GetDist.
+- **Plotting Multiple Samples**: Demonstrate plotting results from multiple searches.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+
+import numpy as np
+import matplotlib.pyplot as plt
+from os import path
+
+from getdist import MCSamples
+from getdist import plots
+
+import autofit as af
+
+"""
+__Model Fit__
+
+First, lets create a result so we have samples to plot, repeating the simple model-fit that is performed in
+the `overview/simple/fit.py` example.
+
+We'll use dynesty in this example, but any MCMC / nested sampling non-linear search which produces samples of
+the posterior could be used.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+model = af.Model(af.ex.Gaussian)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+search = af.DynestyStatic(path_prefix="plot", name="GetDist")
+
+result = search.fit(model=model, analysis=analysis)
+
+samples = result.samples
+
+"""
+__Param Names__
+
+GetDist uses a `model.paramnames` file to load the name of every parameter in the model-fit and pair it with the
+latex symbol used to represent it in plots.
+
+This file is not created by **PyAutoLens** by default, but can be output by the `search.paths` object as shown below.
+"""
+search.paths._save_parameter_names_file(model=model)
+search.paths.zip_remove()
+search.paths._zip()
+
+
+"""
+__GetDist MCSamples__
+
+GetDist uses an `MCSamples` object to store the samples of a non-linear search.
+
+We create this object via a conversion from **PyAutoFit** `Samples`, as well as using the `names`
+and `labels` of parameters in the `Samples` object.
+
+The input `sampler="nested"` is input because we used a nested sampling, `dynesty`. For MCMC this should be
+replaced with "mcmc".
+"""
+
+gd_samples = MCSamples(
+ samples=np.asarray(samples.parameter_lists),
+ loglikes=np.asarray(samples.log_likelihood_list),
+ weights=np.asarray(samples.weight_list),
+ names=samples.model.model_component_and_parameter_names,
+ labels=samples.model.parameter_labels_with_superscripts,
+ sampler="nested",
+)
+
+"""
+__Parameter Names__
+
+Note that in order to customize the figure, we will use the `samples.model.parameter_names` list.
+"""
+print(samples.model.model_component_and_parameter_names)
+
+"""
+__GetDist Plotter__
+
+To make plots we use a GetDist plotter object, which can be customized to change the appearance of the plots.
+"""
+gd_plotter = plots.get_subplot_plotter(width_inch=12)
+
+"""
+__GetDist Subplots__
+
+Using the plotter we can make different plots, for example a triangle plot showing the 1D and 2D PDFs of every
+parameter.
+"""
+gd_plotter.triangle_plot(roots=gd_samples, filled=True)
+
+plt.show()
+plt.close()
+
+"""
+A triangle plot with specific parameters can be plotted by using the `params` input, whereby we specify the specific
+parameter names to plot.
+"""
+gd_plotter.triangle_plot(roots=gd_samples, filled=True, params=["centre", "sigma"])
+
+plt.show()
+plt.close()
+
+"""
+Rectangle plots can be used to show specific 2D combinations of parameters.
+"""
+gd_plotter.rectangle_plot(
+ roots=gd_samples, yparams=["centre"], xparams=["normalization", "sigma"]
+)
+
+plt.show()
+plt.close()
+
+"""
+__GetDist Single Plots__
+
+We can make plots of specific 1D or 2D PDFs, using the single plotter object.
+"""
+gd_plotter = plots.get_single_plotter()
+
+gd_plotter.plot_1d(roots=gd_samples, param="centre")
+
+plt.show()
+plt.close()
+
+gd_plotter = plots.get_single_plotter()
+
+gd_plotter.plot_2d(roots=gd_samples, param1="centre", param2="sigma")
+
+plt.show()
+plt.close()
+
+"""
+We can also make a 3D plot, where the 2D PDF is plotted colored by the value of a third parameter.
+"""
+gd_plotter = plots.get_single_plotter()
+
+gd_plotter.plot_3d(roots=gd_samples, params=["centre", "sigma", "normalization"])
+
+plt.show()
+plt.close()
+
+"""
+__Output__
+
+A figure can be output using standard matplotlib functionality.
+"""
+
+gd_plotter = plots.get_single_plotter()
+
+gd_plotter.plot_3d(roots=gd_samples, params=["centre", "sigma", "normalization"])
+
+output_path = path.join("output")
+
+plt.savefig(path.join(output_path, "getdist.png"))
+plt.close()
+
+"""
+__GetDist Other Plots__
+
+There are many more ways to visualize PDFs possible with GetDist, checkout the official documentation for them all!
+
+ - https://getdist.readthedocs.io/en/latest/
+ - https://getdist.readthedocs.io/en/latest/plots.html
+
+__Plotting Multiple Samples__
+
+Finally, we can plot the results of multiple different non-linear searches on the same plot, using all
+of the functions above.
+
+Lets quickly make a second set of `dynesty` results and plot them on the same figure above with the results
+of the first search.
+"""
+
+search = af.DynestyStatic(path_prefix="plot", name="GetDist_2")
+
+result_extra = search.fit(model=model, analysis=analysis)
+
+samples_extra = result_extra.samples
+
+gd_samples_extra = MCSamples(
+ samples=np.asarray(samples_extra.parameter_lists),
+ loglikes=np.asarray(samples_extra.log_likelihood_list),
+ weights=np.asarray(samples_extra.weight_list),
+ names=samples_extra.model.parameter_names,
+ labels=samples_extra.model.parameter_labels_with_superscripts_latex,
+ sampler="nested",
+)
+
+gd_plotter = plots.get_subplot_plotter(width_inch=12)
+
+gd_plotter.triangle_plot(roots=[gd_samples, gd_samples_extra], filled=True)
+
+plt.show()
+plt.close()
+
+"""
+Note that the models do not need to be the same to make the plots above.
+
+GetDist will clever use the `names` of the parameters to combine the parameters into customizeable PDF plots.
+"""
diff --git a/scripts/plot/NautilusPlotter.py b/scripts/plot/NautilusPlotter.py
index 1dc3f2ff..5fd65ef8 100644
--- a/scripts/plot/NautilusPlotter.py
+++ b/scripts/plot/NautilusPlotter.py
@@ -1,144 +1,140 @@
-"""
-Plots: DynestyPlotter
-======================
-
-This example illustrates how to plot visualization summarizing the results of a nautilus non-linear search using
-the `autofit.plot` module-level functions.
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Notation**: How parameter labels and superscripts are customized for plots.
-- **Plotting**: Using the plot functions to visualize Nautilus search results.
-- **Search Specific Visualization**: Accessing the native Nautilus sampler for custom visualizations.
-- **Plots**: Producing Nautilus-specific diagnostic plots.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import numpy as np
-from os import path
-
-import autofit as af
-import autofit.plot as aplt
-
-"""
-First, lets create a result via nautilus by repeating the simple model-fit that is performed in
-the `overview/simple/fit.py` example.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-model = af.Model(af.ex.Gaussian)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-search = af.Nautilus(
- path_prefix="plot",
- name="NestPlotter",
- n_live=100, # Number of so-called live points. New bounds are constructed so that they encompass the live points.
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-samples = result.samples
-
-"""
-__Notation__
-
-Plot are labeled with short hand parameter names (e.g. the `centre` parameters are plotted using an `x`).
-
-The mappings of every parameter to its shorthand symbol for plots is specified in the `config/notation.yaml` file
-and can be customized.
-
-Each label also has a superscript corresponding to the model component the parameter originates from. For example,
-Gaussians are given the superscript `g`. This can also be customized in the `config/notation.yaml` file.
-
-__Plotting__
-
-We now use the `autofit.plot` module-level functions to visualize the results.
-
-The nautilus readthedocs describes fully all of the methods used below
-
- - https://nautilus-sampler.readthedocs.io/en/stable/guides/crash_course.html
-
-In all the examples below, we use the `kwargs` of this function to pass in any of the input parameters that are
-described in the API docs.
-
-Nautilus plotters use `_kwargs` dictionaries to pass visualization settings to matplotlib lib. For example, below,
-we:
-
- - Set the fontsize of the x and y labels by passing `label_kwargs={"fontsize": 16}`.
- - Set the fontsize of the title by passing `title_kwargs={"fontsize": "10"}`.
-
-There are other `_kwargs` inputs we pass as None, you should check out the Nautilus docs if you need to customize your
-figure.
-"""
-"""
-The `corner_anesthetic` function produces a triangle of 1D and 2D PDF's of every parameter using the library `anesthetic`.
-"""
-aplt.corner_anesthetic(samples=samples)
-
-"""
-The `corner_cornerpy` function produces a triangle of 1D and 2D PDF's of every parameter using the library `corner.py`.
-"""
-aplt.corner_cornerpy(
- samples=samples,
- panelsize=3.5,
- yticksize=16,
- xticksize=16,
- bins=20,
- plot_datapoints=False,
- plot_density=False,
- fill_contours=True,
- levels=(0.68, 0.95),
- labelpad=0.02,
- range=np.ones(model.total_free_parameters) * 0.999,
- label_kwargs={"fontsize": 24},
-)
-
-"""
-__Search Specific Visualization__
-
-The internal sampler can be used to plot the results of the non-linear search.
-
-We do this using the `search_internal` attribute which contains the sampler in its native form.
-
-The first time you run a search, the `search_internal` attribute will be available because it is passed ot the
-result via memory.
-
-If you rerun the fit on a completed result, it will not be available in memory, and therefore
-will be loaded from the `files/search_internal` folder. The `search_internal` entry of the `output.yaml` must be true
-for this to be possible.
-"""
-search_internal = result.search_internal
-
-"""
-__Plots__
-
-Nautilus example plots are not shown explicitly below, so checkout their docs for examples!
-"""
+"""
+Plots: DynestyPlotter
+======================
+
+This example illustrates how to plot visualization summarizing the results of a nautilus non-linear search using
+the `autofit.plot` module-level functions.
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Notation**: How parameter labels and superscripts are customized for plots.
+- **Plotting**: Using the plot functions to visualize Nautilus search results.
+- **Search Specific Visualization**: Accessing the native Nautilus sampler for custom visualizations.
+- **Plots**: Producing Nautilus-specific diagnostic plots.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import numpy as np
+from os import path
+
+import autofit as af
+import autofit.plot as aplt
+
+"""
+First, lets create a result via nautilus by repeating the simple model-fit that is performed in
+the `overview/simple/fit.py` example.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+model = af.Model(af.ex.Gaussian)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+search = af.Nautilus(
+ path_prefix="plot",
+ name="NestPlotter",
+ n_live=100, # Number of so-called live points. New bounds are constructed so that they encompass the live points.
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+samples = result.samples
+
+"""
+__Notation__
+
+Plot are labeled with short hand parameter names (e.g. the `centre` parameters are plotted using an `x`).
+
+The mappings of every parameter to its shorthand symbol for plots is specified in the `config/notation.yaml` file
+and can be customized.
+
+Each label also has a superscript corresponding to the model component the parameter originates from. For example,
+Gaussians are given the superscript `g`. This can also be customized in the `config/notation.yaml` file.
+
+__Plotting__
+
+We now use the `autofit.plot` module-level functions to visualize the results.
+
+The nautilus readthedocs describes fully all of the methods used below
+
+ - https://nautilus-sampler.readthedocs.io/en/stable/guides/crash_course.html
+
+In all the examples below, we use the `kwargs` of this function to pass in any of the input parameters that are
+described in the API docs.
+
+Nautilus plotters use `_kwargs` dictionaries to pass visualization settings to matplotlib lib. For example, below,
+we:
+
+ - Set the fontsize of the x and y labels by passing `label_kwargs={"fontsize": 16}`.
+ - Set the fontsize of the title by passing `title_kwargs={"fontsize": "10"}`.
+
+There are other `_kwargs` inputs we pass as None, you should check out the Nautilus docs if you need to customize your
+figure.
+"""
+"""
+The `corner_anesthetic` function produces a triangle of 1D and 2D PDF's of every parameter using the library `anesthetic`.
+"""
+aplt.corner_anesthetic(samples=samples)
+
+"""
+The `corner_cornerpy` function produces a triangle of 1D and 2D PDF's of every parameter using the library `corner.py`.
+"""
+aplt.corner_cornerpy(
+ samples=samples,
+ panelsize=3.5,
+ yticksize=16,
+ xticksize=16,
+ bins=20,
+ plot_datapoints=False,
+ plot_density=False,
+ fill_contours=True,
+ levels=(0.68, 0.95),
+ labelpad=0.02,
+ range=np.ones(model.total_free_parameters) * 0.999,
+ label_kwargs={"fontsize": 24},
+)
+
+"""
+__Search Specific Visualization__
+
+The internal sampler can be used to plot the results of the non-linear search.
+
+We do this using the `search_internal` attribute which contains the sampler in its native form.
+
+The first time you run a search, the `search_internal` attribute will be available because it is passed ot the
+result via memory.
+
+If you rerun the fit on a completed result, it will not be available in memory, and therefore
+will be loaded from the `files/search_internal` folder. The `search_internal` entry of the `output.yaml` must be true
+for this to be possible.
+"""
+search_internal = result.search_internal
+
+"""
+__Plots__
+
+Nautilus example plots are not shown explicitly below, so checkout their docs for examples!
+"""
diff --git a/scripts/plot/PySwarmsPlotter.py b/scripts/plot/PySwarmsPlotter.py
index a3793f10..bbb02177 100644
--- a/scripts/plot/PySwarmsPlotter.py
+++ b/scripts/plot/PySwarmsPlotter.py
@@ -1,133 +1,129 @@
-"""
-Plots: PySwarmsPlotter
-======================
-
-This example illustrates how to plot visualization summarizing the results of a pyswarms non-linear search using
-the `autofit.plot` module-level functions.
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Notation**: How parameter labels and superscripts are customized for plots.
-- **Plotting**: Using the plot functions to visualize PySwarms search results.
-- **Search Specific Visualization**: Accessing the native PySwarms optimizer for custom visualizations.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import matplotlib.pyplot as plt
-from os import path
-
-import autofit as af
-
-"""
-First, lets create a result via pyswarms by repeating the simple model-fit that is performed in
-the `overview/simple/fit.py` example.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-model = af.Model(af.ex.Gaussian)
-
-model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.normalization = af.UniformPrior(lower_limit=1e-2, upper_limit=1e2)
-model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-search = af.PySwarmsGlobal(
- path_prefix=path.join("plot"), name="MLEPlotter", n_particles=50, iters=10
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-samples = result.samples
-
-"""
-__Notation__
-
-Plot are labeled with short hand parameter names (e.g. the `centre` parameters are plotted using an `x`).
-
-The mappings of every parameter to its shorthand symbol for plots is specified in the `config/notation.yaml` file
-and can be customized.
-
-Each label also has a superscript corresponding to the model component the parameter originates from. For example,
-Gaussians are given the superscript `g`. This can also be customized in the `config/notation.yaml` file.
-
-__Plotting__
-
-We now use the `autofit.plot` module-level functions and pyswarms's in-built plotting libraries to
-make figures.
-
-The pyswarms readthedocs describes fully all of the methods used below
-
- - https://pyswarms.readthedocs.io/en/latest/api/pyswarms.utils.plotters.html
-
-In all the examples below, we use the `kwargs` of this function to pass in any of the input parameters that are
-described in the API docs.
-"""
-"""
-__Search Specific Visualization__
-
-PySwarms has bespoke in-built visualization tools that can be used to plot its results.
-
-The first time you run a search, the `search_internal` attribute will be available because it is passed ot the
-result via memory.
-
-If you rerun the fit on a completed result, it will not be available in memory, and therefore
-will be loaded from the `files/search_internal` folder. The `search_internal` entry of the `output.yaml` must be true
-for this to be possible.
-"""
-search_internal = result.search_internal
-
-"""
-The `contour` method shows a 2D projection of the particle trajectories.
-"""
-from pyswarms.utils import plotters
-
-plotters.plot_contour(
- pos_history=search_internal.pos_history,
- canvas=None,
- title="Trajectories",
- mark=None,
- designer=None,
- mesher=None,
- animator=None,
-)
-plt.show()
-
-plotters.plot_cost_history(
- cost_history=search_internal.cost_history,
- ax=None,
- title="Cost History",
- designer=None,
-)
-plt.show()
-
-"""
-Finish.
-"""
+"""
+Plots: PySwarmsPlotter
+======================
+
+This example illustrates how to plot visualization summarizing the results of a pyswarms non-linear search using
+the `autofit.plot` module-level functions.
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Notation**: How parameter labels and superscripts are customized for plots.
+- **Plotting**: Using the plot functions to visualize PySwarms search results.
+- **Search Specific Visualization**: Accessing the native PySwarms optimizer for custom visualizations.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import matplotlib.pyplot as plt
+from os import path
+
+import autofit as af
+
+"""
+First, lets create a result via pyswarms by repeating the simple model-fit that is performed in
+the `overview/simple/fit.py` example.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+model = af.Model(af.ex.Gaussian)
+
+model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.normalization = af.UniformPrior(lower_limit=1e-2, upper_limit=1e2)
+model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+search = af.PySwarmsGlobal(
+ path_prefix=path.join("plot"), name="MLEPlotter", n_particles=50, iters=10
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+samples = result.samples
+
+"""
+__Notation__
+
+Plot are labeled with short hand parameter names (e.g. the `centre` parameters are plotted using an `x`).
+
+The mappings of every parameter to its shorthand symbol for plots is specified in the `config/notation.yaml` file
+and can be customized.
+
+Each label also has a superscript corresponding to the model component the parameter originates from. For example,
+Gaussians are given the superscript `g`. This can also be customized in the `config/notation.yaml` file.
+
+__Plotting__
+
+We now use the `autofit.plot` module-level functions and pyswarms's in-built plotting libraries to
+make figures.
+
+The pyswarms readthedocs describes fully all of the methods used below
+
+ - https://pyswarms.readthedocs.io/en/latest/api/pyswarms.utils.plotters.html
+
+In all the examples below, we use the `kwargs` of this function to pass in any of the input parameters that are
+described in the API docs.
+"""
+"""
+__Search Specific Visualization__
+
+PySwarms has bespoke in-built visualization tools that can be used to plot its results.
+
+The first time you run a search, the `search_internal` attribute will be available because it is passed ot the
+result via memory.
+
+If you rerun the fit on a completed result, it will not be available in memory, and therefore
+will be loaded from the `files/search_internal` folder. The `search_internal` entry of the `output.yaml` must be true
+for this to be possible.
+"""
+search_internal = result.search_internal
+
+"""
+The `contour` method shows a 2D projection of the particle trajectories.
+"""
+from pyswarms.utils import plotters
+
+plotters.plot_contour(
+ pos_history=search_internal.pos_history,
+ canvas=None,
+ title="Trajectories",
+ mark=None,
+ designer=None,
+ mesher=None,
+ animator=None,
+)
+plt.show()
+
+plotters.plot_cost_history(
+ cost_history=search_internal.cost_history,
+ ax=None,
+ title="Cost History",
+ designer=None,
+)
+plt.show()
+
+"""
+Finish.
+"""
diff --git a/scripts/plot/UltraNestPlotter.py b/scripts/plot/UltraNestPlotter.py
index d07008d7..ea7a436c 100644
--- a/scripts/plot/UltraNestPlotter.py
+++ b/scripts/plot/UltraNestPlotter.py
@@ -1,125 +1,121 @@
-"""
-Plots: DynestyPlotter
-=======================
-
-This example illustrates how to plot visualization summarizing the results of a ultranest non-linear search using
-the `autofit.plot` module-level functions.
-
-Installation
-------------
-
-Because UltraNest is an optional library, you will likely have to install it manually via the command:
-
-`pip install ultranest`
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Notation**: How parameter labels and superscripts are customized for plots.
-- **Plotting**: Using the plot functions to visualize UltraNest search results.
-- **Search Specific Visualization**: Accessing the native UltraNest sampler for custom visualizations.
-- **Plots**: Producing UltraNest-specific diagnostic plots.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-from os import path
-
-import autofit as af
-import autofit.plot as aplt
-
-"""
-First, lets create a result via ultranest by repeating the simple model-fit that is performed in
-the `overview/simple/fit.py` example.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-model = af.Model(af.ex.Gaussian)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-search = af.UltraNest(path_prefix="plot", name="NestPlotter", max_ncalls=10)
-
-result = search.fit(model=model, analysis=analysis)
-
-samples = result.samples
-
-"""
-__Notation__
-
-Plot are labeled with short hand parameter names (e.g. the `centre` parameters are plotted using an `x`).
-
-The mappings of every parameter to its shorthand symbol for plots is specified in the `config/notation.yaml` file
-and can be customized.
-
-Each label also has a superscript corresponding to the model component the parameter originates from. For example,
-Gaussians are given the superscript `g`. This can also be customized in the `config/notation.yaml` file.
-
-__Plotting__
-
-We now use the `autofit.plot` module-level functions to visualize the results.
-
-The ultranest readthedocs describes fully all of the methods used below
-
- - https://johannesbuchner.github.io/UltraNest/readme.html
- - https://johannesbuchner.github.io/UltraNest/ultranest.html#module-ultranest.plot
-
-In all the examples below, we use the `kwargs` of this function to pass in any of the input parameters that are
-described in the API docs.
-"""
-"""
-The `corner_anesthetic` function produces a triangle of 1D and 2D PDF's of every parameter using the library `anesthetic`.
-"""
-aplt.corner_anesthetic(samples=samples)
-
-"""
-The `corner_cornerpy` function produces a triangle of 1D and 2D PDF's of every parameter using the library `corner.py`.
-"""
-aplt.corner_cornerpy(samples=samples)
-
-"""
-__Search Specific Visualization__
-
-The internal sampler can be used to plot the results of the non-linear search.
-
-We do this using the `search_internal` attribute which contains the sampler in its native form.
-
-The first time you run a search, the `search_internal` attribute will be available because it is passed ot the
-result via memory.
-
-If you rerun the fit on a completed result, it will not be available in memory, and therefore
-will be loaded from the `files/search_internal` folder. The `search_internal` entry of the `output.yaml` must be true
-for this to be possible.
-"""
-search_internal = result.search_internal
-
-"""
-__Plots__
-
-UltraNest example plots are not shown explicitly below, so checkout their docs for examples!
-"""
+"""
+Plots: DynestyPlotter
+=======================
+
+This example illustrates how to plot visualization summarizing the results of a ultranest non-linear search using
+the `autofit.plot` module-level functions.
+
+Installation
+------------
+
+Because UltraNest is an optional library, you will likely have to install it manually via the command:
+
+`pip install ultranest`
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Notation**: How parameter labels and superscripts are customized for plots.
+- **Plotting**: Using the plot functions to visualize UltraNest search results.
+- **Search Specific Visualization**: Accessing the native UltraNest sampler for custom visualizations.
+- **Plots**: Producing UltraNest-specific diagnostic plots.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+from os import path
+
+import autofit as af
+import autofit.plot as aplt
+
+"""
+First, lets create a result via ultranest by repeating the simple model-fit that is performed in
+the `overview/simple/fit.py` example.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+model = af.Model(af.ex.Gaussian)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+search = af.UltraNest(path_prefix="plot", name="NestPlotter", max_ncalls=10)
+
+result = search.fit(model=model, analysis=analysis)
+
+samples = result.samples
+
+"""
+__Notation__
+
+Plot are labeled with short hand parameter names (e.g. the `centre` parameters are plotted using an `x`).
+
+The mappings of every parameter to its shorthand symbol for plots is specified in the `config/notation.yaml` file
+and can be customized.
+
+Each label also has a superscript corresponding to the model component the parameter originates from. For example,
+Gaussians are given the superscript `g`. This can also be customized in the `config/notation.yaml` file.
+
+__Plotting__
+
+We now use the `autofit.plot` module-level functions to visualize the results.
+
+The ultranest readthedocs describes fully all of the methods used below
+
+ - https://johannesbuchner.github.io/UltraNest/readme.html
+ - https://johannesbuchner.github.io/UltraNest/ultranest.html#module-ultranest.plot
+
+In all the examples below, we use the `kwargs` of this function to pass in any of the input parameters that are
+described in the API docs.
+"""
+"""
+The `corner_anesthetic` function produces a triangle of 1D and 2D PDF's of every parameter using the library `anesthetic`.
+"""
+aplt.corner_anesthetic(samples=samples)
+
+"""
+The `corner_cornerpy` function produces a triangle of 1D and 2D PDF's of every parameter using the library `corner.py`.
+"""
+aplt.corner_cornerpy(samples=samples)
+
+"""
+__Search Specific Visualization__
+
+The internal sampler can be used to plot the results of the non-linear search.
+
+We do this using the `search_internal` attribute which contains the sampler in its native form.
+
+The first time you run a search, the `search_internal` attribute will be available because it is passed ot the
+result via memory.
+
+If you rerun the fit on a completed result, it will not be available in memory, and therefore
+will be loaded from the `files/search_internal` folder. The `search_internal` entry of the `output.yaml` must be true
+for this to be possible.
+"""
+search_internal = result.search_internal
+
+"""
+__Plots__
+
+UltraNest example plots are not shown explicitly below, so checkout their docs for examples!
+"""
diff --git a/scripts/plot/ZeusPlotter.py b/scripts/plot/ZeusPlotter.py
index 6ff3621d..7466833b 100644
--- a/scripts/plot/ZeusPlotter.py
+++ b/scripts/plot/ZeusPlotter.py
@@ -1,185 +1,181 @@
-"""
-Plots: ZeusPlotter
-==================
-
-This example illustrates how to plot visualization summarizing the results of a zeus non-linear search using
-the `autofit.plot` module-level functions.
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Notation**: How parameter labels and superscripts are customized for plots.
-- **Plotting**: Using the plot functions to visualize Zeus search results.
-- **Search Specific Visualization**: Accessing the native Zeus sampler for custom visualizations.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import matplotlib.pyplot as plt
-from os import path
-
-import autofit as af
-import autofit.plot as aplt
-
-"""
-First, lets create a result via zeus by repeating the simple model-fit that is performed in
-the `overview/simple/fit.py` example.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-model = af.Model(af.ex.Gaussian)
-
-model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.normalization = af.UniformPrior(lower_limit=1e-2, upper_limit=1e2)
-model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-search = af.Zeus(
- path_prefix=path.join("plot"), name="MCMCPlotter", nwalkers=100, nsteps=10000
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-samples = result.samples
-
-"""
-__Notation__
-
-Plot are labeled with short hand parameter names (e.g. the `centre` parameters are plotted using an `x`).
-
-The mappings of every parameter to its shorthand symbol for plots is specified in the `config/notation.yaml` file
-and can be customized.
-
-Each label also has a superscript corresponding to the model component the parameter originates from. For example,
-Gaussians are given the superscript `g`. This can also be customized in the `config/notation.yaml` file.
-
-__Plotting__
-
-We now use the `autofit.plot` module-level functions to visualize the results.
-
-The zeus readthedocs describes fully all of the methods used below
-
- - https://zeus-mcmc.readthedocs.io/en/latest/api/plotting.html#cornerplot
- - https://zeus-mcmc.readthedocs.io/en/latest/notebooks/normal_distribution.html
-
-The `aplt.corner_cornerpy` function wraps the library `corner.py` to make corner plots of the PDF:
-
-- https://corner.readthedocs.io/en/latest/index.html
-
-In all the examples below, we use the `kwargs` of this function to pass in any of the input parameters that are
-described in the API docs.
-"""
-"""
-The `corner_cornerpy` function produces a triangle of 1D and 2D PDF's of every parameter using the library `corner.py`.
-"""
-aplt.corner_cornerpy(
- samples=samples,
- weight_list=None,
- levels=None,
- span=None,
- quantiles=[0.025, 0.5, 0.975],
- truth=None,
- color=None,
- alpha=0.5,
- linewidth=1.5,
- fill=True,
- fontsize=10,
- show_titles=True,
- title_fmt=".2f",
- title_fontsize=12,
- cut=3,
- fig=None,
- size=(10, 10),
-)
-
-
-"""
-__Search Specific Visualization__
-
-The internal sampler can be used to plot the results of the non-linear search.
-
-The first time you run a search, the `search_internal` attribute will be available because it is passed ot the
-result via memory.
-
-If you rerun the fit on a completed result, it will not be available in memory, and therefore
-will be loaded from the `files/search_internal` folder. The `search_internal` entry of the `output.yaml` must be true
-for this to be possible.
-"""
-search_internal = result.search_internal
-
-"""
-The method below shows a 2D projection of the walker trajectories.
-"""
-fig, axes = plt.subplots(result.model.prior_count, figsize=(10, 7))
-
-for i in range(result.model.prior_count):
- for walker_index in range(search_internal.get_log_prob().shape[1]):
- ax = axes[i]
- ax.plot(
- search_internal.get_chain()[:, walker_index, i],
- search_internal.get_log_prob()[:, walker_index],
- alpha=0.3,
- )
-
- ax.set_ylabel("Log Likelihood")
- ax.set_xlabel(result.model.parameter_labels_with_superscripts_latex[i])
-
-plt.show()
-
-"""
-This method shows the likelihood as a series of steps.
-"""
-
-fig, axes = plt.subplots(1, figsize=(10, 7))
-
-for walker_index in range(search_internal.get_log_prob().shape[1]):
- axes.plot(search_internal.get_log_prob()[:, walker_index], alpha=0.3)
-
-axes.set_ylabel("Log Likelihood")
-axes.set_xlabel("step number")
-
-plt.show()
-
-"""
-This method shows the parameter values of every walker at every step.
-"""
-fig, axes = plt.subplots(result.samples.model.prior_count, figsize=(10, 7), sharex=True)
-
-for i in range(result.samples.model.prior_count):
- ax = axes[i]
- ax.plot(search_internal.get_chain()[:, :, i], alpha=0.3)
- ax.set_ylabel(result.model.parameter_labels_with_superscripts_latex[i])
-
-axes[-1].set_xlabel("step number")
-
-plt.show()
-
-"""
-Finish.
-"""
+"""
+Plots: ZeusPlotter
+==================
+
+This example illustrates how to plot visualization summarizing the results of a zeus non-linear search using
+the `autofit.plot` module-level functions.
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Notation**: How parameter labels and superscripts are customized for plots.
+- **Plotting**: Using the plot functions to visualize Zeus search results.
+- **Search Specific Visualization**: Accessing the native Zeus sampler for custom visualizations.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import matplotlib.pyplot as plt
+from os import path
+
+import autofit as af
+import autofit.plot as aplt
+
+"""
+First, lets create a result via zeus by repeating the simple model-fit that is performed in
+the `overview/simple/fit.py` example.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+model = af.Model(af.ex.Gaussian)
+
+model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.normalization = af.UniformPrior(lower_limit=1e-2, upper_limit=1e2)
+model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+search = af.Zeus(
+ path_prefix=path.join("plot"), name="MCMCPlotter", nwalkers=100, nsteps=10000
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+samples = result.samples
+
+"""
+__Notation__
+
+Plot are labeled with short hand parameter names (e.g. the `centre` parameters are plotted using an `x`).
+
+The mappings of every parameter to its shorthand symbol for plots is specified in the `config/notation.yaml` file
+and can be customized.
+
+Each label also has a superscript corresponding to the model component the parameter originates from. For example,
+Gaussians are given the superscript `g`. This can also be customized in the `config/notation.yaml` file.
+
+__Plotting__
+
+We now use the `autofit.plot` module-level functions to visualize the results.
+
+The zeus readthedocs describes fully all of the methods used below
+
+ - https://zeus-mcmc.readthedocs.io/en/latest/api/plotting.html#cornerplot
+ - https://zeus-mcmc.readthedocs.io/en/latest/notebooks/normal_distribution.html
+
+The `aplt.corner_cornerpy` function wraps the library `corner.py` to make corner plots of the PDF:
+
+- https://corner.readthedocs.io/en/latest/index.html
+
+In all the examples below, we use the `kwargs` of this function to pass in any of the input parameters that are
+described in the API docs.
+"""
+"""
+The `corner_cornerpy` function produces a triangle of 1D and 2D PDF's of every parameter using the library `corner.py`.
+"""
+aplt.corner_cornerpy(
+ samples=samples,
+ weight_list=None,
+ levels=None,
+ span=None,
+ quantiles=[0.025, 0.5, 0.975],
+ truth=None,
+ color=None,
+ alpha=0.5,
+ linewidth=1.5,
+ fill=True,
+ fontsize=10,
+ show_titles=True,
+ title_fmt=".2f",
+ title_fontsize=12,
+ cut=3,
+ fig=None,
+ size=(10, 10),
+)
+
+
+"""
+__Search Specific Visualization__
+
+The internal sampler can be used to plot the results of the non-linear search.
+
+The first time you run a search, the `search_internal` attribute will be available because it is passed ot the
+result via memory.
+
+If you rerun the fit on a completed result, it will not be available in memory, and therefore
+will be loaded from the `files/search_internal` folder. The `search_internal` entry of the `output.yaml` must be true
+for this to be possible.
+"""
+search_internal = result.search_internal
+
+"""
+The method below shows a 2D projection of the walker trajectories.
+"""
+fig, axes = plt.subplots(result.model.prior_count, figsize=(10, 7))
+
+for i in range(result.model.prior_count):
+ for walker_index in range(search_internal.get_log_prob().shape[1]):
+ ax = axes[i]
+ ax.plot(
+ search_internal.get_chain()[:, walker_index, i],
+ search_internal.get_log_prob()[:, walker_index],
+ alpha=0.3,
+ )
+
+ ax.set_ylabel("Log Likelihood")
+ ax.set_xlabel(result.model.parameter_labels_with_superscripts_latex[i])
+
+plt.show()
+
+"""
+This method shows the likelihood as a series of steps.
+"""
+
+fig, axes = plt.subplots(1, figsize=(10, 7))
+
+for walker_index in range(search_internal.get_log_prob().shape[1]):
+ axes.plot(search_internal.get_log_prob()[:, walker_index], alpha=0.3)
+
+axes.set_ylabel("Log Likelihood")
+axes.set_xlabel("step number")
+
+plt.show()
+
+"""
+This method shows the parameter values of every walker at every step.
+"""
+fig, axes = plt.subplots(result.samples.model.prior_count, figsize=(10, 7), sharex=True)
+
+for i in range(result.samples.model.prior_count):
+ ax = axes[i]
+ ax.plot(search_internal.get_chain()[:, :, i], alpha=0.3)
+ ax.set_ylabel(result.model.parameter_labels_with_superscripts_latex[i])
+
+axes[-1].set_xlabel("step number")
+
+plt.show()
+
+"""
+Finish.
+"""
diff --git a/scripts/searches/mcmc/Emcee.py b/scripts/searches/mcmc/Emcee.py
index 00b3d30c..16ff3b00 100644
--- a/scripts/searches/mcmc/Emcee.py
+++ b/scripts/searches/mcmc/Emcee.py
@@ -1,167 +1,163 @@
-"""
-Searches: DynestyStatic
-=======================
-
-This example illustrates how to use the MCMC ensamble sampler algorithm Emcee.
-
-Information about Emcee can be found at the following links:
-
- - https://github.com/dfm/emcee
- - https://emcee.readthedocs.io/en/stable/
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Data**: Loading and plotting the 1D Gaussian dataset used to demonstrate the search.
-- **Model + Analysis**: Setting up the model and analysis for the fitting example.
-- **Search**: Configuring and running the Emcee MCMC sampler.
-- **Result**: Inspecting the result and comparing the maximum log likelihood model to the data.
-- **Search Internal**: Accessing the internal Emcee sampler for advanced use.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import matplotlib.pyplot as plt
-import numpy as np
-from os import path
-
-import autofit as af
-
-"""
-__Data__
-
-This example fits a single 1D Gaussian, we therefore load and plot data containing one Gaussian.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-plt.errorbar(
- x=range(data.shape[0]),
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.show()
-plt.close()
-
-"""
-__Model + Analysis__
-
-We create the model and analysis, which in this example is a single `Gaussian` and therefore has dimensionality N=3.
-"""
-model = af.Model(af.ex.Gaussian)
-
-model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.normalization = af.UniformPrior(lower_limit=1e-2, upper_limit=1e2)
-model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-"""
-__Search__
-
-We now create and run the `Emcee` object which acts as our non-linear search.
-
-We manually specify all of the Emcee settings, descriptions of which are provided at the following webpage:
-
- https://emcee.readthedocs.io/en/stable/user/sampler/
- https://emcee.readthedocs.io/en/stable/
-"""
-search = af.Emcee(
- path_prefix="searches",
- name="Emcee",
- nwalkers=30,
- nsteps=1000,
- initializer=af.InitializerBall(lower_limit=0.49, upper_limit=0.51),
- auto_correlations_settings=af.AutoCorrelationsSettings(
- check_for_convergence=True,
- check_size=100,
- required_length=50,
- change_threshold=0.01,
- ),
- number_of_cores=1,
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-"""
-__Result__
-
-The result object returned by the fit provides information on the results of the non-linear search. Lets use it to
-compare the maximum log likelihood `Gaussian` to the data.
-"""
-model_data = result.max_log_likelihood_instance.model_data_from(
- xvalues=np.arange(data.shape[0])
-)
-
-plt.errorbar(
- x=range(data.shape[0]),
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(range(data.shape[0]), model_data, color="r")
-plt.title("Emcee model fit to 1D Gaussian dataset.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-__Search Internal__
-
-The result also contains the internal representation of the non-linear search.
-
-The internal representation of the non-linear search ensures that all sampling info is available in its native form.
-This can be passed to functions which take it as input, for example if the sampling package has bespoke visualization
-functions.
-
-For `Emcee`, this is an instance of the `Sampler` object (`from emcee import EnsembleSampler`).
-"""
-search_internal = result.search_internal
-
-print(search_internal)
-
-"""
-The internal search is by default not saved to hard-disk, because it can often take up quite a lot of hard-disk space
-(significantly more than standard output files).
-
-This means that the search internal will only be available the first time you run the search. If you rerun the code
-and the search is bypassed because the results already exist on hard-disk, the search internal will not be available.
-
-If you are frequently using the search internal you can have it saved to hard-disk by changing the `search_internal`
-setting in `output.yaml` to `True`. The result will then have the search internal available as an attribute,
-irrespective of whether the search is re-run or not.
-"""
+"""
+Searches: DynestyStatic
+=======================
+
+This example illustrates how to use the MCMC ensamble sampler algorithm Emcee.
+
+Information about Emcee can be found at the following links:
+
+ - https://github.com/dfm/emcee
+ - https://emcee.readthedocs.io/en/stable/
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Data**: Loading and plotting the 1D Gaussian dataset used to demonstrate the search.
+- **Model + Analysis**: Setting up the model and analysis for the fitting example.
+- **Search**: Configuring and running the Emcee MCMC sampler.
+- **Result**: Inspecting the result and comparing the maximum log likelihood model to the data.
+- **Search Internal**: Accessing the internal Emcee sampler for advanced use.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import matplotlib.pyplot as plt
+import numpy as np
+from os import path
+
+import autofit as af
+
+"""
+__Data__
+
+This example fits a single 1D Gaussian, we therefore load and plot data containing one Gaussian.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+plt.errorbar(
+ x=range(data.shape[0]),
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.show()
+plt.close()
+
+"""
+__Model + Analysis__
+
+We create the model and analysis, which in this example is a single `Gaussian` and therefore has dimensionality N=3.
+"""
+model = af.Model(af.ex.Gaussian)
+
+model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.normalization = af.UniformPrior(lower_limit=1e-2, upper_limit=1e2)
+model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+"""
+__Search__
+
+We now create and run the `Emcee` object which acts as our non-linear search.
+
+We manually specify all of the Emcee settings, descriptions of which are provided at the following webpage:
+
+ https://emcee.readthedocs.io/en/stable/user/sampler/
+ https://emcee.readthedocs.io/en/stable/
+"""
+search = af.Emcee(
+ path_prefix="searches",
+ name="Emcee",
+ nwalkers=30,
+ nsteps=1000,
+ initializer=af.InitializerBall(lower_limit=0.49, upper_limit=0.51),
+ auto_correlations_settings=af.AutoCorrelationsSettings(
+ check_for_convergence=True,
+ check_size=100,
+ required_length=50,
+ change_threshold=0.01,
+ ),
+ number_of_cores=1,
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+"""
+__Result__
+
+The result object returned by the fit provides information on the results of the non-linear search. Lets use it to
+compare the maximum log likelihood `Gaussian` to the data.
+"""
+model_data = result.max_log_likelihood_instance.model_data_from(
+ xvalues=np.arange(data.shape[0])
+)
+
+plt.errorbar(
+ x=range(data.shape[0]),
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(range(data.shape[0]), model_data, color="r")
+plt.title("Emcee model fit to 1D Gaussian dataset.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+__Search Internal__
+
+The result also contains the internal representation of the non-linear search.
+
+The internal representation of the non-linear search ensures that all sampling info is available in its native form.
+This can be passed to functions which take it as input, for example if the sampling package has bespoke visualization
+functions.
+
+For `Emcee`, this is an instance of the `Sampler` object (`from emcee import EnsembleSampler`).
+"""
+search_internal = result.search_internal
+
+print(search_internal)
+
+"""
+The internal search is by default not saved to hard-disk, because it can often take up quite a lot of hard-disk space
+(significantly more than standard output files).
+
+This means that the search internal will only be available the first time you run the search. If you rerun the code
+and the search is bypassed because the results already exist on hard-disk, the search internal will not be available.
+
+If you are frequently using the search internal you can have it saved to hard-disk by changing the `search_internal`
+setting in `output.yaml` to `True`. The result will then have the search internal available as an attribute,
+irrespective of whether the search is re-run or not.
+"""
diff --git a/scripts/searches/mcmc/Zeus.py b/scripts/searches/mcmc/Zeus.py
index 17ef759e..2760b3a6 100644
--- a/scripts/searches/mcmc/Zeus.py
+++ b/scripts/searches/mcmc/Zeus.py
@@ -1,177 +1,173 @@
-"""
-Searches: DynestyStatic
-=======================
-
-This example illustrates how to use the MCMC ensamble sampler algorithm Zeus.
-
-Information about Zeus can be found at the following links:
-
- - https://github.com/minaskar/zeus
- - https://zeus-mcmc.readthedocs.io/en/latest/
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Data**: Loading and plotting the 1D Gaussian dataset used to demonstrate the search.
-- **Model + Analysis**: Setting up the model and analysis for the fitting example.
-- **Search**: Configuring and running the Zeus MCMC sampler.
-- **Result**: Inspecting the result and comparing the maximum log likelihood model to the data.
-- **Search Internal**: Accessing the internal Zeus sampler for advanced use.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import matplotlib.pyplot as plt
-import numpy as np
-from os import path
-
-import autofit as af
-
-"""
-__Data__
-
-This example fits a single 1D Gaussian, we therefore load and plot data containing one Gaussian.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-plt.errorbar(
- x=range(data.shape[0]),
- y=data,
- yerr=noise_map,
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.show()
-plt.close()
-
-"""
-__Model + Analysis__
-
-We create the model and analysis, which in this example is a single `Gaussian` and therefore has dimensionality N=3.
-"""
-model = af.Model(af.ex.Gaussian)
-
-model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.normalization = af.UniformPrior(lower_limit=1e-2, upper_limit=1e2)
-model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-"""
-__Search__
-
-We now create and run the `Zeus` object which acts as our non-linear search.
-
-We manually specify all of the Zeus settings, descriptions of which are provided at the following webpage:
-
- https://zeus-mcmc.readthedocs.io/en/latest/
- https://zeus-mcmc.readthedocs.io/en/latest/api/sampler.html
-"""
-search = af.Zeus(
- path_prefix="searches",
- name="Zeus",
- nwalkers=30,
- nsteps=1001,
- initializer=af.InitializerBall(lower_limit=0.49, upper_limit=0.51),
- auto_correlations_settings=af.AutoCorrelationsSettings(
- check_for_convergence=True,
- check_size=100,
- required_length=50,
- change_threshold=0.01,
- ),
- tune=False,
- tolerance=0.05,
- patience=5,
- maxsteps=10000,
- mu=1.0,
- maxiter=10000,
- vectorize=False,
- check_walkers=True,
- shuffle_ensemble=True,
- light_mode=False,
- iterations_per_full_update=501,
- number_of_cores=1,
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-"""
-__Result__
-
-The result object returned by the fit provides information on the results of the non-linear search. Lets use it to
-compare the maximum log likelihood `Gaussian` to the data.
-"""
-model_data = result.max_log_likelihood_instance.model_data_from(
- xvalues=np.arange(data.shape[0])
-)
-
-plt.errorbar(
- x=range(data.shape[0]),
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(range(data.shape[0]), model_data, color="r")
-plt.title("DynestyStatic model fit to 1D Gaussian dataset.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-__Search Internal__
-
-The result also contains the internal representation of the non-linear search.
-
-The internal representation of the non-linear search ensures that all sampling info is available in its native form.
-This can be passed to functions which take it as input, for example if the sampling package has bespoke visualization
-functions.
-
-For `Emcee`, this is an instance of the `Sampler` object (`from zeus import EnsembleSampler`).
-"""
-search_internal = result.search_internal
-
-print(search_internal)
-
-"""
-The internal search is by default not saved to hard-disk, because it can often take up quite a lot of hard-disk space
-(significantly more than standard output files).
-
-This means that the search internal will only be available the first time you run the search. If you rerun the code
-and the search is bypassed because the results already exist on hard-disk, the search internal will not be available.
-
-If you are frequently using the search internal you can have it saved to hard-disk by changing the `search_internal`
-setting in `output.yaml` to `True`. The result will then have the search internal available as an attribute,
-irrespective of whether the search is re-run or not.
-"""
+"""
+Searches: DynestyStatic
+=======================
+
+This example illustrates how to use the MCMC ensamble sampler algorithm Zeus.
+
+Information about Zeus can be found at the following links:
+
+ - https://github.com/minaskar/zeus
+ - https://zeus-mcmc.readthedocs.io/en/latest/
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Data**: Loading and plotting the 1D Gaussian dataset used to demonstrate the search.
+- **Model + Analysis**: Setting up the model and analysis for the fitting example.
+- **Search**: Configuring and running the Zeus MCMC sampler.
+- **Result**: Inspecting the result and comparing the maximum log likelihood model to the data.
+- **Search Internal**: Accessing the internal Zeus sampler for advanced use.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import matplotlib.pyplot as plt
+import numpy as np
+from os import path
+
+import autofit as af
+
+"""
+__Data__
+
+This example fits a single 1D Gaussian, we therefore load and plot data containing one Gaussian.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+plt.errorbar(
+ x=range(data.shape[0]),
+ y=data,
+ yerr=noise_map,
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.show()
+plt.close()
+
+"""
+__Model + Analysis__
+
+We create the model and analysis, which in this example is a single `Gaussian` and therefore has dimensionality N=3.
+"""
+model = af.Model(af.ex.Gaussian)
+
+model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.normalization = af.UniformPrior(lower_limit=1e-2, upper_limit=1e2)
+model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+"""
+__Search__
+
+We now create and run the `Zeus` object which acts as our non-linear search.
+
+We manually specify all of the Zeus settings, descriptions of which are provided at the following webpage:
+
+ https://zeus-mcmc.readthedocs.io/en/latest/
+ https://zeus-mcmc.readthedocs.io/en/latest/api/sampler.html
+"""
+search = af.Zeus(
+ path_prefix="searches",
+ name="Zeus",
+ nwalkers=30,
+ nsteps=1001,
+ initializer=af.InitializerBall(lower_limit=0.49, upper_limit=0.51),
+ auto_correlations_settings=af.AutoCorrelationsSettings(
+ check_for_convergence=True,
+ check_size=100,
+ required_length=50,
+ change_threshold=0.01,
+ ),
+ tune=False,
+ tolerance=0.05,
+ patience=5,
+ maxsteps=10000,
+ mu=1.0,
+ maxiter=10000,
+ vectorize=False,
+ check_walkers=True,
+ shuffle_ensemble=True,
+ light_mode=False,
+ iterations_per_full_update=501,
+ number_of_cores=1,
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+"""
+__Result__
+
+The result object returned by the fit provides information on the results of the non-linear search. Lets use it to
+compare the maximum log likelihood `Gaussian` to the data.
+"""
+model_data = result.max_log_likelihood_instance.model_data_from(
+ xvalues=np.arange(data.shape[0])
+)
+
+plt.errorbar(
+ x=range(data.shape[0]),
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(range(data.shape[0]), model_data, color="r")
+plt.title("DynestyStatic model fit to 1D Gaussian dataset.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+__Search Internal__
+
+The result also contains the internal representation of the non-linear search.
+
+The internal representation of the non-linear search ensures that all sampling info is available in its native form.
+This can be passed to functions which take it as input, for example if the sampling package has bespoke visualization
+functions.
+
+For `Emcee`, this is an instance of the `Sampler` object (`from zeus import EnsembleSampler`).
+"""
+search_internal = result.search_internal
+
+print(search_internal)
+
+"""
+The internal search is by default not saved to hard-disk, because it can often take up quite a lot of hard-disk space
+(significantly more than standard output files).
+
+This means that the search internal will only be available the first time you run the search. If you rerun the code
+and the search is bypassed because the results already exist on hard-disk, the search internal will not be available.
+
+If you are frequently using the search internal you can have it saved to hard-disk by changing the `search_internal`
+setting in `output.yaml` to `True`. The result will then have the search internal available as an attribute,
+irrespective of whether the search is re-run or not.
+"""
diff --git a/scripts/searches/mle/Drawer.py b/scripts/searches/mle/Drawer.py
index 9bbb9c2d..3d10ca3e 100644
--- a/scripts/searches/mle/Drawer.py
+++ b/scripts/searches/mle/Drawer.py
@@ -1,138 +1,134 @@
-"""
-Searches: Drawer
-================
-
-This example illustrates how to use the Drawer search, simply draws a fixed number of samples from the model uniformly
-from the priors.
-
-Therefore, it does not seek to determine model parameters which maximize the likelihood or map out the
-posterior of the overall parameter space.
-
-Whilst this is not the typical use case of a non-linear search, it has certain niche applications, for example:
-
- - Given a model one can determine how much variation there is in the log likelihood / log posterior values.
- By visualizing this as a histogram one can therefore quantify the behaviour of that
- model's `log_likelihood_function`.
-
- - If the `log_likelihood_function` of a model is stochastic (e.g. different values of likelihood may be
- computed for an identical model due to randomness in the likelihood evaluation) this search can quantify
- the behaviour of that stochasticity.
-
- - For advanced modeling tools, for example sensitivity mapping performed via the `Sensitivity` object,
- the `Drawer` search may be sufficient to perform the overall modeling task, without the need of performing
- an actual parameter space search.
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Data**: Loading and plotting the 1D Gaussian dataset used to demonstrate the search.
-- **Model + Analysis**: Setting up the model and analysis for the fitting example.
-- **Search**: Configuring and running the Drawer search to draw samples from the priors.
-- **Result**: Inspecting the result and comparing the maximum log likelihood model to the data.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import matplotlib.pyplot as plt
-import numpy as np
-from os import path
-
-import autofit as af
-
-"""
-__Data__
-
-This example fits a single 1D Gaussian, we therefore load and plot data containing one Gaussian.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-plt.errorbar(
- x=range(data.shape[0]),
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.show()
-plt.close()
-
-"""
-__Model + Analysis__
-
-We create the model and analysis, which in this example is a single `Gaussian` and therefore has dimensionality N=3.
-"""
-model = af.Model(af.ex.Gaussian)
-
-model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.normalization = af.UniformPrior(lower_limit=1e-2, upper_limit=1e2)
-model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-"""
-__Search__
-
-We now create and run the `Drawer` object which acts as our non-linear search.
-
-We manually specify all of the Drawer settings, descriptions of which are provided at the following webpage:
-
-https://github.com/rhayes777/PyAutoFit/blob/main/autofit/non_linear/optimize/drawer/drawer.py
-"""
-search = af.Drawer(path_prefix="searches", name="Drawer", total_draws=3)
-
-result = search.fit(model=model, analysis=analysis)
-
-"""
-__Result__
-
-The result object returned by the fit provides information on the results of the non-linear search. Lets use it to
-compare the maximum log likelihood `Gaussian` to the data.
-"""
-model_data = result.max_log_likelihood_instance.model_data_from(
- xvalues=np.arange(data.shape[0])
-)
-
-plt.errorbar(
- x=range(data.shape[0]),
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(range(data.shape[0]), model_data, color="r")
-plt.title("PySwarmsLocal model fit to 1D Gaussian dataset.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
+"""
+Searches: Drawer
+================
+
+This example illustrates how to use the Drawer search, simply draws a fixed number of samples from the model uniformly
+from the priors.
+
+Therefore, it does not seek to determine model parameters which maximize the likelihood or map out the
+posterior of the overall parameter space.
+
+Whilst this is not the typical use case of a non-linear search, it has certain niche applications, for example:
+
+ - Given a model one can determine how much variation there is in the log likelihood / log posterior values.
+ By visualizing this as a histogram one can therefore quantify the behaviour of that
+ model's `log_likelihood_function`.
+
+ - If the `log_likelihood_function` of a model is stochastic (e.g. different values of likelihood may be
+ computed for an identical model due to randomness in the likelihood evaluation) this search can quantify
+ the behaviour of that stochasticity.
+
+ - For advanced modeling tools, for example sensitivity mapping performed via the `Sensitivity` object,
+ the `Drawer` search may be sufficient to perform the overall modeling task, without the need of performing
+ an actual parameter space search.
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Data**: Loading and plotting the 1D Gaussian dataset used to demonstrate the search.
+- **Model + Analysis**: Setting up the model and analysis for the fitting example.
+- **Search**: Configuring and running the Drawer search to draw samples from the priors.
+- **Result**: Inspecting the result and comparing the maximum log likelihood model to the data.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import matplotlib.pyplot as plt
+import numpy as np
+from os import path
+
+import autofit as af
+
+"""
+__Data__
+
+This example fits a single 1D Gaussian, we therefore load and plot data containing one Gaussian.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+plt.errorbar(
+ x=range(data.shape[0]),
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.show()
+plt.close()
+
+"""
+__Model + Analysis__
+
+We create the model and analysis, which in this example is a single `Gaussian` and therefore has dimensionality N=3.
+"""
+model = af.Model(af.ex.Gaussian)
+
+model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.normalization = af.UniformPrior(lower_limit=1e-2, upper_limit=1e2)
+model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+"""
+__Search__
+
+We now create and run the `Drawer` object which acts as our non-linear search.
+
+We manually specify all of the Drawer settings, descriptions of which are provided at the following webpage:
+
+https://github.com/rhayes777/PyAutoFit/blob/main/autofit/non_linear/optimize/drawer/drawer.py
+"""
+search = af.Drawer(path_prefix="searches", name="Drawer", total_draws=3)
+
+result = search.fit(model=model, analysis=analysis)
+
+"""
+__Result__
+
+The result object returned by the fit provides information on the results of the non-linear search. Lets use it to
+compare the maximum log likelihood `Gaussian` to the data.
+"""
+model_data = result.max_log_likelihood_instance.model_data_from(
+ xvalues=np.arange(data.shape[0])
+)
+
+plt.errorbar(
+ x=range(data.shape[0]),
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(range(data.shape[0]), model_data, color="r")
+plt.title("PySwarmsLocal model fit to 1D Gaussian dataset.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
diff --git a/scripts/searches/mle/LBFGS.py b/scripts/searches/mle/LBFGS.py
index 2caec90f..d94c577c 100644
--- a/scripts/searches/mle/LBFGS.py
+++ b/scripts/searches/mle/LBFGS.py
@@ -1,166 +1,162 @@
-"""
-Searches: LBFGS
-===============
-
-This example illustrates how to use the scipy optimize.minimize algorithm L-BFGS.
-
-Information about the L-BFGS method can be found at the following links:
-
- - https://docs.scipy.org/doc/scipy/reference/optimize.minimize-lbfgsb.html
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Data**: Loading and plotting the 1D Gaussian dataset used to demonstrate the search.
-- **Model + Analysis**: Setting up the model and analysis for the fitting example.
-- **Search**: Configuring and running the L-BFGS optimization algorithm.
-- **Result**: Inspecting the result and comparing the maximum log likelihood model to the data.
-- **Search Internal**: Accessing the internal scipy optimizer for advanced use.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import matplotlib.pyplot as plt
-import numpy as np
-from os import path
-
-import autofit as af
-
-"""
-__Data__
-
-This example fits a single 1D Gaussian, we therefore load and plot data containing one Gaussian.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-plt.errorbar(
- x=range(data.shape[0]),
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.show()
-plt.close()
-
-"""
-__Model + Analysis__
-
-We create the model and analysis, which in this example is a single `Gaussian` and therefore has dimensionality N=3.
-"""
-model = af.Model(af.ex.Gaussian)
-
-model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.normalization = af.UniformPrior(lower_limit=1e-2, upper_limit=1e2)
-model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map, use_jax=False)
-
-"""
-__Search__
-
-We now create and run the `LBFGS` object which acts as our non-linear search.
-
-We manually specify all of the LBFGS settings, descriptions of which are provided at the following webpage:
-
- https://docs.scipy.org/doc/scipy/reference/optimize.minimize-lbfgsb.html
-"""
-search = af.LBFGS(
- path_prefix="searches",
- name="LBFGS",
- tol=None,
- disp=None,
- maxcor=10,
- ftol=2.220446049250313e-09,
- gtol=1e-05,
- eps=1e-08,
- maxfun=15000,
- maxiter=15000,
- iprint=-1,
- maxls=20,
- iterations_per_full_update=1000,
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-"""
-__Result__
-
-The result object returned by the fit provides information on the results of the non-linear search. Lets use it to
-compare the maximum log likelihood `Gaussian` to the data.
-"""
-model_data = result.max_log_likelihood_instance.model_data_from(
- xvalues=np.arange(data.shape[0])
-)
-
-plt.errorbar(
- x=range(data.shape[0]),
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(range(data.shape[0]), model_data, color="r")
-plt.title("PySwarmsLocal model fit to 1D Gaussian dataset.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-__Search Internal__
-
-The result also contains the internal representation of the non-linear search.
-
-The internal representation of the non-linear search ensures that all sampling info is available in its native form.
-This can be passed to functions which take it as input, for example if the sampling package has bespoke visualization
-functions.
-
-For `PySwarms`, this is an instance of the `Sampler` object (`from pyswarms import GlobalBestPSO`).
-"""
-search_internal = result.search_internal
-
-print(search_internal)
-
-"""
-The internal search is by default not saved to hard-disk, because it can often take up quite a lot of hard-disk space
-(significantly more than standard output files).
-
-This means that the search internal will only be available the first time you run the search. If you rerun the code
-and the search is bypassed because the results already exist on hard-disk, the search internal will not be available.
-
-If you are frequently using the search internal you can have it saved to hard-disk by changing the `search_internal`
-setting in `output.yaml` to `True`. The result will then have the search internal available as an attribute,
-irrespective of whether the search is re-run or not.
-"""
+"""
+Searches: LBFGS
+===============
+
+This example illustrates how to use the scipy optimize.minimize algorithm L-BFGS.
+
+Information about the L-BFGS method can be found at the following links:
+
+ - https://docs.scipy.org/doc/scipy/reference/optimize.minimize-lbfgsb.html
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Data**: Loading and plotting the 1D Gaussian dataset used to demonstrate the search.
+- **Model + Analysis**: Setting up the model and analysis for the fitting example.
+- **Search**: Configuring and running the L-BFGS optimization algorithm.
+- **Result**: Inspecting the result and comparing the maximum log likelihood model to the data.
+- **Search Internal**: Accessing the internal scipy optimizer for advanced use.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import matplotlib.pyplot as plt
+import numpy as np
+from os import path
+
+import autofit as af
+
+"""
+__Data__
+
+This example fits a single 1D Gaussian, we therefore load and plot data containing one Gaussian.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+plt.errorbar(
+ x=range(data.shape[0]),
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.show()
+plt.close()
+
+"""
+__Model + Analysis__
+
+We create the model and analysis, which in this example is a single `Gaussian` and therefore has dimensionality N=3.
+"""
+model = af.Model(af.ex.Gaussian)
+
+model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.normalization = af.UniformPrior(lower_limit=1e-2, upper_limit=1e2)
+model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map, use_jax=False)
+
+"""
+__Search__
+
+We now create and run the `LBFGS` object which acts as our non-linear search.
+
+We manually specify all of the LBFGS settings, descriptions of which are provided at the following webpage:
+
+ https://docs.scipy.org/doc/scipy/reference/optimize.minimize-lbfgsb.html
+"""
+search = af.LBFGS(
+ path_prefix="searches",
+ name="LBFGS",
+ tol=None,
+ disp=None,
+ maxcor=10,
+ ftol=2.220446049250313e-09,
+ gtol=1e-05,
+ eps=1e-08,
+ maxfun=15000,
+ maxiter=15000,
+ iprint=-1,
+ maxls=20,
+ iterations_per_full_update=1000,
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+"""
+__Result__
+
+The result object returned by the fit provides information on the results of the non-linear search. Lets use it to
+compare the maximum log likelihood `Gaussian` to the data.
+"""
+model_data = result.max_log_likelihood_instance.model_data_from(
+ xvalues=np.arange(data.shape[0])
+)
+
+plt.errorbar(
+ x=range(data.shape[0]),
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(range(data.shape[0]), model_data, color="r")
+plt.title("PySwarmsLocal model fit to 1D Gaussian dataset.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+__Search Internal__
+
+The result also contains the internal representation of the non-linear search.
+
+The internal representation of the non-linear search ensures that all sampling info is available in its native form.
+This can be passed to functions which take it as input, for example if the sampling package has bespoke visualization
+functions.
+
+For `PySwarms`, this is an instance of the `Sampler` object (`from pyswarms import GlobalBestPSO`).
+"""
+search_internal = result.search_internal
+
+print(search_internal)
+
+"""
+The internal search is by default not saved to hard-disk, because it can often take up quite a lot of hard-disk space
+(significantly more than standard output files).
+
+This means that the search internal will only be available the first time you run the search. If you rerun the code
+and the search is bypassed because the results already exist on hard-disk, the search internal will not be available.
+
+If you are frequently using the search internal you can have it saved to hard-disk by changing the `search_internal`
+setting in `output.yaml` to `True`. The result will then have the search internal available as an attribute,
+irrespective of whether the search is re-run or not.
+"""
diff --git a/scripts/searches/mle/PySwarmsGlobal.py b/scripts/searches/mle/PySwarmsGlobal.py
index d43cc1ff..a4fc13bb 100644
--- a/scripts/searches/mle/PySwarmsGlobal.py
+++ b/scripts/searches/mle/PySwarmsGlobal.py
@@ -1,165 +1,161 @@
-"""
-Searches: PySwarmsGlobal
-========================
-
-This example illustrates how to use the particle swarm optimization algorithm PySwarmsGlobal.
-
-Information about PySwarms can be found at the following links:
-
- - https://github.com/ljvmiranda921/pyswarms
- - https://pyswarms.readthedocs.io/en/latest/index.html
- - https://pyswarms.readthedocs.io/en/latest/api/pyswarms.single.html#module-pyswarms.single.global_best
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Data**: Loading and plotting the 1D Gaussian dataset used to demonstrate the search.
-- **Model + Analysis**: Setting up the model and analysis for the fitting example.
-- **Search**: Configuring and running the PySwarmsGlobal particle swarm optimizer.
-- **Result**: Inspecting the result and comparing the maximum log likelihood model to the data.
-- **Search Internal**: Accessing the internal PySwarms optimizer for advanced use.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import matplotlib.pyplot as plt
-import numpy as np
-from os import path
-
-import autofit as af
-
-"""
-__Data__
-
-This example fits a single 1D Gaussian, we therefore load and plot data containing one Gaussian.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-plt.errorbar(
- x=range(data.shape[0]),
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.show()
-plt.close()
-
-"""
-__Model + Analysis__
-
-We create the model and analysis, which in this example is a single `Gaussian` and therefore has dimensionality N=3.
-"""
-model = af.Model(af.ex.Gaussian)
-
-model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.normalization = af.UniformPrior(lower_limit=1e-2, upper_limit=1e2)
-model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-"""
-__Search__
-
-We now create and run the `PySwarmsGlobal` object which acts as our non-linear search.
-
-We manually specify all of the PySwarms settings, descriptions of which are provided at the following webpage:
-
- https://pyswarms.readthedocs.io/en/latest/api/pyswarms.single.html#module-pyswarms.single.global_best
-"""
-search = af.PySwarmsGlobal(
- path_prefix="searches",
- name="PySwarmsGlobal",
- n_particles=50,
- iters=1000,
- cognitive=0.5,
- social=0.3,
- inertia=0.9,
- ftol=-np.inf,
- iterations_per_full_update=100,
- number_of_cores=1,
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-"""
-__Result__
-
-The result object returned by the fit provides information on the results of the non-linear search. Lets use it to
-compare the maximum log likelihood `Gaussian` to the data.
-"""
-model_data = result.max_log_likelihood_instance.model_data_from(
- xvalues=np.arange(data.shape[0])
-)
-
-plt.errorbar(
- x=range(data.shape[0]),
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(range(data.shape[0]), model_data, color="r")
-plt.title("PySwarmsGlobal model fit to 1D Gaussian dataset.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-__Search Internal__
-
-The result also contains the internal representation of the non-linear search.
-
-The internal representation of the non-linear search ensures that all sampling info is available in its native form.
-This can be passed to functions which take it as input, for example if the sampling package has bespoke visualization
-functions.
-
-For `PySwarms`, this is an instance of the `Sampler` object (`from pyswarms import GlobalBestPSO`).
-"""
-search_internal = result.search_internal
-
-print(search_internal)
-
-"""
-The internal search is by default not saved to hard-disk, because it can often take up quite a lot of hard-disk space
-(significantly more than standard output files).
-
-This means that the search internal will only be available the first time you run the search. If you rerun the code
-and the search is bypassed because the results already exist on hard-disk, the search internal will not be available.
-
-If you are frequently using the search internal you can have it saved to hard-disk by changing the `search_internal`
-setting in `output.yaml` to `True`. The result will then have the search internal available as an attribute,
-irrespective of whether the search is re-run or not.
-"""
+"""
+Searches: PySwarmsGlobal
+========================
+
+This example illustrates how to use the particle swarm optimization algorithm PySwarmsGlobal.
+
+Information about PySwarms can be found at the following links:
+
+ - https://github.com/ljvmiranda921/pyswarms
+ - https://pyswarms.readthedocs.io/en/latest/index.html
+ - https://pyswarms.readthedocs.io/en/latest/api/pyswarms.single.html#module-pyswarms.single.global_best
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Data**: Loading and plotting the 1D Gaussian dataset used to demonstrate the search.
+- **Model + Analysis**: Setting up the model and analysis for the fitting example.
+- **Search**: Configuring and running the PySwarmsGlobal particle swarm optimizer.
+- **Result**: Inspecting the result and comparing the maximum log likelihood model to the data.
+- **Search Internal**: Accessing the internal PySwarms optimizer for advanced use.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import matplotlib.pyplot as plt
+import numpy as np
+from os import path
+
+import autofit as af
+
+"""
+__Data__
+
+This example fits a single 1D Gaussian, we therefore load and plot data containing one Gaussian.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+plt.errorbar(
+ x=range(data.shape[0]),
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.show()
+plt.close()
+
+"""
+__Model + Analysis__
+
+We create the model and analysis, which in this example is a single `Gaussian` and therefore has dimensionality N=3.
+"""
+model = af.Model(af.ex.Gaussian)
+
+model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.normalization = af.UniformPrior(lower_limit=1e-2, upper_limit=1e2)
+model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+"""
+__Search__
+
+We now create and run the `PySwarmsGlobal` object which acts as our non-linear search.
+
+We manually specify all of the PySwarms settings, descriptions of which are provided at the following webpage:
+
+ https://pyswarms.readthedocs.io/en/latest/api/pyswarms.single.html#module-pyswarms.single.global_best
+"""
+search = af.PySwarmsGlobal(
+ path_prefix="searches",
+ name="PySwarmsGlobal",
+ n_particles=50,
+ iters=1000,
+ cognitive=0.5,
+ social=0.3,
+ inertia=0.9,
+ ftol=-np.inf,
+ iterations_per_full_update=100,
+ number_of_cores=1,
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+"""
+__Result__
+
+The result object returned by the fit provides information on the results of the non-linear search. Lets use it to
+compare the maximum log likelihood `Gaussian` to the data.
+"""
+model_data = result.max_log_likelihood_instance.model_data_from(
+ xvalues=np.arange(data.shape[0])
+)
+
+plt.errorbar(
+ x=range(data.shape[0]),
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(range(data.shape[0]), model_data, color="r")
+plt.title("PySwarmsGlobal model fit to 1D Gaussian dataset.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+__Search Internal__
+
+The result also contains the internal representation of the non-linear search.
+
+The internal representation of the non-linear search ensures that all sampling info is available in its native form.
+This can be passed to functions which take it as input, for example if the sampling package has bespoke visualization
+functions.
+
+For `PySwarms`, this is an instance of the `Sampler` object (`from pyswarms import GlobalBestPSO`).
+"""
+search_internal = result.search_internal
+
+print(search_internal)
+
+"""
+The internal search is by default not saved to hard-disk, because it can often take up quite a lot of hard-disk space
+(significantly more than standard output files).
+
+This means that the search internal will only be available the first time you run the search. If you rerun the code
+and the search is bypassed because the results already exist on hard-disk, the search internal will not be available.
+
+If you are frequently using the search internal you can have it saved to hard-disk by changing the `search_internal`
+setting in `output.yaml` to `True`. The result will then have the search internal available as an attribute,
+irrespective of whether the search is re-run or not.
+"""
diff --git a/scripts/searches/mle/PySwarmsLocal.py b/scripts/searches/mle/PySwarmsLocal.py
index ced238f0..efeea756 100644
--- a/scripts/searches/mle/PySwarmsLocal.py
+++ b/scripts/searches/mle/PySwarmsLocal.py
@@ -1,167 +1,163 @@
-"""
-Searches: PySwarmsLocal
-========================
-
-This example illustrates how to use the particle swarm optimization algorithm PySwarmsLocal.
-
-Information about PySwarms can be found at the following links:
-
- - https://github.com/ljvmiranda921/pyswarms
- - https://pyswarms.readthedocs.io/en/latest/index.html
- - https://pyswarms.readthedocs.io/en/latest/api/pyswarms.single.html#module-pyswarms.single.local_best
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Data**: Loading and plotting the 1D Gaussian dataset used to demonstrate the search.
-- **Model + Analysis**: Setting up the model and analysis for the fitting example.
-- **Search**: Configuring and running the PySwarmsLocal particle swarm optimizer.
-- **Result**: Inspecting the result and comparing the maximum log likelihood model to the data.
-- **Search Internal**: Accessing the internal PySwarms optimizer for advanced use.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import matplotlib.pyplot as plt
-import numpy as np
-from os import path
-
-import autofit as af
-
-"""
-__Data__
-
-This example fits a single 1D Gaussian, we therefore load and plot data containing one Gaussian.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-plt.errorbar(
- x=range(data.shape[0]),
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.show()
-plt.close()
-
-"""
-__Model + Analysis__
-
-We create the model and analysis, which in this example is a single `Gaussian` and therefore has dimensionality N=3.
-"""
-model = af.Model(af.ex.Gaussian)
-
-model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.normalization = af.UniformPrior(lower_limit=1e-2, upper_limit=1e2)
-model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-"""
-__Search__
-
-We now create and run the `PySwarmsLocal` object which acts as our non-linear search.
-
-We manually specify all of the PySwarms settings, descriptions of which are provided at the following webpage:
-
- https://pyswarms.readthedocs.io/en/latest/api/pyswarms.single.html#module-pyswarms.single.local_best
-"""
-search = af.PySwarmsLocal(
- path_prefix="searches",
- name="PySwarmsLocal",
- n_particles=50,
- iters=1000,
- cognitive=0.5,
- social=0.3,
- inertia=0.9,
- number_of_k_neighbors=3,
- minkowski_p_norm=2,
- ftol=-np.inf,
- iterations_per_full_update=1000,
- number_of_cores=1,
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-"""
-__Result__
-
-The result object returned by the fit provides information on the results of the non-linear search. Lets use it to
-compare the maximum log likelihood `Gaussian` to the data.
-"""
-model_data = result.max_log_likelihood_instance.model_data_from(
- xvalues=np.arange(data.shape[0])
-)
-
-plt.errorbar(
- x=range(data.shape[0]),
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(range(data.shape[0]), model_data, color="r")
-plt.title("PySwarmsLocal model fit to 1D Gaussian dataset.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-__Search Internal__
-
-The result also contains the internal representation of the non-linear search.
-
-The internal representation of the non-linear search ensures that all sampling info is available in its native form.
-This can be passed to functions which take it as input, for example if the sampling package has bespoke visualization
-functions.
-
-For `PySwarms`, this is an instance of the `Sampler` object (`from pyswarms import LocalBestPSO`).
-"""
-search_internal = result.search_internal
-
-print(search_internal)
-
-"""
-The internal search is by default not saved to hard-disk, because it can often take up quite a lot of hard-disk space
-(significantly more than standard output files).
-
-This means that the search internal will only be available the first time you run the search. If you rerun the code
-and the search is bypassed because the results already exist on hard-disk, the search internal will not be available.
-
-If you are frequently using the search internal you can have it saved to hard-disk by changing the `search_internal`
-setting in `output.yaml` to `True`. The result will then have the search internal available as an attribute,
-irrespective of whether the search is re-run or not.
-"""
+"""
+Searches: PySwarmsLocal
+========================
+
+This example illustrates how to use the particle swarm optimization algorithm PySwarmsLocal.
+
+Information about PySwarms can be found at the following links:
+
+ - https://github.com/ljvmiranda921/pyswarms
+ - https://pyswarms.readthedocs.io/en/latest/index.html
+ - https://pyswarms.readthedocs.io/en/latest/api/pyswarms.single.html#module-pyswarms.single.local_best
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Data**: Loading and plotting the 1D Gaussian dataset used to demonstrate the search.
+- **Model + Analysis**: Setting up the model and analysis for the fitting example.
+- **Search**: Configuring and running the PySwarmsLocal particle swarm optimizer.
+- **Result**: Inspecting the result and comparing the maximum log likelihood model to the data.
+- **Search Internal**: Accessing the internal PySwarms optimizer for advanced use.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import matplotlib.pyplot as plt
+import numpy as np
+from os import path
+
+import autofit as af
+
+"""
+__Data__
+
+This example fits a single 1D Gaussian, we therefore load and plot data containing one Gaussian.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+plt.errorbar(
+ x=range(data.shape[0]),
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.show()
+plt.close()
+
+"""
+__Model + Analysis__
+
+We create the model and analysis, which in this example is a single `Gaussian` and therefore has dimensionality N=3.
+"""
+model = af.Model(af.ex.Gaussian)
+
+model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.normalization = af.UniformPrior(lower_limit=1e-2, upper_limit=1e2)
+model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+"""
+__Search__
+
+We now create and run the `PySwarmsLocal` object which acts as our non-linear search.
+
+We manually specify all of the PySwarms settings, descriptions of which are provided at the following webpage:
+
+ https://pyswarms.readthedocs.io/en/latest/api/pyswarms.single.html#module-pyswarms.single.local_best
+"""
+search = af.PySwarmsLocal(
+ path_prefix="searches",
+ name="PySwarmsLocal",
+ n_particles=50,
+ iters=1000,
+ cognitive=0.5,
+ social=0.3,
+ inertia=0.9,
+ number_of_k_neighbors=3,
+ minkowski_p_norm=2,
+ ftol=-np.inf,
+ iterations_per_full_update=1000,
+ number_of_cores=1,
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+"""
+__Result__
+
+The result object returned by the fit provides information on the results of the non-linear search. Lets use it to
+compare the maximum log likelihood `Gaussian` to the data.
+"""
+model_data = result.max_log_likelihood_instance.model_data_from(
+ xvalues=np.arange(data.shape[0])
+)
+
+plt.errorbar(
+ x=range(data.shape[0]),
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(range(data.shape[0]), model_data, color="r")
+plt.title("PySwarmsLocal model fit to 1D Gaussian dataset.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+__Search Internal__
+
+The result also contains the internal representation of the non-linear search.
+
+The internal representation of the non-linear search ensures that all sampling info is available in its native form.
+This can be passed to functions which take it as input, for example if the sampling package has bespoke visualization
+functions.
+
+For `PySwarms`, this is an instance of the `Sampler` object (`from pyswarms import LocalBestPSO`).
+"""
+search_internal = result.search_internal
+
+print(search_internal)
+
+"""
+The internal search is by default not saved to hard-disk, because it can often take up quite a lot of hard-disk space
+(significantly more than standard output files).
+
+This means that the search internal will only be available the first time you run the search. If you rerun the code
+and the search is bypassed because the results already exist on hard-disk, the search internal will not be available.
+
+If you are frequently using the search internal you can have it saved to hard-disk by changing the `search_internal`
+setting in `output.yaml` to `True`. The result will then have the search internal available as an attribute,
+irrespective of whether the search is re-run or not.
+"""
diff --git a/scripts/searches/nest/DynestyDynamic.py b/scripts/searches/nest/DynestyDynamic.py
index 966eea57..315eccce 100644
--- a/scripts/searches/nest/DynestyDynamic.py
+++ b/scripts/searches/nest/DynestyDynamic.py
@@ -1,169 +1,165 @@
-"""
-Searches: DynestyDynamic
-=======================
-
-This example illustrates how to use the nested sampling algorithm DynestyDynamic.
-
-Information about Dynesty can be found at the following links:
-
- - https://github.com/joshspeagle/dynesty
- - https://dynesty.readthedocs.io/en/latest/
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Data**: Loading and plotting the 1D Gaussian dataset used to demonstrate the search.
-- **Model + Analysis**: Setting up the model and analysis for the fitting example.
-- **Search**: Configuring and running the DynestyDynamic nested sampler.
-- **Result**: Inspecting the result and comparing the maximum log likelihood model to the data.
-- **Search Internal**: Accessing the internal Dynesty sampler for advanced use.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import matplotlib.pyplot as plt
-import numpy as np
-from os import path
-
-import autofit as af
-
-"""
-__Data__
-
-This example fits a single 1D Gaussian, we therefore load and plot data containing one Gaussian.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-plt.errorbar(
- x=range(data.shape[0]),
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.show()
-plt.close()
-
-"""
-__Model + Analysis__
-
-We create the model and analysis, which in this example is a single `Gaussian` and therefore has dimensionality N=3.
-"""
-model = af.Model(af.ex.Gaussian)
-
-model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.normalization = af.LogUniformPrior(lower_limit=1e-2, upper_limit=1e2)
-model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-"""
-__Search__
-
-We now create and run the `DynestyDynamic` object which acts as our non-linear search.
-
-We manually specify all of the Dynesty settings, descriptions of which are provided at the following webpage:
-
- https://dynesty.readthedocs.io/en/latest/api.html
- https://dynesty.readthedocs.io/en/latest/api.html#module-dynesty.dynamicsampler
-"""
-search = af.DynestyDynamic(
- path_prefix="searches",
- name="DynestyDynamic",
- nlive=50,
- bound="multi",
- sample="auto",
- bootstrap=None,
- enlarge=None,
- update_interval=None,
- walks=25,
- facc=0.5,
- slices=5,
- fmove=0.9,
- max_move=100,
- number_of_cores=1,
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-"""
-__Result__
-
-The result object returned by the fit provides information on the results of the non-linear search. Lets use it to
-compare the maximum log likelihood `Gaussian` to the data.
-"""
-model_data = result.max_log_likelihood_instance.model_data_from(
- xvalues=np.arange(data.shape[0])
-)
-
-plt.errorbar(
- x=range(data.shape[0]),
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(range(data.shape[0]), model_data, color="r")
-plt.title("DynestyDynamic model fit to 1D Gaussian dataset.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-__Search Internal__
-
-The result also contains the internal representation of the non-linear search.
-
-The internal representation of the non-linear search ensures that all sampling info is available in its native form.
-This can be passed to functions which take it as input, for example if the sampling package has bespoke visualization
-functions.
-
-For `DynestyStatic`, this is an instance of the `NestedSampler` object (`from dynesty import DynamicNestedSampler`).
-"""
-search_internal = result.search_internal
-
-print(search_internal)
-
-"""
-The internal search is by default not saved to hard-disk, because it can often take up quite a lot of hard-disk space
-(significantly more than standard output files).
-
-This means that the search internal will only be available the first time you run the search. If you rerun the code
-and the search is bypassed because the results already exist on hard-disk, the search internal will not be available.
-
-If you are frequently using the search internal you can have it saved to hard-disk by changing the `search_internal`
-setting in `output.yaml` to `True`. The result will then have the search internal available as an attribute,
-irrespective of whether the search is re-run or not.
-"""
+"""
+Searches: DynestyDynamic
+=======================
+
+This example illustrates how to use the nested sampling algorithm DynestyDynamic.
+
+Information about Dynesty can be found at the following links:
+
+ - https://github.com/joshspeagle/dynesty
+ - https://dynesty.readthedocs.io/en/latest/
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Data**: Loading and plotting the 1D Gaussian dataset used to demonstrate the search.
+- **Model + Analysis**: Setting up the model and analysis for the fitting example.
+- **Search**: Configuring and running the DynestyDynamic nested sampler.
+- **Result**: Inspecting the result and comparing the maximum log likelihood model to the data.
+- **Search Internal**: Accessing the internal Dynesty sampler for advanced use.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import matplotlib.pyplot as plt
+import numpy as np
+from os import path
+
+import autofit as af
+
+"""
+__Data__
+
+This example fits a single 1D Gaussian, we therefore load and plot data containing one Gaussian.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+plt.errorbar(
+ x=range(data.shape[0]),
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.show()
+plt.close()
+
+"""
+__Model + Analysis__
+
+We create the model and analysis, which in this example is a single `Gaussian` and therefore has dimensionality N=3.
+"""
+model = af.Model(af.ex.Gaussian)
+
+model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.normalization = af.LogUniformPrior(lower_limit=1e-2, upper_limit=1e2)
+model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+"""
+__Search__
+
+We now create and run the `DynestyDynamic` object which acts as our non-linear search.
+
+We manually specify all of the Dynesty settings, descriptions of which are provided at the following webpage:
+
+ https://dynesty.readthedocs.io/en/latest/api.html
+ https://dynesty.readthedocs.io/en/latest/api.html#module-dynesty.dynamicsampler
+"""
+search = af.DynestyDynamic(
+ path_prefix="searches",
+ name="DynestyDynamic",
+ nlive=50,
+ bound="multi",
+ sample="auto",
+ bootstrap=None,
+ enlarge=None,
+ update_interval=None,
+ walks=25,
+ facc=0.5,
+ slices=5,
+ fmove=0.9,
+ max_move=100,
+ number_of_cores=1,
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+"""
+__Result__
+
+The result object returned by the fit provides information on the results of the non-linear search. Lets use it to
+compare the maximum log likelihood `Gaussian` to the data.
+"""
+model_data = result.max_log_likelihood_instance.model_data_from(
+ xvalues=np.arange(data.shape[0])
+)
+
+plt.errorbar(
+ x=range(data.shape[0]),
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(range(data.shape[0]), model_data, color="r")
+plt.title("DynestyDynamic model fit to 1D Gaussian dataset.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+__Search Internal__
+
+The result also contains the internal representation of the non-linear search.
+
+The internal representation of the non-linear search ensures that all sampling info is available in its native form.
+This can be passed to functions which take it as input, for example if the sampling package has bespoke visualization
+functions.
+
+For `DynestyStatic`, this is an instance of the `NestedSampler` object (`from dynesty import DynamicNestedSampler`).
+"""
+search_internal = result.search_internal
+
+print(search_internal)
+
+"""
+The internal search is by default not saved to hard-disk, because it can often take up quite a lot of hard-disk space
+(significantly more than standard output files).
+
+This means that the search internal will only be available the first time you run the search. If you rerun the code
+and the search is bypassed because the results already exist on hard-disk, the search internal will not be available.
+
+If you are frequently using the search internal you can have it saved to hard-disk by changing the `search_internal`
+setting in `output.yaml` to `True`. The result will then have the search internal available as an attribute,
+irrespective of whether the search is re-run or not.
+"""
diff --git a/scripts/searches/nest/DynestyStatic.py b/scripts/searches/nest/DynestyStatic.py
index d303750b..d2071b4d 100644
--- a/scripts/searches/nest/DynestyStatic.py
+++ b/scripts/searches/nest/DynestyStatic.py
@@ -1,171 +1,167 @@
-"""
-Searches: DynestyStatic
-=======================
-
-This example illustrates how to use the nested sampling algorithm DynestyStatic.
-
-Information about Dynesty can be found at the following links:
-
- - https://github.com/joshspeagle/dynesty
- - https://dynesty.readthedocs.io/en/latest/
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Data**: Loading and plotting the 1D Gaussian dataset used to demonstrate the search.
-- **Model + Analysis**: Setting up the model and analysis for the fitting example.
-- **Search**: Configuring and running the DynestyStatic nested sampler.
-- **Result**: Inspecting the result and comparing the maximum log likelihood model to the data.
-- **Search Internal**: Accessing the internal Dynesty sampler for advanced use.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import matplotlib.pyplot as plt
-import numpy as np
-from os import path
-
-import autofit as af
-
-"""
-__Data__
-
-This example fits a single 1D Gaussian, we therefore load and plot data containing one Gaussian.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-plt.errorbar(
- x=range(data.shape[0]),
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.show()
-plt.close()
-
-"""
-__Model + Analysis__
-
-We create the model and analysis, which in this example is a single `Gaussian` and therefore has dimensionality N=3.
-"""
-model = af.Model(af.ex.Gaussian)
-
-model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.normalization = af.LogUniformPrior(lower_limit=1e-2, upper_limit=1e2)
-model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-"""
-__Search__
-
-We now create and run the `DynestyStatic` object which acts as our non-linear search.
-
-We manually specify all of the Dynesty settings, descriptions of which are provided at the following webpage:
-
- https://dynesty.readthedocs.io/en/latest/api.html
- https://dynesty.readthedocs.io/en/latest/api.html#module-dynesty.nestedsamplers
-"""
-search = af.DynestyStatic(
- path_prefix=path.join("searches"),
- name="DynestyStatic",
- nlive=50,
- bound="multi",
- sample="auto",
- bootstrap=None,
- enlarge=None,
- update_interval=None,
- walks=25,
- facc=0.5,
- slices=5,
- fmove=0.9,
- max_move=100,
- iterations_per_full_update=2500,
- number_of_cores=1,
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-"""
-__Result__
-
-The result object returned by the fit provides information on the results of the non-linear search. Lets use it to
-compare the maximum log likelihood `Gaussian` to the data.
-"""
-model_data = result.max_log_likelihood_instance.model_data_from(
- xvalues=np.arange(data.shape[0])
-)
-
-plt.errorbar(
- x=range(data.shape[0]),
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(range(data.shape[0]), model_data, color="r")
-plt.title("DynestyStatic model fit to 1D Gaussian dataset.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-
-"""
-__Search Internal__
-
-The result also contains the internal representation of the non-linear search.
-
-The internal representation of the non-linear search ensures that all sampling info is available in its native form.
-This can be passed to functions which take it as input, for example if the sampling package has bespoke visualization
-functions.
-
-For `DynestyStatic`, this is an instance of the `NestedSampler` object (`from dynesty import NestedSampler`).
-"""
-search_internal = result.search_internal
-
-print(search_internal)
-
-"""
-The internal search is by default not saved to hard-disk, because it can often take up quite a lot of hard-disk space
-(significantly more than standard output files).
-
-This means that the search internal will only be available the first time you run the search. If you rerun the code
-and the search is bypassed because the results already exist on hard-disk, the search internal will not be available.
-
-If you are frequently using the search internal you can have it saved to hard-disk by changing the `search_internal`
-setting in `output.yaml` to `True`. The result will then have the search internal available as an attribute,
-irrespective of whether the search is re-run or not.
-"""
+"""
+Searches: DynestyStatic
+=======================
+
+This example illustrates how to use the nested sampling algorithm DynestyStatic.
+
+Information about Dynesty can be found at the following links:
+
+ - https://github.com/joshspeagle/dynesty
+ - https://dynesty.readthedocs.io/en/latest/
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Data**: Loading and plotting the 1D Gaussian dataset used to demonstrate the search.
+- **Model + Analysis**: Setting up the model and analysis for the fitting example.
+- **Search**: Configuring and running the DynestyStatic nested sampler.
+- **Result**: Inspecting the result and comparing the maximum log likelihood model to the data.
+- **Search Internal**: Accessing the internal Dynesty sampler for advanced use.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import matplotlib.pyplot as plt
+import numpy as np
+from os import path
+
+import autofit as af
+
+"""
+__Data__
+
+This example fits a single 1D Gaussian, we therefore load and plot data containing one Gaussian.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+plt.errorbar(
+ x=range(data.shape[0]),
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.show()
+plt.close()
+
+"""
+__Model + Analysis__
+
+We create the model and analysis, which in this example is a single `Gaussian` and therefore has dimensionality N=3.
+"""
+model = af.Model(af.ex.Gaussian)
+
+model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.normalization = af.LogUniformPrior(lower_limit=1e-2, upper_limit=1e2)
+model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+"""
+__Search__
+
+We now create and run the `DynestyStatic` object which acts as our non-linear search.
+
+We manually specify all of the Dynesty settings, descriptions of which are provided at the following webpage:
+
+ https://dynesty.readthedocs.io/en/latest/api.html
+ https://dynesty.readthedocs.io/en/latest/api.html#module-dynesty.nestedsamplers
+"""
+search = af.DynestyStatic(
+ path_prefix=path.join("searches"),
+ name="DynestyStatic",
+ nlive=50,
+ bound="multi",
+ sample="auto",
+ bootstrap=None,
+ enlarge=None,
+ update_interval=None,
+ walks=25,
+ facc=0.5,
+ slices=5,
+ fmove=0.9,
+ max_move=100,
+ iterations_per_full_update=2500,
+ number_of_cores=1,
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+"""
+__Result__
+
+The result object returned by the fit provides information on the results of the non-linear search. Lets use it to
+compare the maximum log likelihood `Gaussian` to the data.
+"""
+model_data = result.max_log_likelihood_instance.model_data_from(
+ xvalues=np.arange(data.shape[0])
+)
+
+plt.errorbar(
+ x=range(data.shape[0]),
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(range(data.shape[0]), model_data, color="r")
+plt.title("DynestyStatic model fit to 1D Gaussian dataset.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+
+"""
+__Search Internal__
+
+The result also contains the internal representation of the non-linear search.
+
+The internal representation of the non-linear search ensures that all sampling info is available in its native form.
+This can be passed to functions which take it as input, for example if the sampling package has bespoke visualization
+functions.
+
+For `DynestyStatic`, this is an instance of the `NestedSampler` object (`from dynesty import NestedSampler`).
+"""
+search_internal = result.search_internal
+
+print(search_internal)
+
+"""
+The internal search is by default not saved to hard-disk, because it can often take up quite a lot of hard-disk space
+(significantly more than standard output files).
+
+This means that the search internal will only be available the first time you run the search. If you rerun the code
+and the search is bypassed because the results already exist on hard-disk, the search internal will not be available.
+
+If you are frequently using the search internal you can have it saved to hard-disk by changing the `search_internal`
+setting in `output.yaml` to `True`. The result will then have the search internal available as an attribute,
+irrespective of whether the search is re-run or not.
+"""
diff --git a/scripts/searches/nest/Nautilus.py b/scripts/searches/nest/Nautilus.py
index 5a7b1bc7..cdacda7f 100644
--- a/scripts/searches/nest/Nautilus.py
+++ b/scripts/searches/nest/Nautilus.py
@@ -1,173 +1,169 @@
-"""
-Searches=Nautilus
-=======================
-
-This example illustrates how to use the nested sampling algorithm Nautilus.
-
-Information about Nautilus can be found at the following links:
-
- - https://nautilus-sampler.readthedocs.io/en/stable/index.html
- - https://github.com/johannesulf/nautilus
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Data**: Loading and plotting the 1D Gaussian dataset used to demonstrate the search.
-- **Model + Analysis**: Setting up the model and analysis for the fitting example.
-- **Search**: Configuring and running the Nautilus nested sampler.
-- **Result**: Inspecting the result and comparing the maximum log likelihood model to the data.
-- **Search Internal**: Accessing the internal Nautilus sampler for advanced use.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import matplotlib.pyplot as plt
-import numpy as np
-from os import path
-
-import autofit as af
-
-"""
-__Data__
-
-This example fits a single 1D Gaussian, we therefore load and plot data containing one Gaussian.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-plt.errorbar(
- x=range(data.shape[0]),
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.show()
-plt.close()
-
-"""
-__Model + Analysis__
-
-We create the model and analysis, which in this example is a single `Gaussian` and therefore has dimensionality N=3.
-"""
-model = af.Model(af.ex.Gaussian)
-
-model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.normalization = af.LogUniformPrior(lower_limit=1e-2, upper_limit=1e2)
-model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-"""
-__Search__
-
-We now create and run the `Nautilus` object which acts as our non-linear search.
-
-We manually specify all of the Nautilus settings, descriptions of which are provided at the following webpage:
-
-https://github.com/johannesulf/nautilus
-"""
-search = af.Nautilus(
- path_prefix=path.join("searches"),
- name="Nautilus",
- number_of_cores=4,
- n_live=100, # Number of so-called live points. New bounds are constructed so that they encompass the live points.
- n_update=None, # The maximum number of additions to the live set before a new bound is created
- enlarge_per_dim=1.1, # Along each dimension, outer ellipsoidal bounds are enlarged by this factor.
- n_points_min=None, # The minimum number of points each ellipsoid should have. Effectively, ellipsoids with less than twice that number will not be split further.
- split_threshold=100, # Threshold used for splitting the multi-ellipsoidal bound used for sampling.
- n_networks=4, # Number of networks used in the estimator.
- n_batch=100, # Number of likelihood evaluations that are performed at each step. If likelihood evaluations are parallelized, should be multiple of the number of parallel processes.
- n_like_new_bound=None, # The maximum number of likelihood calls before a new bounds is created. If None, use 10 times n_live.
- vectorized=False, # If True, the likelihood function can receive multiple input sets at once.
- seed=None, # Seed for random number generation used for reproducible results accross different runs.
- f_live=0.01, # Maximum fraction of the evidence contained in the live set before building the initial shells terminates.
- n_shell=1, # Minimum number of points in each shell. The algorithm will sample from the shells until this is reached. Default is 1.
- n_eff=500, # Minimum effective sample size. The algorithm will sample from the shells until this is reached. Default is 10000.
- discard_exploration=False, # Whether to discard points drawn in the exploration phase. This is required for a fully unbiased posterior and evidence estimate.
- verbose=True, # Whether to print information about the run.
- n_like_max=np.inf, # Maximum number of likelihood evaluations. Regardless of progress, the sampler will stop if this value is reached. Default is infinity.
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-"""
-__Result__
-
-The result object returned by the fit provides information on the results of the non-linear search. Lets use it to
-compare the maximum log likelihood `Gaussian` to the data.
-"""
-model_data = result.max_log_likelihood_instance.model_data_from(
- xvalues=np.arange(data.shape[0])
-)
-
-plt.errorbar(
- x=range(data.shape[0]),
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(range(data.shape[0]), model_data, color="r")
-plt.title("Nautilus model fit to 1D Gaussian dataset.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
-
-"""
-__Search Internal__
-
-The result also contains the internal representation of the non-linear search.
-
-The internal representation of the non-linear search ensures that all sampling info is available in its native form.
-This can be passed to functions which take it as input, for example if the sampling package has bespoke visualization
-functions.
-
-For `DynestyStatic`, this is an instance of the `Sampler` object (`from nautilus import Sampler`).
-"""
-search_internal = result.search_internal
-
-print(search_internal)
-
-"""
-The internal search is by default not saved to hard-disk, because it can often take up quite a lot of hard-disk space
-(significantly more than standard output files).
-
-This means that the search internal will only be available the first time you run the search. If you rerun the code
-and the search is bypassed because the results already exist on hard-disk, the search internal will not be available.
-
-If you are frequently using the search internal you can have it saved to hard-disk by changing the `search_internal`
-setting in `output.yaml` to `True`. The result will then have the search internal available as an attribute,
-irrespective of whether the search is re-run or not.
-"""
+"""
+Searches=Nautilus
+=======================
+
+This example illustrates how to use the nested sampling algorithm Nautilus.
+
+Information about Nautilus can be found at the following links:
+
+ - https://nautilus-sampler.readthedocs.io/en/stable/index.html
+ - https://github.com/johannesulf/nautilus
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Data**: Loading and plotting the 1D Gaussian dataset used to demonstrate the search.
+- **Model + Analysis**: Setting up the model and analysis for the fitting example.
+- **Search**: Configuring and running the Nautilus nested sampler.
+- **Result**: Inspecting the result and comparing the maximum log likelihood model to the data.
+- **Search Internal**: Accessing the internal Nautilus sampler for advanced use.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import matplotlib.pyplot as plt
+import numpy as np
+from os import path
+
+import autofit as af
+
+"""
+__Data__
+
+This example fits a single 1D Gaussian, we therefore load and plot data containing one Gaussian.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+plt.errorbar(
+ x=range(data.shape[0]),
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.show()
+plt.close()
+
+"""
+__Model + Analysis__
+
+We create the model and analysis, which in this example is a single `Gaussian` and therefore has dimensionality N=3.
+"""
+model = af.Model(af.ex.Gaussian)
+
+model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.normalization = af.LogUniformPrior(lower_limit=1e-2, upper_limit=1e2)
+model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+"""
+__Search__
+
+We now create and run the `Nautilus` object which acts as our non-linear search.
+
+We manually specify all of the Nautilus settings, descriptions of which are provided at the following webpage:
+
+https://github.com/johannesulf/nautilus
+"""
+search = af.Nautilus(
+ path_prefix=path.join("searches"),
+ name="Nautilus",
+ number_of_cores=4,
+ n_live=100, # Number of so-called live points. New bounds are constructed so that they encompass the live points.
+ n_update=None, # The maximum number of additions to the live set before a new bound is created
+ enlarge_per_dim=1.1, # Along each dimension, outer ellipsoidal bounds are enlarged by this factor.
+ n_points_min=None, # The minimum number of points each ellipsoid should have. Effectively, ellipsoids with less than twice that number will not be split further.
+ split_threshold=100, # Threshold used for splitting the multi-ellipsoidal bound used for sampling.
+ n_networks=4, # Number of networks used in the estimator.
+ n_batch=100, # Number of likelihood evaluations that are performed at each step. If likelihood evaluations are parallelized, should be multiple of the number of parallel processes.
+ n_like_new_bound=None, # The maximum number of likelihood calls before a new bounds is created. If None, use 10 times n_live.
+ vectorized=False, # If True, the likelihood function can receive multiple input sets at once.
+ seed=None, # Seed for random number generation used for reproducible results accross different runs.
+ f_live=0.01, # Maximum fraction of the evidence contained in the live set before building the initial shells terminates.
+ n_shell=1, # Minimum number of points in each shell. The algorithm will sample from the shells until this is reached. Default is 1.
+ n_eff=500, # Minimum effective sample size. The algorithm will sample from the shells until this is reached. Default is 10000.
+ discard_exploration=False, # Whether to discard points drawn in the exploration phase. This is required for a fully unbiased posterior and evidence estimate.
+ verbose=True, # Whether to print information about the run.
+ n_like_max=np.inf, # Maximum number of likelihood evaluations. Regardless of progress, the sampler will stop if this value is reached. Default is infinity.
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+"""
+__Result__
+
+The result object returned by the fit provides information on the results of the non-linear search. Lets use it to
+compare the maximum log likelihood `Gaussian` to the data.
+"""
+model_data = result.max_log_likelihood_instance.model_data_from(
+ xvalues=np.arange(data.shape[0])
+)
+
+plt.errorbar(
+ x=range(data.shape[0]),
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(range(data.shape[0]), model_data, color="r")
+plt.title("Nautilus model fit to 1D Gaussian dataset.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
+
+"""
+__Search Internal__
+
+The result also contains the internal representation of the non-linear search.
+
+The internal representation of the non-linear search ensures that all sampling info is available in its native form.
+This can be passed to functions which take it as input, for example if the sampling package has bespoke visualization
+functions.
+
+For `DynestyStatic`, this is an instance of the `Sampler` object (`from nautilus import Sampler`).
+"""
+search_internal = result.search_internal
+
+print(search_internal)
+
+"""
+The internal search is by default not saved to hard-disk, because it can often take up quite a lot of hard-disk space
+(significantly more than standard output files).
+
+This means that the search internal will only be available the first time you run the search. If you rerun the code
+and the search is bypassed because the results already exist on hard-disk, the search internal will not be available.
+
+If you are frequently using the search internal you can have it saved to hard-disk by changing the `search_internal`
+setting in `output.yaml` to `True`. The result will then have the search internal available as an attribute,
+irrespective of whether the search is re-run or not.
+"""
diff --git a/scripts/searches/nest/UltraNest.py b/scripts/searches/nest/UltraNest.py
index f6fc3f89..f3a94438 100644
--- a/scripts/searches/nest/UltraNest.py
+++ b/scripts/searches/nest/UltraNest.py
@@ -1,163 +1,159 @@
-"""
-Searches: UltraNest
-=======================
-
-This example illustrates how to use the nested sampling algorithm UltraNest.
-
-UltraNest is an optional requirement and must be installed manually via the command `pip install ultranest`.
-It is optional as it has certain dependencies which are generally straight forward to install (e.g. Cython).
-
-Information about UltraNest can be found at the following links:
-
- - https://github.com/JohannesBuchner/UltraNest
- - https://johannesbuchner.github.io/UltraNest/readme.html
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Data**: Loading and plotting the 1D Gaussian dataset used to demonstrate the search.
-- **Model + Analysis**: Setting up the model and analysis for the fitting example.
-- **Search**: Configuring and running the UltraNest nested sampler.
-- **Result**: Inspecting the result and comparing the maximum log likelihood model to the data.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import matplotlib.pyplot as plt
-import numpy as np
-from os import path
-
-import autofit as af
-
-"""
-__Data__
-
-This example fits a single 1D Gaussian, we therefore load and plot data containing one Gaussian.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-plt.errorbar(
- x=range(data.shape[0]),
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.show()
-plt.close()
-
-"""
-__Model + Analysis__
-
-We create the model and analysis, which in this example is a single `Gaussian` and therefore has dimensionality N=3.
-"""
-model = af.Model(af.ex.Gaussian)
-
-model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.normalization = af.LogUniformPrior(lower_limit=1e-2, upper_limit=1e2)
-model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-"""
-__Search__
-
-We now create and run the `UltraNest` object which acts as our non-linear search.
-
-We manually specify all of the Dynesty settings, descriptions of which are provided at the following webpage:
-
-- https://johannesbuchner.github.io/UltraNest/readme.html
-- https://johannesbuchner.github.io/UltraNest/ultranest.html#ultranest.integrator.ReactiveNestedSampler
-
-"""
-search = af.UltraNest(
- path_prefix="searches",
- name="UltraNest",
- resume=True,
- run_num=None,
- num_test_samples=2,
- draw_multiple=True,
- num_bootstraps=30,
- vectorized=False,
- ndraw_min=128,
- ndraw_max=65536,
- storage_backend="hdf5",
- warmstart_max_tau=-1,
- update_interval_volume_fraction=0.8,
- update_interval_ncall=None,
- log_interval=None,
- show_status=True,
- viz_callback="auto",
- dlogz=0.5,
- dKL=0.5,
- frac_remain=0.01,
- Lepsilon=0.001,
- min_ess=400,
- max_iters=None,
- max_ncalls=None,
- max_num_improvement_loops=-1,
- min_num_live_points=50,
- cluster_num_live_points=40,
- insertion_test_window=10,
- insertion_test_zscore_threshold=2,
- stepsampler_cls="RegionMHSampler",
- nsteps=11,
- number_of_cores=1,
-)
-
-result = search.fit(model=model, analysis=analysis)
-
-"""
-__Result__
-
-The result object returned by the fit provides information on the results of the non-linear search. Lets use it to
-compare the maximum log likelihood `Gaussian` to the data.
-"""
-model_data = result.max_log_likelihood_instance.model_data_from(
- xvalues=np.arange(data.shape[0])
-)
-
-plt.errorbar(
- x=range(data.shape[0]),
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.plot(range(data.shape[0]), model_data, color="r")
-plt.title("UltraNest model fit to 1D Gaussian dataset.")
-plt.xlabel("x values of profile")
-plt.ylabel("Profile normalization")
-plt.show()
-plt.close()
+"""
+Searches: UltraNest
+=======================
+
+This example illustrates how to use the nested sampling algorithm UltraNest.
+
+UltraNest is an optional requirement and must be installed manually via the command `pip install ultranest`.
+It is optional as it has certain dependencies which are generally straight forward to install (e.g. Cython).
+
+Information about UltraNest can be found at the following links:
+
+ - https://github.com/JohannesBuchner/UltraNest
+ - https://johannesbuchner.github.io/UltraNest/readme.html
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Data**: Loading and plotting the 1D Gaussian dataset used to demonstrate the search.
+- **Model + Analysis**: Setting up the model and analysis for the fitting example.
+- **Search**: Configuring and running the UltraNest nested sampler.
+- **Result**: Inspecting the result and comparing the maximum log likelihood model to the data.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import matplotlib.pyplot as plt
+import numpy as np
+from os import path
+
+import autofit as af
+
+"""
+__Data__
+
+This example fits a single 1D Gaussian, we therefore load and plot data containing one Gaussian.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+plt.errorbar(
+ x=range(data.shape[0]),
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.show()
+plt.close()
+
+"""
+__Model + Analysis__
+
+We create the model and analysis, which in this example is a single `Gaussian` and therefore has dimensionality N=3.
+"""
+model = af.Model(af.ex.Gaussian)
+
+model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.normalization = af.LogUniformPrior(lower_limit=1e-2, upper_limit=1e2)
+model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+"""
+__Search__
+
+We now create and run the `UltraNest` object which acts as our non-linear search.
+
+We manually specify all of the Dynesty settings, descriptions of which are provided at the following webpage:
+
+- https://johannesbuchner.github.io/UltraNest/readme.html
+- https://johannesbuchner.github.io/UltraNest/ultranest.html#ultranest.integrator.ReactiveNestedSampler
+
+"""
+search = af.UltraNest(
+ path_prefix="searches",
+ name="UltraNest",
+ resume=True,
+ run_num=None,
+ num_test_samples=2,
+ draw_multiple=True,
+ num_bootstraps=30,
+ vectorized=False,
+ ndraw_min=128,
+ ndraw_max=65536,
+ storage_backend="hdf5",
+ warmstart_max_tau=-1,
+ update_interval_volume_fraction=0.8,
+ update_interval_ncall=None,
+ log_interval=None,
+ show_status=True,
+ viz_callback="auto",
+ dlogz=0.5,
+ dKL=0.5,
+ frac_remain=0.01,
+ Lepsilon=0.001,
+ min_ess=400,
+ max_iters=None,
+ max_ncalls=None,
+ max_num_improvement_loops=-1,
+ min_num_live_points=50,
+ cluster_num_live_points=40,
+ insertion_test_window=10,
+ insertion_test_zscore_threshold=2,
+ stepsampler_cls="RegionMHSampler",
+ nsteps=11,
+ number_of_cores=1,
+)
+
+result = search.fit(model=model, analysis=analysis)
+
+"""
+__Result__
+
+The result object returned by the fit provides information on the results of the non-linear search. Lets use it to
+compare the maximum log likelihood `Gaussian` to the data.
+"""
+model_data = result.max_log_likelihood_instance.model_data_from(
+ xvalues=np.arange(data.shape[0])
+)
+
+plt.errorbar(
+ x=range(data.shape[0]),
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.plot(range(data.shape[0]), model_data, color="r")
+plt.title("UltraNest model fit to 1D Gaussian dataset.")
+plt.xlabel("x values of profile")
+plt.ylabel("Profile normalization")
+plt.show()
+plt.close()
diff --git a/scripts/searches/start_point.py b/scripts/searches/start_point.py
index 5f4100fc..ea1a6f11 100644
--- a/scripts/searches/start_point.py
+++ b/scripts/searches/start_point.py
@@ -1,196 +1,192 @@
-"""
-Feature: Start Point
-====================
-
-For maximum likelihood estimator (MLE) and Markov Chain Monte Carlo (MCMC) non-linear searches, parameter space
-sampling is built around having a "location" in parameter space.
-
-This could simply be the parameters of the current maximum likelihood model in an MLE fit, or the locations of many
-walkers in parameter space (e.g. MCMC).
-
-For many model-fitting problems, we may have an expectation of where correct solutions lie in parameter space and
-therefore want our non-linear search to start near that location of parameter space. Alternatively, we may want to
-sample a specific region of parameter space, to determine what solutions look like there.
-
-The start-point API allows us to do this, by manually specifying the start-point of an MLE fit or the start-point of
-the walkers in an MCMC fit. Because nested sampling draws from priors, it cannot use the start-point API.
-
-__Comparison to Priors__
-
-Similar behaviour can be achieved by customizing the priors of a model-fit. We could place `TruncatedGaussianPrior`'s
-centred on the regions of parameter space we want to sample, or we could place tight `UniformPrior`'s on regions
-of parameter space we believe the correct answer lies.
-
-The downside of using priors is that our priors have a direct influence on the parameters we infer and the size
-of the inferred parameter errors. By using priors to control the location of our model-fit, we therefore risk
-inferring a non-representative model.
-
-For users more familiar with statistical inference, adjusting ones priors in the way described above leads to
-changes in the posterior, which therefore impacts the model inferred.
-
-__Example Source Code (`af.ex`)__
-
-The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
-
- - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
- `visualize` functions.
-
- - `Gaussian`: a model component representing a 1D Gaussian profile.
-
-These are functionally identical to the `Analysis` and `Gaussian` objects you have seen elsewhere in the workspace.
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Comparison to Priors**: Explain the differences between start-point and prior customization approaches.
-- **Example Source Code (`af.ex`)**: The example objects used in this script.
-- **Start Here Notebook**: Reference to the related tutorial notebook.
-- **Data**: Load and plot the 1D Gaussian dataset.
-- **Start Point Priors**: Define a model with broad uniform priors for start-point demonstration.
-- **Start Point**: Set parameter start point ranges for initializing the search.
-- **Search + Analysis + Model-Fit**: Perform the model-fit with the configured start point.
-- **Result**: Extract and display the initial walker samples and fit results.
-
-__Start Here Notebook__
-
-If any code in this script is unclear, refer to the `modeling/start_here.ipynb` notebook.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import matplotlib.pyplot as plt
-from os import path
-import autofit as af
-
-"""
-__Data__
-
-This example fits a single 1D Gaussian, we therefore load and plot data containing one Gaussian.
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-
-"""
-__Dataset Auto-Simulation__
-
-If the dataset does not already exist on your system, it will be created by running the corresponding
-simulator script. This ensures that all example scripts can be run without manually simulating data first.
-"""
-if not path.exists(dataset_path):
- import subprocess
- import sys
-
- subprocess.run(
- [sys.executable, "scripts/simulators/simulators.py"],
- check=True,
- )
-
-data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
-noise_map = af.util.numpy_array_from_json(
- file_path=path.join(dataset_path, "noise_map.json")
-)
-
-plt.errorbar(
- x=range(data.shape[0]),
- y=data,
- yerr=noise_map,
- linestyle="",
- color="k",
- ecolor="k",
- elinewidth=1,
- capsize=2,
-)
-plt.show()
-plt.close()
-
-"""
-__Start Point Priors__
-
-The start-point API does not conflict with the use of priors, which are still associated with every parameter.
-
-We manually customize the priors of the model used by the non-linear search.
-
-We use broad `UniformPriors`'s so that our priors do not impact our inferred model and errors (which would be
-the case with tight `TruncatedGaussianPrior`'s.
-"""
-model = af.Model(af.ex.Gaussian)
-
-model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
-model.normalization = af.UniformPrior(lower_limit=1e-2, upper_limit=1e2)
-model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
-
-"""
-We can inspect the model (with customized priors) via its `.info` attribute.
-"""
-print(model.info)
-
-
-"""
-__Start Point__
-
-We now define the start point of certain parameters in the model:
-
- - The 1D Gaussian is centred near pixel 50, so we set a start point there.
-
- - The sigma value of the Gaussian looks around 10, so we set a start point there.
-
-For all parameters where the start-point is not specified (in this case the `normalization`, their
-parameter values are drawn randomly from the prior when determining the initial locations of the parameters.
-"""
-initializer = af.InitializerParamBounds(
- {model.centre: (49.0, 51.0), model.sigma: (9.0, 11.0)}
-)
-
-"""
-A quick look at the model's `info` attribute shows that the starting points above do not change
-the priors or model info.
-"""
-print(model.info)
-
-"""
-Information on the initializer can be extracted and printed, which is shown below, where the start points are
-clearly visible.
-"""
-print(initializer.info_from_model(model=model))
-
-
-"""
-__Search + Analysis + Model-Fit__
-
-The code below performs the normal steps to set up a model-fit. We omit comments of this code as you should be
-familiar with it and it is not specific to this example!
-"""
-search = af.Emcee(
- path_prefix="searches",
- name="start_point",
- nwalkers=30,
- nsteps=1000,
- initializer=initializer,
- number_of_cores=1,
-)
-
-analysis = af.ex.Analysis(data=data, noise_map=noise_map)
-
-result = search.fit(model=model, analysis=analysis)
-
-"""
-__Result__
-
-We can print the initial `parameter_lists` of the result's `Samples` object to check that the initial
-walker samples were set within the start point ranges above.
-"""
-samples = result.samples
-
-print(samples.model.parameter_names)
-
-print(samples.parameter_lists[0])
-print(samples.parameter_lists[1])
-print(samples.parameter_lists[2])
-
-"""
-Finish.
-"""
+"""
+Feature: Start Point
+====================
+
+For maximum likelihood estimator (MLE) and Markov Chain Monte Carlo (MCMC) non-linear searches, parameter space
+sampling is built around having a "location" in parameter space.
+
+This could simply be the parameters of the current maximum likelihood model in an MLE fit, or the locations of many
+walkers in parameter space (e.g. MCMC).
+
+For many model-fitting problems, we may have an expectation of where correct solutions lie in parameter space and
+therefore want our non-linear search to start near that location of parameter space. Alternatively, we may want to
+sample a specific region of parameter space, to determine what solutions look like there.
+
+The start-point API allows us to do this, by manually specifying the start-point of an MLE fit or the start-point of
+the walkers in an MCMC fit. Because nested sampling draws from priors, it cannot use the start-point API.
+
+__Comparison to Priors__
+
+Similar behaviour can be achieved by customizing the priors of a model-fit. We could place `TruncatedGaussianPrior`'s
+centred on the regions of parameter space we want to sample, or we could place tight `UniformPrior`'s on regions
+of parameter space we believe the correct answer lies.
+
+The downside of using priors is that our priors have a direct influence on the parameters we infer and the size
+of the inferred parameter errors. By using priors to control the location of our model-fit, we therefore risk
+inferring a non-representative model.
+
+For users more familiar with statistical inference, adjusting ones priors in the way described above leads to
+changes in the posterior, which therefore impacts the model inferred.
+
+__Example Source Code (`af.ex`)__
+
+The **PyAutoFit** source code has the following example objects (accessed via `af.ex`) used in this tutorial:
+
+ - `Analysis`: an analysis object which fits noisy 1D datasets, including `log_likelihood_function` and
+ `visualize` functions.
+
+ - `Gaussian`: a model component representing a 1D Gaussian profile.
+
+These are functionally identical to the `Analysis` and `Gaussian` objects you have seen elsewhere in the workspace.
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Comparison to Priors**: Explain the differences between start-point and prior customization approaches.
+- **Example Source Code (`af.ex`)**: The example objects used in this script.
+- **Start Here Notebook**: Reference to the related tutorial notebook.
+- **Data**: Load and plot the 1D Gaussian dataset.
+- **Start Point Priors**: Define a model with broad uniform priors for start-point demonstration.
+- **Start Point**: Set parameter start point ranges for initializing the search.
+- **Search + Analysis + Model-Fit**: Perform the model-fit with the configured start point.
+- **Result**: Extract and display the initial walker samples and fit results.
+
+__Start Here Notebook__
+
+If any code in this script is unclear, refer to the `modeling/start_here.ipynb` notebook.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import matplotlib.pyplot as plt
+from os import path
+import autofit as af
+
+"""
+__Data__
+
+This example fits a single 1D Gaussian, we therefore load and plot data containing one Gaussian.
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+
+"""
+__Dataset Auto-Simulation__
+
+If the dataset does not already exist on your system, it will be created by running the corresponding
+simulator script. This ensures that all example scripts can be run without manually simulating data first.
+"""
+if not path.exists(dataset_path):
+ import subprocess
+ import sys
+
+ subprocess.run(
+ [sys.executable, "scripts/simulators/simulators.py"],
+ check=True,
+ )
+
+data = af.util.numpy_array_from_json(file_path=path.join(dataset_path, "data.json"))
+noise_map = af.util.numpy_array_from_json(
+ file_path=path.join(dataset_path, "noise_map.json")
+)
+
+plt.errorbar(
+ x=range(data.shape[0]),
+ y=data,
+ yerr=noise_map,
+ linestyle="",
+ color="k",
+ ecolor="k",
+ elinewidth=1,
+ capsize=2,
+)
+plt.show()
+plt.close()
+
+"""
+__Start Point Priors__
+
+The start-point API does not conflict with the use of priors, which are still associated with every parameter.
+
+We manually customize the priors of the model used by the non-linear search.
+
+We use broad `UniformPriors`'s so that our priors do not impact our inferred model and errors (which would be
+the case with tight `TruncatedGaussianPrior`'s.
+"""
+model = af.Model(af.ex.Gaussian)
+
+model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
+model.normalization = af.UniformPrior(lower_limit=1e-2, upper_limit=1e2)
+model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
+
+"""
+We can inspect the model (with customized priors) via its `.info` attribute.
+"""
+print(model.info)
+
+
+"""
+__Start Point__
+
+We now define the start point of certain parameters in the model:
+
+ - The 1D Gaussian is centred near pixel 50, so we set a start point there.
+
+ - The sigma value of the Gaussian looks around 10, so we set a start point there.
+
+For all parameters where the start-point is not specified (in this case the `normalization`, their
+parameter values are drawn randomly from the prior when determining the initial locations of the parameters.
+"""
+initializer = af.InitializerParamBounds(
+ {model.centre: (49.0, 51.0), model.sigma: (9.0, 11.0)}
+)
+
+"""
+A quick look at the model's `info` attribute shows that the starting points above do not change
+the priors or model info.
+"""
+print(model.info)
+
+"""
+Information on the initializer can be extracted and printed, which is shown below, where the start points are
+clearly visible.
+"""
+print(initializer.info_from_model(model=model))
+
+
+"""
+__Search + Analysis + Model-Fit__
+
+The code below performs the normal steps to set up a model-fit. We omit comments of this code as you should be
+familiar with it and it is not specific to this example!
+"""
+search = af.Emcee(
+ path_prefix="searches",
+ name="start_point",
+ nwalkers=30,
+ nsteps=1000,
+ initializer=initializer,
+ number_of_cores=1,
+)
+
+analysis = af.ex.Analysis(data=data, noise_map=noise_map)
+
+result = search.fit(model=model, analysis=analysis)
+
+"""
+__Result__
+
+We can print the initial `parameter_lists` of the result's `Samples` object to check that the initial
+walker samples were set within the start point ranges above.
+"""
+samples = result.samples
+
+print(samples.model.parameter_names)
+
+print(samples.parameter_lists[0])
+print(samples.parameter_lists[1])
+print(samples.parameter_lists[2])
+
+"""
+Finish.
+"""
diff --git a/scripts/simulators/simulators.py b/scripts/simulators/simulators.py
index b4fadfbd..14b8a00e 100644
--- a/scripts/simulators/simulators.py
+++ b/scripts/simulators/simulators.py
@@ -1,215 +1,211 @@
-"""
-__Simulators__
-
-These scripts simulate the 1D Gaussian datasets used to demonstrate model-fitting.
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Gaussian x1**: Simulate a single 1D Gaussian dataset.
-- **Gaussian x1 (0)**: Simulate a single Gaussian with sigma=1.0.
-- **Gaussian x1 (1)**: Simulate a single Gaussian with sigma=5.0.
-- **Gaussian x1 (2)**: Simulate a single Gaussian with sigma=10.0.
-- **Gaussian x1 (Identical 0)**: Simulate an identical single Gaussian dataset (copy 0).
-- **Gaussian x1 (Identical 1)**: Simulate an identical single Gaussian dataset (copy 1).
-- **Gaussian x1 (Identical 2)**: Simulate an identical single Gaussian dataset (copy 2).
-- **Gaussian x1 + Exponential x1**: Simulate a dataset with one Gaussian and one Exponential.
-- **Gaussian x2 + Exponential x1**: Simulate a dataset with two Gaussians and one Exponential.
-- **Gaussian x2**: Simulate a dataset with two Gaussians.
-- **Gaussian x3**: Simulate a dataset with three Gaussians.
-- **Gaussian x5**: Simulate a dataset with five Gaussians.
-- **Gaussian x1 unconvolved**: Simulate a single Gaussian without convolution.
-- **Gaussian x1 convolved**: Simulate a single Gaussian with kernel convolution.
-- **Gaussian x1 with feature**: Simulate a Gaussian with a small feature bump.
-- **Gaussian x2 split**: Simulate two separated Gaussians.
-- **Gaussian x1 time**: Simulate time-varying Gaussian datasets.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import util
-from os import path
-
-import autofit as af
-
-"""
-__Gaussian x1__
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
-gaussian = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=10.0)
-util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
-
-"""
-__Gaussian x1 (0)__
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1_0")
-gaussian = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=1.0)
-util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
-
-"""
-__Gaussian x1 (1)__
-"""
-gaussian = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=5.0)
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1_1")
-util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
-
-"""
-__Gaussian x1 (2)__
-"""
-gaussian = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=10.0)
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1_2")
-util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
-
-"""
-__Gaussian x1 (Identical 0)__
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1_identical_0")
-gaussian = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=10.0)
-util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
-
-"""
-__Gaussian x1 (Identical 1)__
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1_identical_1")
-gaussian = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=10.0)
-util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
-
-"""
-__Gaussian x1 (Identical 2)__
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1_identical_2")
-gaussian = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=10.0)
-util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
-
-"""
-__Gaussian x1 + Exponential x1__
-"""
-gaussian = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=10.0)
-exponential = af.ex.Exponential(centre=50.0, normalization=40.0, rate=0.05)
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1__exponential_x1")
-util.simulate_dataset_1d_via_profile_1d_list_from(
- profile_1d_list=[gaussian, exponential], dataset_path=dataset_path
-)
-
-"""
-__Gaussian x2 + Exponential x1__
-"""
-gaussian_0 = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=10.0)
-gaussian_1 = af.ex.Gaussian(centre=20.0, normalization=30.0, sigma=5.0)
-exponential = af.ex.Exponential(centre=70.0, normalization=40.0, rate=0.005)
-dataset_path = path.join("dataset", "example_1d", "gaussian_x2__exponential_x1")
-util.simulate_dataset_1d_via_profile_1d_list_from(
- profile_1d_list=[gaussian_0, gaussian_1, exponential], dataset_path=dataset_path
-)
-
-"""
-__Gaussian x2__
-"""
-gaussian_0 = af.ex.Gaussian(centre=50.0, normalization=20.0, sigma=1.0)
-gaussian_1 = af.ex.Gaussian(centre=50.0, normalization=40.0, sigma=5.0)
-dataset_path = path.join("dataset", "example_1d", "gaussian_x2")
-util.simulate_dataset_1d_via_profile_1d_list_from(
- profile_1d_list=[gaussian_0, gaussian_1], dataset_path=dataset_path
-)
-
-"""
-__Gaussian x3__
-"""
-gaussian_0 = af.ex.Gaussian(centre=50.0, normalization=20.0, sigma=1.0)
-gaussian_1 = af.ex.Gaussian(centre=50.0, normalization=40.0, sigma=5.0)
-gaussian_2 = af.ex.Gaussian(centre=50.0, normalization=60.0, sigma=10.0)
-dataset_path = path.join("dataset", "example_1d", "gaussian_x3")
-util.simulate_dataset_1d_via_profile_1d_list_from(
- profile_1d_list=[gaussian_0, gaussian_1, gaussian_2], dataset_path=dataset_path
-)
-
-"""
-__Gaussian x5__
-"""
-gaussian_0 = af.ex.Gaussian(centre=50.0, normalization=20.0, sigma=1.0)
-gaussian_1 = af.ex.Gaussian(centre=50.0, normalization=40.0, sigma=5.0)
-gaussian_2 = af.ex.Gaussian(centre=50.0, normalization=60.0, sigma=10.0)
-gaussian_3 = af.ex.Gaussian(centre=50.0, normalization=80.0, sigma=15.0)
-gaussian_4 = af.ex.Gaussian(centre=50.0, normalization=100.0, sigma=20.0)
-dataset_path = path.join("dataset", "example_1d", "gaussian_x5")
-util.simulate_dataset_1d_via_profile_1d_list_from(
- profile_1d_list=[gaussian_0, gaussian_1, gaussian_2, gaussian_3, gaussian_4],
- dataset_path=dataset_path,
-)
-
-"""
-__Gaussian x1 unconvolved__
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1_unconvolved")
-gaussian = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=3.0)
-util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
-
-"""
-__Gaussian x1 convolved__
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1_convolved")
-gaussian = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=3.0)
-util.simulate_data_1d_with_kernel_via_gaussian_from(
- gaussian=gaussian, dataset_path=dataset_path
-)
-
-"""
-__Gaussian x1 with feature__
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1_with_feature")
-gaussian = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=10.0)
-gaussian_feature = af.ex.Gaussian(centre=70.0, normalization=0.3, sigma=0.5)
-util.simulate_dataset_1d_via_profile_1d_list_from(
- profile_1d_list=[gaussian, gaussian_feature], dataset_path=dataset_path
-)
-
-"""
-__Gaussian x2 split__
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x2_split")
-gaussian_0 = af.ex.Gaussian(centre=25.0, normalization=50.0, sigma=12.5)
-gaussian_1 = af.ex.Gaussian(centre=75.0, normalization=50.0, sigma=12.5)
-util.simulate_dataset_1d_via_profile_1d_list_from(
- profile_1d_list=[gaussian_0, gaussian_1], dataset_path=dataset_path
-)
-
-
-"""
-__Gaussian x1 time__
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1_time", "time_0")
-gaussian = af.ex.Gaussian(centre=40.0, normalization=50.0, sigma=20.0)
-util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
-
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1_time", "time_1")
-gaussian = af.ex.Gaussian(centre=50.0, normalization=50.0, sigma=20.0)
-util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
-
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1_time", "time_2")
-gaussian = af.ex.Gaussian(centre=60.0, normalization=50.0, sigma=20.0)
-util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
-
-
-"""
-__Gaussian x1 time__
-"""
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1_variable", "sigma_0")
-gaussian = af.ex.Gaussian(centre=50.0, normalization=50.0, sigma=10.0)
-util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
-
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1_variable", "sigma_1")
-gaussian = af.ex.Gaussian(centre=50.0, normalization=50.0, sigma=20.0)
-util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
-
-dataset_path = path.join("dataset", "example_1d", "gaussian_x1_variable", "sigma_2")
-gaussian = af.ex.Gaussian(centre=50.0, normalization=50.0, sigma=30.0)
-util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
-
-"""
-Finish.
-"""
+"""
+__Simulators__
+
+These scripts simulate the 1D Gaussian datasets used to demonstrate model-fitting.
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Gaussian x1**: Simulate a single 1D Gaussian dataset.
+- **Gaussian x1 (0)**: Simulate a single Gaussian with sigma=1.0.
+- **Gaussian x1 (1)**: Simulate a single Gaussian with sigma=5.0.
+- **Gaussian x1 (2)**: Simulate a single Gaussian with sigma=10.0.
+- **Gaussian x1 (Identical 0)**: Simulate an identical single Gaussian dataset (copy 0).
+- **Gaussian x1 (Identical 1)**: Simulate an identical single Gaussian dataset (copy 1).
+- **Gaussian x1 (Identical 2)**: Simulate an identical single Gaussian dataset (copy 2).
+- **Gaussian x1 + Exponential x1**: Simulate a dataset with one Gaussian and one Exponential.
+- **Gaussian x2 + Exponential x1**: Simulate a dataset with two Gaussians and one Exponential.
+- **Gaussian x2**: Simulate a dataset with two Gaussians.
+- **Gaussian x3**: Simulate a dataset with three Gaussians.
+- **Gaussian x5**: Simulate a dataset with five Gaussians.
+- **Gaussian x1 unconvolved**: Simulate a single Gaussian without convolution.
+- **Gaussian x1 convolved**: Simulate a single Gaussian with kernel convolution.
+- **Gaussian x1 with feature**: Simulate a Gaussian with a small feature bump.
+- **Gaussian x2 split**: Simulate two separated Gaussians.
+- **Gaussian x1 time**: Simulate time-varying Gaussian datasets.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import util
+from os import path
+
+import autofit as af
+
+"""
+__Gaussian x1__
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1")
+gaussian = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=10.0)
+util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
+
+"""
+__Gaussian x1 (0)__
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1_0")
+gaussian = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=1.0)
+util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
+
+"""
+__Gaussian x1 (1)__
+"""
+gaussian = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=5.0)
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1_1")
+util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
+
+"""
+__Gaussian x1 (2)__
+"""
+gaussian = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=10.0)
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1_2")
+util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
+
+"""
+__Gaussian x1 (Identical 0)__
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1_identical_0")
+gaussian = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=10.0)
+util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
+
+"""
+__Gaussian x1 (Identical 1)__
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1_identical_1")
+gaussian = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=10.0)
+util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
+
+"""
+__Gaussian x1 (Identical 2)__
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1_identical_2")
+gaussian = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=10.0)
+util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
+
+"""
+__Gaussian x1 + Exponential x1__
+"""
+gaussian = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=10.0)
+exponential = af.ex.Exponential(centre=50.0, normalization=40.0, rate=0.05)
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1__exponential_x1")
+util.simulate_dataset_1d_via_profile_1d_list_from(
+ profile_1d_list=[gaussian, exponential], dataset_path=dataset_path
+)
+
+"""
+__Gaussian x2 + Exponential x1__
+"""
+gaussian_0 = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=10.0)
+gaussian_1 = af.ex.Gaussian(centre=20.0, normalization=30.0, sigma=5.0)
+exponential = af.ex.Exponential(centre=70.0, normalization=40.0, rate=0.005)
+dataset_path = path.join("dataset", "example_1d", "gaussian_x2__exponential_x1")
+util.simulate_dataset_1d_via_profile_1d_list_from(
+ profile_1d_list=[gaussian_0, gaussian_1, exponential], dataset_path=dataset_path
+)
+
+"""
+__Gaussian x2__
+"""
+gaussian_0 = af.ex.Gaussian(centre=50.0, normalization=20.0, sigma=1.0)
+gaussian_1 = af.ex.Gaussian(centre=50.0, normalization=40.0, sigma=5.0)
+dataset_path = path.join("dataset", "example_1d", "gaussian_x2")
+util.simulate_dataset_1d_via_profile_1d_list_from(
+ profile_1d_list=[gaussian_0, gaussian_1], dataset_path=dataset_path
+)
+
+"""
+__Gaussian x3__
+"""
+gaussian_0 = af.ex.Gaussian(centre=50.0, normalization=20.0, sigma=1.0)
+gaussian_1 = af.ex.Gaussian(centre=50.0, normalization=40.0, sigma=5.0)
+gaussian_2 = af.ex.Gaussian(centre=50.0, normalization=60.0, sigma=10.0)
+dataset_path = path.join("dataset", "example_1d", "gaussian_x3")
+util.simulate_dataset_1d_via_profile_1d_list_from(
+ profile_1d_list=[gaussian_0, gaussian_1, gaussian_2], dataset_path=dataset_path
+)
+
+"""
+__Gaussian x5__
+"""
+gaussian_0 = af.ex.Gaussian(centre=50.0, normalization=20.0, sigma=1.0)
+gaussian_1 = af.ex.Gaussian(centre=50.0, normalization=40.0, sigma=5.0)
+gaussian_2 = af.ex.Gaussian(centre=50.0, normalization=60.0, sigma=10.0)
+gaussian_3 = af.ex.Gaussian(centre=50.0, normalization=80.0, sigma=15.0)
+gaussian_4 = af.ex.Gaussian(centre=50.0, normalization=100.0, sigma=20.0)
+dataset_path = path.join("dataset", "example_1d", "gaussian_x5")
+util.simulate_dataset_1d_via_profile_1d_list_from(
+ profile_1d_list=[gaussian_0, gaussian_1, gaussian_2, gaussian_3, gaussian_4],
+ dataset_path=dataset_path,
+)
+
+"""
+__Gaussian x1 unconvolved__
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1_unconvolved")
+gaussian = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=3.0)
+util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
+
+"""
+__Gaussian x1 convolved__
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1_convolved")
+gaussian = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=3.0)
+util.simulate_data_1d_with_kernel_via_gaussian_from(
+ gaussian=gaussian, dataset_path=dataset_path
+)
+
+"""
+__Gaussian x1 with feature__
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1_with_feature")
+gaussian = af.ex.Gaussian(centre=50.0, normalization=25.0, sigma=10.0)
+gaussian_feature = af.ex.Gaussian(centre=70.0, normalization=0.3, sigma=0.5)
+util.simulate_dataset_1d_via_profile_1d_list_from(
+ profile_1d_list=[gaussian, gaussian_feature], dataset_path=dataset_path
+)
+
+"""
+__Gaussian x2 split__
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x2_split")
+gaussian_0 = af.ex.Gaussian(centre=25.0, normalization=50.0, sigma=12.5)
+gaussian_1 = af.ex.Gaussian(centre=75.0, normalization=50.0, sigma=12.5)
+util.simulate_dataset_1d_via_profile_1d_list_from(
+ profile_1d_list=[gaussian_0, gaussian_1], dataset_path=dataset_path
+)
+
+
+"""
+__Gaussian x1 time__
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1_time", "time_0")
+gaussian = af.ex.Gaussian(centre=40.0, normalization=50.0, sigma=20.0)
+util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
+
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1_time", "time_1")
+gaussian = af.ex.Gaussian(centre=50.0, normalization=50.0, sigma=20.0)
+util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
+
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1_time", "time_2")
+gaussian = af.ex.Gaussian(centre=60.0, normalization=50.0, sigma=20.0)
+util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
+
+
+"""
+__Gaussian x1 time__
+"""
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1_variable", "sigma_0")
+gaussian = af.ex.Gaussian(centre=50.0, normalization=50.0, sigma=10.0)
+util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
+
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1_variable", "sigma_1")
+gaussian = af.ex.Gaussian(centre=50.0, normalization=50.0, sigma=20.0)
+util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
+
+dataset_path = path.join("dataset", "example_1d", "gaussian_x1_variable", "sigma_2")
+gaussian = af.ex.Gaussian(centre=50.0, normalization=50.0, sigma=30.0)
+util.simulate_dataset_1d_via_gaussian_from(gaussian=gaussian, dataset_path=dataset_path)
+
+"""
+Finish.
+"""
diff --git a/scripts/simulators/simulators_sample.py b/scripts/simulators/simulators_sample.py
index bbea0f31..b785f10d 100644
--- a/scripts/simulators/simulators_sample.py
+++ b/scripts/simulators/simulators_sample.py
@@ -1,113 +1,109 @@
-"""
-__Simulators__
-
-These scripts simulates many 1D Gaussian datasets with a low signal to noise ratio, which are used to demonstrate
-model-fitting.
-
-__Contents__
-
-This script is split into the following sections:
-
-- **Gaussian x1 low snr (centre fixed to 50.0)**: Simulate low signal-to-noise Gaussian datasets with a fixed centre.
-- **Gaussian x1 low snr (centre drawn from parent Gaussian distribution to 50.0)**: Simulate hierarchical Gaussian datasets with centres drawn from a parent distribution.
-- **Gaussian x2 offset centre**: Simulate datasets with two Gaussians with offset centres for graphical model demonstrations.
-"""
-
-# %matplotlib inline
-# from pyprojroot import here
-# workspace_path = str(here())
-# %cd $workspace_path
-# print(f"Working Directory has been set to `{workspace_path}`")
-
-import numpy as np
-from os import path
-
-import autofit as af
-import util
-
-"""
-__Gaussian x1 low snr (centre fixed to 50.0)__
-
-This is used for demonstrating expectation propagation, whereby a shared `centre` parameter is inferred from a sample
-of `total_datasets` 1D Gaussian datasets.
-"""
-total_datasets = 50
-
-for i in range(total_datasets):
- dataset_path = path.join(
- "dataset", "example_1d", f"gaussian_x1__low_snr", f"dataset_{i}"
- )
- gaussian = af.ex.Gaussian(centre=50.0, normalization=0.5, sigma=5.0)
- util.simulate_dataset_1d_via_gaussian_from(
- gaussian=gaussian, dataset_path=dataset_path
- )
-
-"""
-__Gaussian x1 low snr (centre drawn from parent Gaussian distribution to 50.0)__
-
-This is used for demonstrating expectation propagation and hierachical modeling, whereby a the `centre` parameters
-of a sample of `total_datasets` 1D Gaussian datasets are drawn from a Gaussian distribution.
-"""
-
-total_datasets = 10
-
-gaussian_parent_model = af.Model(
- af.ex.Gaussian,
- centre=af.TruncatedGaussianPrior(
- mean=50.0, sigma=10.0, lower_limit=0.0, upper_limit=100.0
- ),
- normalization=0.5,
- sigma=5.0,
-)
-
-for i in range(total_datasets):
- dataset_path = path.join(
- "dataset", "example_1d", f"gaussian_x1__hierarchical", f"dataset_{i}"
- )
-
- gaussian = gaussian_parent_model.random_instance()
-
- util.simulate_dataset_1d_via_gaussian_from(
- gaussian=gaussian, dataset_path=dataset_path
- )
-
-
-"""
-__Gaussian x2 offset centre__
-
-This is used for demonstrating the benefits of graphical models over fitting one-by-one, because it creates a
-degeneracy in the offset of the centres of the two Gaussians.
-"""
-total_datasets = 10
-
-for i in range(total_datasets):
- dataset_path = path.join(
- "dataset", "example_1d", f"gaussian_x2__offset_centres", f"dataset_{i}"
- )
-
- sigma_0_prior = af.TruncatedGaussianPrior(
- lower_limit=0.0, upper_limit=20.0, mean=10.0, sigma=10.0
- )
- while True:
- try:
- sigma_0_value = sigma_0_prior.value_for(unit=np.random.random(1))
- break
- except af.exc.PriorLimitException:
- continue
-
- sigma_1_prior = af.TruncatedGaussianPrior(
- lower_limit=0.0, upper_limit=20.0, mean=10.0, sigma=10.0
- )
- while True:
- try:
- sigma_1_value = sigma_1_prior.value_for(unit=np.random.random(1))
- break
- except af.exc.PriorLimitException:
- continue
-
- gaussian_0 = af.ex.Gaussian(centre=40.0, normalization=1.0, sigma=sigma_0_value)
- gaussian_1 = af.ex.Gaussian(centre=60.0, normalization=1.0, sigma=sigma_1_value)
-
- util.simulate_dataset_1d_via_profile_1d_list_from(
- profile_1d_list=[gaussian_0, gaussian_1], dataset_path=dataset_path
- )
+"""
+__Simulators__
+
+These scripts simulates many 1D Gaussian datasets with a low signal to noise ratio, which are used to demonstrate
+model-fitting.
+
+__Contents__
+
+This script is split into the following sections:
+
+- **Gaussian x1 low snr (centre fixed to 50.0)**: Simulate low signal-to-noise Gaussian datasets with a fixed centre.
+- **Gaussian x1 low snr (centre drawn from parent Gaussian distribution to 50.0)**: Simulate hierarchical Gaussian datasets with centres drawn from a parent distribution.
+- **Gaussian x2 offset centre**: Simulate datasets with two Gaussians with offset centres for graphical model demonstrations.
+"""
+
+# from autoconf import setup_notebook; setup_notebook()
+
+import numpy as np
+from os import path
+
+import autofit as af
+import util
+
+"""
+__Gaussian x1 low snr (centre fixed to 50.0)__
+
+This is used for demonstrating expectation propagation, whereby a shared `centre` parameter is inferred from a sample
+of `total_datasets` 1D Gaussian datasets.
+"""
+total_datasets = 50
+
+for i in range(total_datasets):
+ dataset_path = path.join(
+ "dataset", "example_1d", f"gaussian_x1__low_snr", f"dataset_{i}"
+ )
+ gaussian = af.ex.Gaussian(centre=50.0, normalization=0.5, sigma=5.0)
+ util.simulate_dataset_1d_via_gaussian_from(
+ gaussian=gaussian, dataset_path=dataset_path
+ )
+
+"""
+__Gaussian x1 low snr (centre drawn from parent Gaussian distribution to 50.0)__
+
+This is used for demonstrating expectation propagation and hierachical modeling, whereby a the `centre` parameters
+of a sample of `total_datasets` 1D Gaussian datasets are drawn from a Gaussian distribution.
+"""
+
+total_datasets = 10
+
+gaussian_parent_model = af.Model(
+ af.ex.Gaussian,
+ centre=af.TruncatedGaussianPrior(
+ mean=50.0, sigma=10.0, lower_limit=0.0, upper_limit=100.0
+ ),
+ normalization=0.5,
+ sigma=5.0,
+)
+
+for i in range(total_datasets):
+ dataset_path = path.join(
+ "dataset", "example_1d", f"gaussian_x1__hierarchical", f"dataset_{i}"
+ )
+
+ gaussian = gaussian_parent_model.random_instance()
+
+ util.simulate_dataset_1d_via_gaussian_from(
+ gaussian=gaussian, dataset_path=dataset_path
+ )
+
+
+"""
+__Gaussian x2 offset centre__
+
+This is used for demonstrating the benefits of graphical models over fitting one-by-one, because it creates a
+degeneracy in the offset of the centres of the two Gaussians.
+"""
+total_datasets = 10
+
+for i in range(total_datasets):
+ dataset_path = path.join(
+ "dataset", "example_1d", f"gaussian_x2__offset_centres", f"dataset_{i}"
+ )
+
+ sigma_0_prior = af.TruncatedGaussianPrior(
+ lower_limit=0.0, upper_limit=20.0, mean=10.0, sigma=10.0
+ )
+ while True:
+ try:
+ sigma_0_value = sigma_0_prior.value_for(unit=np.random.random(1))
+ break
+ except af.exc.PriorLimitException:
+ continue
+
+ sigma_1_prior = af.TruncatedGaussianPrior(
+ lower_limit=0.0, upper_limit=20.0, mean=10.0, sigma=10.0
+ )
+ while True:
+ try:
+ sigma_1_value = sigma_1_prior.value_for(unit=np.random.random(1))
+ break
+ except af.exc.PriorLimitException:
+ continue
+
+ gaussian_0 = af.ex.Gaussian(centre=40.0, normalization=1.0, sigma=sigma_0_value)
+ gaussian_1 = af.ex.Gaussian(centre=60.0, normalization=1.0, sigma=sigma_1_value)
+
+ util.simulate_dataset_1d_via_profile_1d_list_from(
+ profile_1d_list=[gaussian_0, gaussian_1], dataset_path=dataset_path
+ )























